7TCS
 
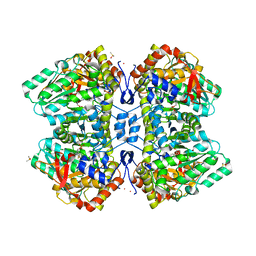 | | M379A mutant tyrosine phenol-lyase complexed with L-methionine | | Descriptor: | (4Z)-4-({[(1E)-1-carboxy-3-(methylsulfanyl)propylidene]azaniumyl}methylidene)-2-methyl-5-[(phosphonooxy)methyl]-1,4-dihydropyridin-3-olate, CESIUM ION, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2021-12-28 | | Release date: | 2022-06-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | M379A Mutant Tyrosine Phenol-lyase from Citrobacter freundii Has Altered Conformational Dynamics.
Chembiochem, 23, 2022
|
|
7TDL
 
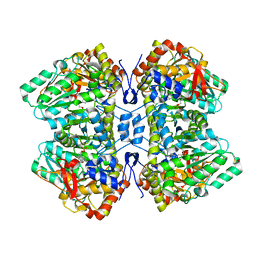 | | M379A mutant tyrosine phenol-lyase complexed with 3-bromo-DL-phenylalanine | | Descriptor: | (4Z)-4-({[(1E)-2-(3-bromophenyl)-1-carboxyethylidene]azaniumyl}methylidene)-2-methyl-5-[(phosphonooxy)methyl]-1,4-dihydropyridin-3-olate, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2022-01-01 | | Release date: | 2022-06-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | M379A Mutant Tyrosine Phenol-lyase from Citrobacter freundii Has Altered Conformational Dynamics.
Chembiochem, 23, 2022
|
|
5JVP
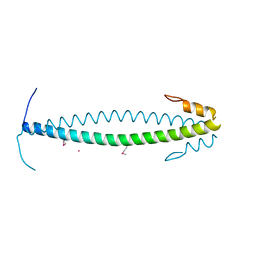 
 | | The neck-linker and alpha 7 helix of Homo sapiens CENP-E | | Descriptor: | CADMIUM ION, Chimera protein of Centromere-associated protein E and Microtubule-associated protein RP/EB family member 1 | | Authors: | Phillips, R.K, Rayment, I. | | Deposit date: | 2016-05-11 | | Release date: | 2016-08-03 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Family-specific Kinesin Structures Reveal Neck-linker Length Based on Initiation of the Coiled-coil.
J.Biol.Chem., 291, 2016
|
|
5JX1
 
