1KEB
 
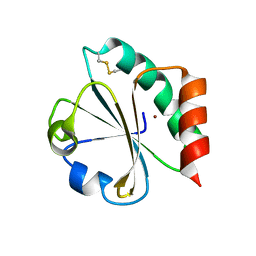 | | Crystal Structure of Double Mutant M37L,P40S E.coli Thioredoxin | | Descriptor: | COPPER (II) ION, Thioredoxin 1 | | Authors: | Rudresh, Jain, R, Dani, V, Mitra, A, Srivastava, S, Sarma, S.P, Varadarajan, R, Ramakumar, S. | | Deposit date: | 2001-11-15 | | Release date: | 2002-11-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Consequences of Replacement of an alpha-helical Pro Residue in E.coli Thioredoxin
PROTEIN ENG., 15, 2002
|
|
3AKC
 
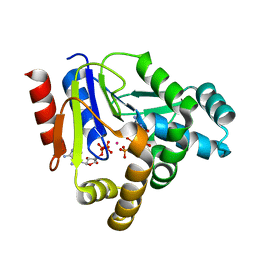 | | Crystal structure of CMP kinase in complex with CDP and ADP from Thermus thermophilus HB8 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CYTIDINE-5'-DIPHOSPHATE, Cytidylate kinase, ... | | Authors: | Mega, R, Nakagawa, N, Kuramitsu, S, Masui, R. | | Deposit date: | 2010-07-12 | | Release date: | 2011-07-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The crystal structure of the tertiary complex of CMP kinase with a phosphoryl group acceptor and a donor from Thermus thermophilus HB8
To be Published
|
|
1KPK
 
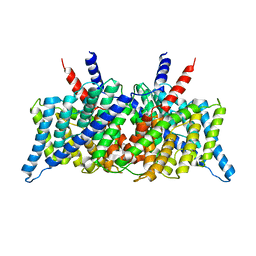 | | Crystal Structure of the ClC Chloride Channel from E. coli | | Descriptor: | putative channel transporter | | Authors: | Dutzler, R, Campbell, E.B, Cadene, M, Chait, B.T, MacKinnon, R. | | Deposit date: | 2001-12-31 | | Release date: | 2002-01-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | X-ray structure of a ClC chloride channel at 3.0 A reveals the molecular basis of
anion selectivity.
Nature, 415, 2002
|
|
1KPL
 
 | | Crystal Structure of the ClC Chloride Channel from S. typhimurium | | Descriptor: | CHLORIDE ION, N-OCTANE, PENTADECANE, ... | | Authors: | Dutzler, R, Campbell, E.B, Cadene, M, Chait, B.T, MacKinnon, R. | | Deposit date: | 2001-12-31 | | Release date: | 2002-01-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | X-ray structure of a ClC chloride channel at 3.0 A reveals the molecular basis of
anion selectivity.
Nature, 415, 2002
|
|
2Y8Q
 
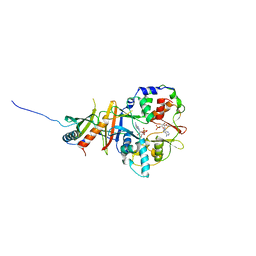 | | Structure of the regulatory fragment of mammalian AMPK in complex with one ADP | | Descriptor: | 5'-AMP-ACTIVATED PROTEIN KINASE CATALYTIC SUBUNIT ALPHA-1, 5'-AMP-ACTIVATED PROTEIN KINASE SUBUNIT BETA-2, 5'-AMP-ACTIVATED PROTEIN KINASE SUBUNIT GAMMA-1, ... | | Authors: | Xiao, B, Sanders, M.J, Underwood, E, Heath, R, Mayer, F, Carmena, D, Jing, C, Walker, P.A, Eccleston, J.F, Haire, L.F, Saiu, P, Howell, S.A, Aasland, R, Martin, S.R, Carling, D, Gamblin, S.J. | | Deposit date: | 2011-02-09 | | Release date: | 2011-03-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of Mammalian Ampk and its Regulation by Adp
Nature, 472, 2011
|
|
7VWW
 
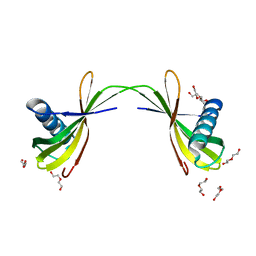 | |
1JV2
 
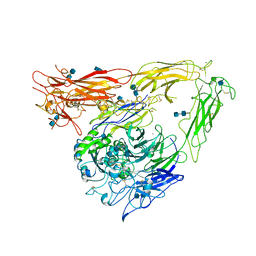 | | CRYSTAL STRUCTURE OF THE EXTRACELLULAR SEGMENT OF INTEGRIN ALPHAVBETA3 | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Xiong, J.P, Stehle, T, Diefenbach, B, Zhang, R, Dunker, R, Scott, D, Joachimiak, A, Goodman, S.L, Arnaout, M.A. | | Deposit date: | 2001-08-28 | | Release date: | 2001-10-17 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of the extracellular segment of integrin alpha Vbeta3.
Science, 294, 2001
|
|
1JZY
 
