3LVC
 
 | |
3LVA
 
 | |
3PIB
 | |
3PJ7
 | |
3PJ5
 
 | |
3PJB
 
 | |
2BQP
 
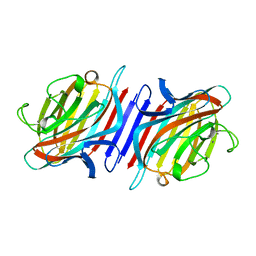 | | THE STRUCTURE OF THE PEA LECTIN-D-GLUCOPYRANOSE COMPLEX | | Descriptor: | CALCIUM ION, MANGANESE (II) ION, PROTEIN (PEA LECTIN), ... | | Authors: | Ruzeinikov, S.N, Mikhailova Yu, I, Tsygannik, I.N, Pangborn, W, Duax, W, Pletnev, V.Z. | | Deposit date: | 1998-12-08 | | Release date: | 1998-12-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Structure of Pea Lectin-D-Glucopyranose Complex at a 1.9 A Resolution
RUSS.J.BIOORGANIC CHEM., 23, 1997
|
|
7SFA
 
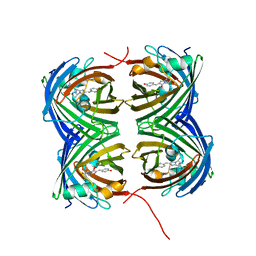 | |
7SF9
 
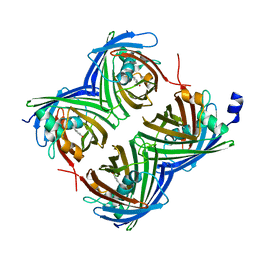 | |
2PXW
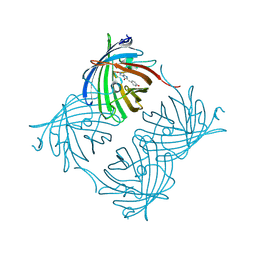 
 | | Crystal Structure of N66D Mutant of Green Fluorescent Protein from Zoanthus sp. at 2.4 A Resolution (Transition State) | | Descriptor: | GFP-like fluorescent chromoprotein FP506 | | Authors: | Pletnev, S.V, Pletneva, N.V, Tikhonova, T.V, Pletnev, V.Z. | | Deposit date: | 2007-05-14 | | Release date: | 2007-09-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Refined crystal structures of red and green fluorescent proteins from the button polyp Zoanthus.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2PXS
 
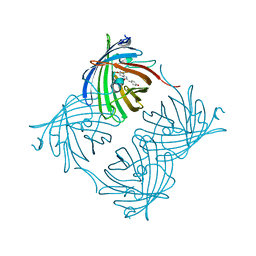 | | Crystal Structure of N66D Mutant of Green Fluorescent Protein from Zoanthus sp. at 2.2 A Resolution (Mature State) | | Descriptor: | GFP-like fluorescent chromoprotein FP506 | | Authors: | Pletnev, S.V, Pletneva, N.V, Tikhonova, T.V, Pletnev, V.Z. | | Deposit date: | 2007-05-14 | | Release date: | 2007-09-25 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Refined crystal structures of red and green fluorescent proteins from the button polyp Zoanthus.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2OJK
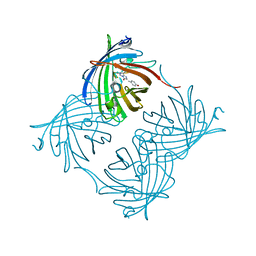 
 | | Crystal Structure of Green Fluorescent Protein from Zoanthus sp at 2.2 A Resolution | | Descriptor: | GFP-like fluorescent chromoprotein FP506 | | Authors: | Pletneva, N.V, Pletnev, S.V, Tikhonova, T.V, Pletnev, V.Z. | | Deposit date: | 2007-01-12 | | Release date: | 2007-09-25 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Refined crystal structures of red and green fluorescent proteins from the button polyp Zoanthus.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
4EDS
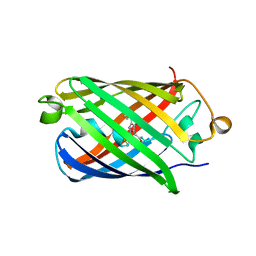 
 | |
4EDO
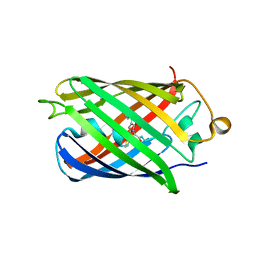 
 | |
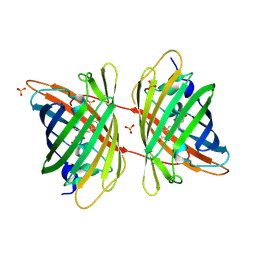
3GB3
 
 | |
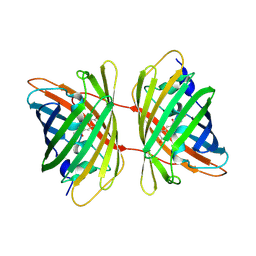
3GL4
 
 | |
1F90
 
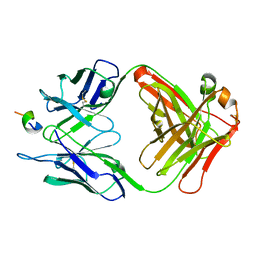 | | FAB FRAGMENT OF MONOCLONAL ANTIBODY (LNKB-2) AGAINST HUMAN INTERLEUKIN-2 IN COMPLEX WITH ANTIGENIC PEPTIDE | | Descriptor: | ANTIGENIC NONAPEPTIDE, FAB FRAGMENT OF MONOCLONAL ANTIBODY | | Authors: | Afonin, P.V, Fokin, A.V, Tsigannik, I.N, Mikhailova, I.Y, Onoprienko, L.V, Mikhaleva, I.I, Ivanov, V.T, Mareeva, T.Y, Nesmeyanov, V.A, Li, N, Duax, W.L, Pletnev, V.Z. | | Deposit date: | 2000-07-06 | | Release date: | 2001-07-11 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of an anti-interleukin-2 monoclonal antibody Fab complexed with an antigenic nonapeptide.
Protein Sci., 10, 2001
|
|
1BQP
 
 | | THE STRUCTURE OF THE PEA LECTIN-D-MANNOPYRANOSE COMPLEX | | Descriptor: | CALCIUM ION, MANGANESE (II) ION, PROTEIN (LECTIN), ... | | Authors: | Ruzeinikov, S.N, Mikhailova, I.Y, Tsygannik, I.N, Pangborn, W, Duax, W, Pletnev, V.Z. | | Deposit date: | 1998-08-17 | | Release date: | 1998-08-26 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Structure of the Pea Lectin-D-Mannopyranose Complex at a 2.1 A Resolution
RUSS.J.BIOORGANIC CHEM., 24, 1998
|
|
