1LOS
 
 | |
1LOQ
 
 | |
1LP6
 
 | |
1LOL
 
 | | Crystal structure of orotidine monophosphate decarboxylase complex with XMP | | Descriptor: | 1,3-BUTANEDIOL, XANTHOSINE-5'-MONOPHOSPHATE, orotidine 5'-monophosphate decarboxylase | | Authors: | Wu, N, Pai, E.F. | | Deposit date: | 2002-05-06 | | Release date: | 2002-08-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structures of inhibitor complexes reveal an alternate binding mode in orotidine-5'-monophosphate decarboxylase.
J.Biol.Chem., 277, 2002
|
|
1LOR
 
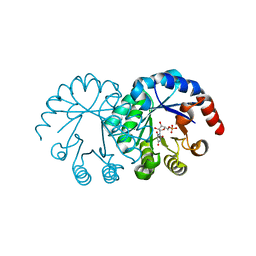 | | crystal structure of orotidine 5'-monophosphate complexed with BMP | | Descriptor: | 6-HYDROXYURIDINE-5'-PHOSPHATE, orotidine monophosphate decarboxylase | | Authors: | Wu, N, Pai, E.F. | | Deposit date: | 2002-05-06 | | Release date: | 2002-08-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures of inhibitor complexes reveal an alternate binding mode in orotidine-5'-monophosphate decarboxylase.
J.Biol.Chem., 277, 2002
|
|
1M8K
 
 | |
1LVW
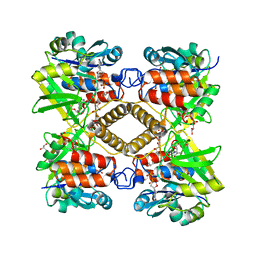 
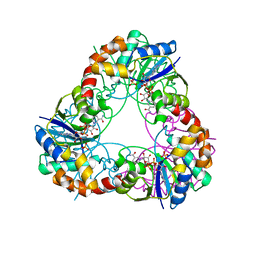
 | | Crystal structure of glucose-1-phosphate thymidylyltransferase, RmlA, complex with dTDP | | Descriptor: | CHLORIDE ION, GLYCEROL, SULFATE ION, ... | | Authors: | Dong, A, Christendat, D, Pai, E.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2002-05-29 | | Release date: | 2003-07-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of glucose-1-phosphate thymidylyltransferase, RmlA, complex with dTDP
To be Published
|
|
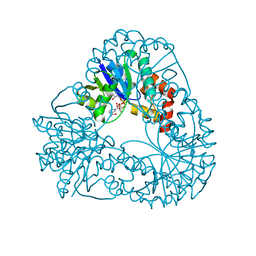
1M8F
 
 | |
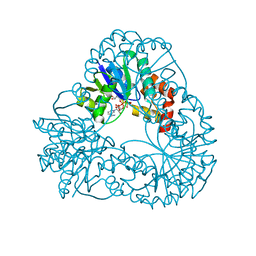
1M8J
 
 | |
1M8G
 
 | |
1N5X
 
 | | Xanthine Dehydrogenase from Bovine Milk with Inhibitor TEI-6720 Bound | | Descriptor: | 2-(3-CYANO-4-ISOBUTOXY-PHENYL)-4-METHYL-5-THIAZOLE-CARBOXYLIC ACID, DIOXOTHIOMOLYBDENUM(VI) ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Okamoto, K, Eger, B.T, Nishino, T, Kondo, S, Pai, E.F, Nishino, T. | | Deposit date: | 2002-11-07 | | Release date: | 2003-03-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | An Extremely Potent Inhibitor of Xanthine Oxidoreductase: Crystal Structure of the Enzyme-Inhibitor Complex and Mechanism of Inhibition
J.BIOL.CHEM., 278, 2003
|
|
3N2M
 
 | | Crystal structure of Plasmodium falciparum orotidine 5'-monophosphate decarboxylase complexed with 5-fluoro-6-amino-UMP | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 6-amino-5-fluorouridine 5'-(dihydrogen phosphate), ... | | Authors: | Liu, Y, Kotra, L.P, Pai, E.F. | | Deposit date: | 2010-05-18 | | Release date: | 2011-04-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Plasmodium falciparum orotidine 5'-monophosphate decarboxylase complexed with 5-fluoro-6-amino-UMP
To be Published
|
|
3N34
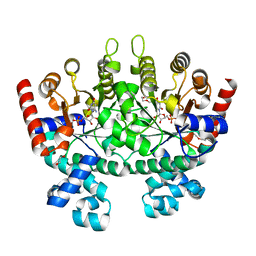 
 | | Crystal structure of Plasmodium falciparum orotidine 5'-monophosphate decarboxylase complexed with 5-fluoro-6-amino-UMP, produced from 5-fluoro-6-azido-UMP | | Descriptor: | 1,2-ETHANEDIOL, 6-amino-5-fluorouridine 5'-(dihydrogen phosphate), DI(HYDROXYETHYL)ETHER, ... | | Authors: | Liu, Y, Kotra, L.P, Pai, E.F. | | Deposit date: | 2010-05-19 | | Release date: | 2011-04-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal structure of Plasmodium falciparum orotidine 5'-monophosphate decarboxylase complexed with 5-fluoro-6-amino-UMP, produced from 5-fluoro-6-azido-UMP
To be Published
|
|
3MWA
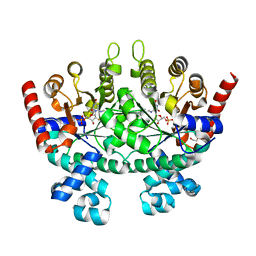 
 | |
3N3M
 