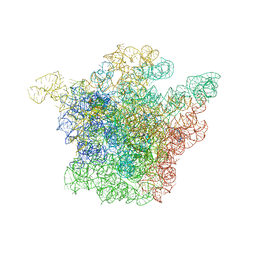 | | Structural Basis for the Interaction of Antibiotics with the Peptidyl Transferase Center in Eubacteria | | Descriptor: | 23S rRNA, ERYTHROMYCIN A, MAGNESIUM ION, ... | | Authors: | Schluenzen, F, Zarivach, R, Harms, J, Bashan, A, Tocilj, A, Albrecht, R, Yonath, A, Franceschi, F. | | Deposit date: | 2001-09-17 | | Release date: | 2001-10-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria.
Nature, 413, 2001
|
|
1JFX
 
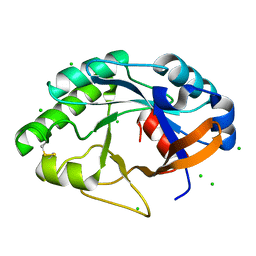 | | Crystal structure of the bacterial lysozyme from Streptomyces coelicolor at 1.65 A resolution | | Descriptor: | 1,4-beta-N-Acetylmuramidase M1, CHLORIDE ION | | Authors: | Rau, A, Hogg, T, Marquardt, R, Hilgenfeld, R. | | Deposit date: | 2001-06-22 | | Release date: | 2001-09-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A new lysozyme fold. Crystal structure of the muramidase from Streptomyces coelicolor at 1.65 A resolution.
J.Biol.Chem., 276, 2001
|
|
1JV5
 
 | | Anti-blood group A Fv | | Descriptor: | Ig chain heavy chain precursor V region, Ig kappa chain precursor V region | | Authors: | Thomas, R, Patenaude, S.I, MacKenzie, C.R, To, R, Hirama, T, Young, N.M, Evans, S.V. | | Deposit date: | 2001-08-28 | | Release date: | 2002-01-09 | | Last modified: | 2018-04-18 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of an anti-blood group A Fv and improvement of its binding affinity without loss of specificity.
J.Biol.Chem., 277, 2002
|
|
1P03
 
 | |
6IDH
 
 | | Antibody 64M-5 Fab in ligand-free form | | Descriptor: | Anti-(6-4) photoproduct antibody 64M-5 Fab (heavy chain), Anti-(6-4) photoproduct antibody 64M-5 Fab (light chain) | | Authors: | Yokoyama, H, Mizutani, R, Noguchi, S, Hayashida, N. | | Deposit date: | 2018-09-10 | | Release date: | 2019-02-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structures of the antibody 64M-5 Fab and its complex with dT(6-4)T indicate induced-fit and high-affinity mechanisms.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
1P04
 | |
1P06
 
 | |
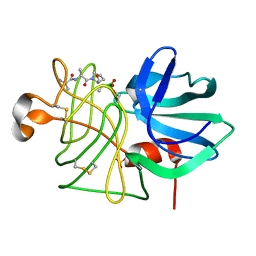
1P02
 
 | |
2ZIG
 
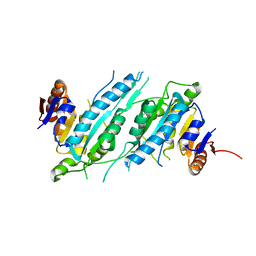 | | Crystal Structure of TTHA0409, Putative DNA Modification Methylase from Thermus thermophilus HB8 | | Descriptor: | Putative modification methylase | | Authors: | Morita, R, Ishikawa, H, Nakagawa, N, Masui, R, Yokoyama, S, Kuramitsu, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2008-02-15 | | Release date: | 2008-07-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of a putative DNA methylase TTHA0409 from Thermus thermophilus HB8
Proteins, 73, 2008
|
|
4MY6
 
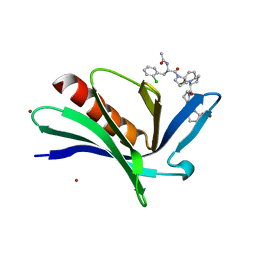 | | EnaH-EVH1 in complex with peptidomimetic low-molecular weight inhibitor Ac-[2-Cl-F]-[ProM-2]-[ProM-1]-OH | | Descriptor: | (3aR,5aS,8S,10aS)-1-[(3S,6R,8aS)-1'-[(2S)-2-acetamido-3-(2-chlorophenyl)propanoyl]-5-oxidanylidene-spiro[1,2,3,8a-tetrahydroindolizine-6,2'-pyrrolidine]-3-yl]carbonyl-10-oxidanylidene-2,3,3a,5a,8,10a-hexahydrodipyrrolo[3,2-b:3',1'-f]azepine-8-carboxylic acid, BROMIDE ION, Protein enabled homolog | | Authors: | Barone, M, Roske, Y, Kuehne, R. | | Deposit date: | 2013-09-27 | | Release date: | 2014-10-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A modular toolkit to inhibit proline-rich motif-mediated protein-protein interactions.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
5EUU
 