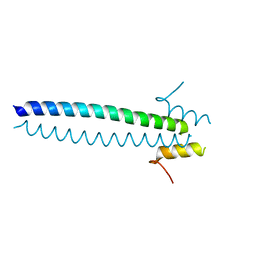 | |
9BNJ
 
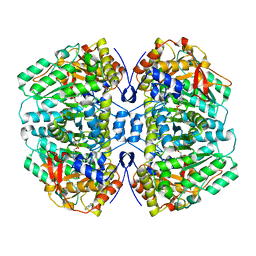 | |
8V2K
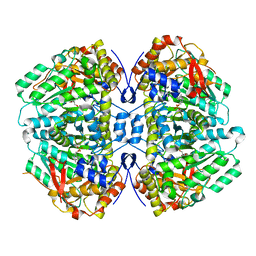 
 | | Proteus vulgaris tryptophan indole-lyase complexed with L-alanine | | Descriptor: | (2E)-2-{[(Z)-{3-HYDROXY-2-METHYL-5-[(PHOSPHONOOXY)METHYL]PYRIDIN-4(1H)-YLIDENE}METHYL]IMINO}PROPANOIC ACID, (E)-N-({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene)-L-alanine, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2023-11-22 | | Release date: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Proteus vulgaris tryptophan indole-lyase complexed with L-alanine
To Be Published
|
|
5W19
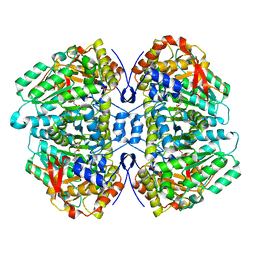 
 | | Tryptophan indole-lyase complex with oxindolyl-L-alanine | | Descriptor: | 1-carboxy-1-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]azaniumyl}-2-[(3R)-2-oxo-2,3-dihydro-1H-indol-3-yl]ethan-1-ide, POTASSIUM ION, Tryptophanase | | Authors: | Phillips, R.S, Wood, Z.A. | | Deposit date: | 2017-06-02 | | Release date: | 2018-06-06 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of Proteus vulgaris tryptophan indole-lyase complexed with oxindolyl-L-alanine: implications for the reaction mechanism.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
8V9P
 | | Proteus vulgaris tryptophan indole-lyase complexed with (3S)-dioxindolyl-L-alanine | | Descriptor: | (2~{E})-2-[(~{Z})-[2-methyl-3-oxidanyl-5-[[oxidanyl-bis(oxidanylidene)-$l^{6}-phosphanyl]oxymethyl]-1~{H}-pyridin-4-ylidene]methyl]imino-3-[(3~{S})-3-oxidanyl-2-oxidanylidene-1~{H}-indol-3-yl]propanoic acid, DIMETHYL SULFOXIDE, POTASSIUM ION, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2023-12-08 | | Release date: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Proteus vulgaris tryptophan indole-lyase complexed with L-ethionine
To Be Published
|
|
8V4A
 | | Proteus vulgaris tryptophan indole-lyase complexed with L-ethionine | | Descriptor: | (2E)-4-(ethylsulfanyl)-2-{[(Z)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4(1H)-ylidene}methyl]imino}butanoic acid, (E)-S-ethyl-N-({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene)-L-homocysteine, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2023-11-28 | | Release date: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Proteus vulgaris tryptophan indole-lyase complexed with L-ethionine
To Be Published
|
|
8V6P
 
 | | Proteus vulgaris tryptophan indole-lyase complexed with 7-aza-L-tryptophan | | Descriptor: | (2E)-2-{[(Z)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4(1H)-ylidene}methyl]imino}-3-(1H-pyrrolo[2,3-b]pyridin-3-yl)propanoic acid, (E)-N-({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene)-3-(1H-pyrrolo[2,3-b]pyridin-3-yl)-L-alanine, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2023-12-02 | | Release date: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Proteus vulgaris tryptophan indole-lyase complexed with L-alanine
To Be Published
|
|
6N2H
 
 | | Structure of D-ornithine/D-lysine decarboxylase from Salmonella typhimurium | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, D-ornithine/D-lysine decarboxylase, DIMETHYL SULFOXIDE | | Authors: | Phillips, R.S, Hoover, T.R. | | Deposit date: | 2018-11-13 | | Release date: | 2019-10-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Crystal Structure of d-Ornithine/d-Lysine Decarboxylase, a Stereoinverting Decarboxylase: Implications for Substrate Specificity and Stereospecificity of Fold III Decarboxylases.
Biochemistry, 58, 2019
|
|
5W1B
 | | Tryptophan indole-lyase | | Descriptor: | POTASSIUM ION, Tryptophanase | | Authors: | Phillips, R.S, Wood, Z.A. | | Deposit date: | 2017-06-02 | | Release date: | 2018-06-06 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of Proteus vulgaris tryptophan indole-lyase complexed with oxindolyl-L-alanine: implications for the reaction mechanism.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
8D4I
 
 | | Structure of Y430F D-ornithine/D-lysine decarboxylase complex with putrescine | | Descriptor: | 1,2-ETHANEDIOL, 1,4-DIAMINOBUTANE, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Phillips, R.S, Nguyen Hoang, K.N. | | Deposit date: | 2022-06-02 | | Release date: | 2022-11-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | The Y430F mutant of Salmonella d-ornithine/d-lysine decarboxylase has altered stereospecificity and a putrescine allosteric activation site.
Arch.Biochem.Biophys., 731, 2022
|
|
8D5R
 