 | |
2OBA
 
 | | Pseudomonas aeruginosa 6-pyruvoyl tetrahydrobiopterin synthase | | Descriptor: | Probable 6-pyruvoyl tetrahydrobiopterin synthase, ZINC ION | | Authors: | McGrath, T.E, Kisselman, G, Battaile, K, Romanov, V, Wu-Brown, J, Guthrie, J, Virag, C, Mansoury, K, Edwards, A.M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2006-12-18 | | Release date: | 2007-01-30 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Pseudomonas aeruginosa 6-pyruvoyl tetrahydrobiopterin synthase
TO BE PUBLISHED
|
|
2O4D
 
 | | Crystal Structure of a hypothetical protein from Pseudomonas aeruginosa | | Descriptor: | Hypothetical protein PA0269 | | Authors: | McGrath, T.E, Battaile, K, Kisselman, G, Romanov, V, Wu-Brown, J, Virag, C, Ng, I, Kimber, M, Edwards, A.M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2006-12-04 | | Release date: | 2007-01-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of a hypothetical protein from Pseudomonas aeruginosa
To be Published
|
|
2O1B
 
 | | Structure of aminotransferase from Staphylococcus aureus | | Descriptor: | Aminotransferase, class I, PYRIDOXAL-5'-PHOSPHATE | | Authors: | McGrath, T.E, Dharamsi, A, Thambipillai, D, Edwards, A.M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2006-11-28 | | Release date: | 2006-12-12 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure of aminotransferase from Staphylococcus aureus
To be Published
|
|
3NA8
 
 | | Crystal Structure of a putative dihydrodipicolinate synthetase from Pseudomonas aeruginosa | | Descriptor: | D-MALATE, MAGNESIUM ION, putative dihydrodipicolinate synthetase | | Authors: | Qiu, W, Lam, R, Romanov, V, Jones, K, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2010-06-01 | | Release date: | 2011-06-01 | | Last modified: | 2012-02-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of a putative dihydrodipicolinate synthetase from Pseudomonas aeruginosa
To be Published
|
|
3NTS
 
 | |
3NTV
 
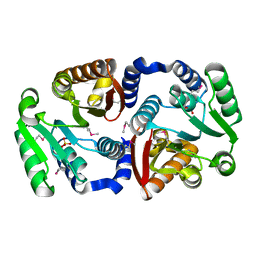 | | Crystal structure of a putative caffeoyl-CoA O-methyltransferase from Staphylococcus aureus | | Descriptor: | MW1564 protein, SULFATE ION | | Authors: | Qiu, W, Lam, R, Romanov, V, Jones, K, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2010-07-05 | | Release date: | 2011-07-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal structure of a putative caffeoyl-CoA O-methyltransferase from Staphylococcus aureus
TO BE PUBLISHED
|
|
3NUR
 
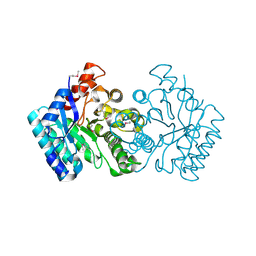 | | Crystal structure of a putative amidohydrolase from Staphylococcus aureus | | Descriptor: | Amidohydrolase, CALCIUM ION | | Authors: | Qiu, W, Lam, R, Romanov, V, Lam, K, Soloveychik, M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2010-07-07 | | Release date: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of a putative amidohydrolase from Staphylococcus aureus
To be Published
|
|
3O79
 
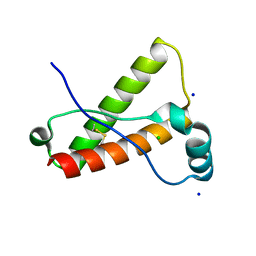 | | Crystal Structure of Wild-type Rabbit PrP 126-230 | | Descriptor: | CHLORIDE ION, GLYCEROL, Rabbit PrP, ... | | Authors: | Sweeting, B, Chakrabartty, A, Pai, E.F. | | Deposit date: | 2010-07-30 | | Release date: | 2010-11-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Prion disease susceptibility is affected by beta-structure folding propensity and local side-chain interactions in PrP.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2P8P
 
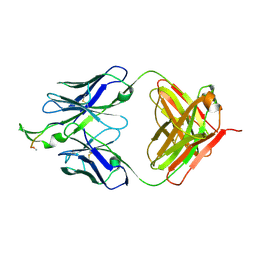 | | Crystal structure of the HIV-1 Cross Neutralizing Monoclonal Antibody 2F5 in complex with gp41 Peptide LELDKWASLW[N-Ac] | | Descriptor: | gp41 peptide, nmAb 2F5, heavy chain, ... | | Authors: | Bryson, S, Julien, J.-P, Pai, E.F. | | Deposit date: | 2007-03-22 | | Release date: | 2007-05-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural details of HIV-1 recognition by the broadly neutralizing monoclonal antibody 2F5: epitope conformation, antigen-recognition loop mobility, and anion-binding site.
J.Mol.Biol., 384, 2008
|
|
2P8L
 
 | | Crystal structure of the HIV-1 Cross Neutralizing Monoclonal Antibody 2F5 in complex with gp41 Peptide ELLELDKWASLWN | | Descriptor: | gp41 peptide, nmAb 2F5, heavy chain, ... | | Authors: | Julien, J.P, Bryson, S, Pai, E.F. | | Deposit date: | 2007-03-22 | | Release date: | 2007-05-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structural details of HIV-1 recognition by the broadly neutralizing monoclonal antibody 2F5: epitope conformation, antigen-recognition loop mobility, and anion-binding site.
J.Mol.Biol., 384, 2008
|
|