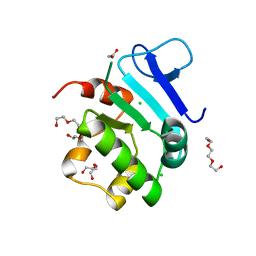 | | Rat prestin STAS domain in complex with chloride | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Prestin,Prestin, ... | | Authors: | Lolli, G, Pasqualetto, E, Costanzi, E, Bonetto, G, Battistutta, R. | | Deposit date: | 2015-11-19 | | Release date: | 2015-12-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | The STAS domain of mammalian SLC26A5 prestin harbours an anion-binding site.
Biochem.J., 473, 2016
|
|
3ZXI
 
 | | Crystal structure of human mitochondrial tyrosyl-tRNA synthetase in complex with a tyrosyl-adenylate analog | | Descriptor: | PHOSPHORIC ACID 2-AMINO-3-(4-HYDROXY-PHENYL)-PROPYL ESTER ADENOSIN-5'YL ESTER, TYROSYL-TRNA SYNTHETASE, MITOCHONDRIAL | | Authors: | Bonnefond, L, Frugier, M, Rudinger-Thirion, J, Balg, C, Chenevert, R, Lorber, B, Giege, R, Sauter, C. | | Deposit date: | 2011-08-11 | | Release date: | 2011-11-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Exploiting Protein Engineering and Crystal Polymorphism for Successful X-Ray Structure Determination
Cryst.Growth Des., 11, 2011
|
|
5EUS
 
 | | Rat prestin STAS domain in complex with bromide | | Descriptor: | 1,2-ETHANEDIOL, BROMIDE ION, Prestin,Rat prestin STAS domain, ... | | Authors: | Lolli, G, Pasqualetto, E, Costanzi, E, Bonetto, G, Battistutta, R. | | Deposit date: | 2015-11-19 | | Release date: | 2015-12-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.833 Å) | | Cite: | The STAS domain of mammalian SLC26A5 prestin harbours an anion-binding site.
Biochem.J., 473, 2016
|
|
5EUW
 
 | | Rat prestin STAS domain in complex with nitrate | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, NITRATE ION, ... | | Authors: | Lolli, G, Pasqualetto, E, Costanzi, E, Bonetto, G, Battistutta, R. | | Deposit date: | 2015-11-19 | | Release date: | 2015-12-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | The STAS domain of mammalian SLC26A5 prestin harbours an anion-binding site.
Biochem.J., 473, 2016
|
|
5EUX
 
 | | Rat prestin STAS domain in complex with thiocyanate | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, Prestin,Prestin, ... | | Authors: | Lolli, G, Pasqualetto, E, Costanzi, E, Bonetto, G, Battistutta, R. | | Deposit date: | 2015-11-19 | | Release date: | 2015-12-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.038 Å) | | Cite: | The STAS domain of mammalian SLC26A5 prestin harbours an anion-binding site.
Biochem.J., 473, 2016
|
|
2ZQN
 
 | | Crystal structure of the earthworm R-type lectin C-half in complex with Lactose | | Descriptor: | 29-kDa galactose-binding lectin, IMIDAZOLE, PHOSPHATE ION, ... | | Authors: | Suzuki, R, Kuno, A, Hasegawa, T, Hirabayashi, J, Kasai, K, Momma, M, Fujimoto, Z. | | Deposit date: | 2008-08-13 | | Release date: | 2008-09-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Sugar-complex structures of the C-half domain of the galactose-binding lectin EW29 from the earthworm Lumbricus terrestris
Acta Crystallogr.,Sect.D, 65, 2009
|
|
1X9Y
 
 | | The prostaphopain B structure | | Descriptor: | cysteine proteinase | | Authors: | Filipek, R, Szczepanowski, R, Sabat, A, Potempa, J, Bochtler, M. | | Deposit date: | 2004-08-24 | | Release date: | 2004-11-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Prostaphopain B structure: a comparison of proregion-mediated and staphostatin-mediated protease inhibition.
Biochemistry, 43, 2004
|
|
2Y5C
 
 | | Structure of human ferredoxin 2 (Fdx2)in complex with 2Fe2S cluster | | Descriptor: | ADRENODOXIN-LIKE PROTEIN, MITOCHONDRIAL, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Webert, H, Hobler, A, Sheftel, A.D, Molik, S, Maestre-Reyna, M, Essen, L.-O, Vorniscescu, D, Keusgen, M, Hannemann, F, Bernhardt, R, Lill, R. | | Deposit date: | 2011-01-12 | | Release date: | 2012-02-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure and Functional Studies on Human Mitochondrial Ferredoxins
To be Published
|
|