 | | Structure of Y430F D-ornithine/D-lysine decarboxylase complex with D-ornithine | | Descriptor: | 1,2-ETHANEDIOL, 1,4-DIAMINOBUTANE, ACETATE ION, ... | | Authors: | Phillips, R.S, Nguyen Hoang, K.N. | | Deposit date: | 2022-06-06 | | Release date: | 2022-11-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | The Y430F mutant of Salmonella d-ornithine/d-lysine decarboxylase has altered stereospecificity and a putrescine allosteric activation site.
Arch.Biochem.Biophys., 731, 2022
|
|
8D88
 | |
8D2Y
 
 | | Y430F mutant of D-ornithine/D-lysine decarboxylase | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACETATE ION, ... | | Authors: | Phillips, R.S, Nguyen Hoang, K.N. | | Deposit date: | 2022-05-31 | | Release date: | 2022-11-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | The Y430F mutant of Salmonella d-ornithine/d-lysine decarboxylase has altered stereospecificity and a putrescine allosteric activation site.
Arch.Biochem.Biophys., 731, 2022
|
|
6MQQ
 
 | | Citrobacter freundii F448A mutant tyrosine phenol-lyase complexed with 4-hydroxypyridine and aminoacrylate from S-ethyl-L-cysteine | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, POTASSIUM ION, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2018-10-10 | | Release date: | 2019-10-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
6NV8
 
 | | Perdeuterated tyrosine phenol-lyase from Citrobacter freundii complexed with an aminoacrylate intermediate formed from S-ethyl-L-cysteine and 4-hydroxypyridine | | Descriptor: | 2-AMINO-ACRYLIC ACID, 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2019-02-04 | | Release date: | 2020-02-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
6XRH
 
 | | Salmonella typhimurium tryptophan synthase complexed with oxindolyl-L-alanine and D-glycerol-3-phosphate | | Descriptor: | (E)-N-({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene)-3-[(3S)-2-oxo-2,3-dihydro-1H-indol-3-yl]-L-alanine, 4-[(E)-({1-carboxy-2-[(3S)-2-oxo-2,3-dihydro-1H-indol-3-yl]ethan-1-id-1-yl}iminio)methyl]-2-methyl-5-[(phosphonooxy)methyl]pyridin-1-ium-3-olate, DIMETHYL SULFOXIDE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2020-07-12 | | Release date: | 2021-02-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Structural Basis of the Stereochemistry of Inhibition of Tryptophan Synthase by Tryptophan and Derivatives.
Biochemistry, 60, 2021
|
|
6XT0
 
 | |
7UX4
 
 | |
6MO3
 
 | | Citrobacter freundii tyrosine phenol-lyase complexed with 4-hydroxypyridine and aminoacrylate from L-serine | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, POTASSIUM ION, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2018-10-04 | | Release date: | 2019-10-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
6MLS
 
 | | Citrobacter freundii tyrosine phenol-lyase complexed with 4-hydroxypyridine and aminoacrylate from L-tyrosine | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, POTASSIUM ION, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2018-09-27 | | Release date: | 2019-10-02 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
6MPD
 
 | | Citrobacter freundii tyrosine phenol-lyase complexed with 4-hydroxypyridine and aminoacrylate from 3-F-L-tyrosine | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, 3-FLUOROTYROSINE, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2018-10-05 | | Release date: | 2019-10-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
6MME
 
 | | Citrobacter freundii tyrosine phenol-lyase complexed with 4-hydroxypyridine and aminoacrylate from S-ethyl-L-cysteine | | Descriptor: | 2-{[(E)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methylidene]amino}prop-2-enoic acid, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, POTASSIUM ION, ... | | Authors: | Phillips, R.S. | | Deposit date: | 2018-09-30 | | Release date: | 2019-10-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Pressure and Temperature Effects on the Formation of Aminoacrylate Intermediates of Tyrosine Phenol-lyase Demonstrate Reaction Dynamics
Acs Catalysis, 10, 2020
|
|
