1D1F
 | |
1OWT
 
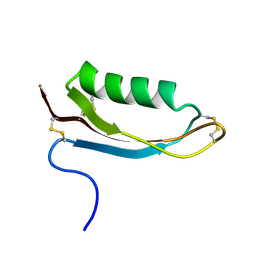 | | Structure of the Alzheimer's disease amyloid precursor protein copper binding domain | | Descriptor: | Amyloid beta A4 protein | | Authors: | Barnham, K.J, McKinstry, W.J, Multhaup, G, Galatis, D, Morton, C.J, Curtain, C.C, Williamson, N.A, White, A.R, Hinds, M.G, Norton, R.S, Beyreuther, K, Masters, C.L, Parker, M.W, Cappai, R. | | Deposit date: | 2003-03-30 | | Release date: | 2003-05-13 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure of the Alzheimer's Disease Amyloid Precursor Protein Copper Binding Domain. A REGULATOR OF NEURONAL COPPER HOMEOSTASIS.
J.Biol.Chem., 278, 2003
|
|
1PSM
 
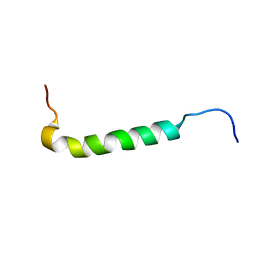 | |
1DG0
 
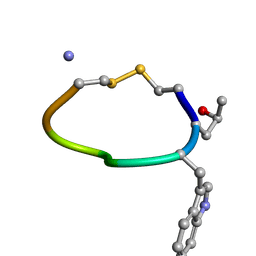 | | NMR STRUCTURE OF DES[GLY1]-CONTRYPHAN-R CYCLIC PEPTIDE (MAJOR FORM) | | Descriptor: | DES[GLY1]-CONTRYPHAN-R | | Authors: | Pallaghy, P.K, He, W, Jimenez, E.C, Olivera, B.M, Norton, R.S. | | Deposit date: | 1999-11-22 | | Release date: | 2003-09-09 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structures of the contryphan family of cyclic peptides. Role of electrostatic interactions in cis-trans isomerism
Biochemistry, 39, 2000
|
|
4Y1S
 | | Structural basis for Ca2+-mediated interaction of the perforin C2 domain with lipid membranes | | Descriptor: | CALCIUM ION, Perforin-1 | | Authors: | Conroy, P.J, Yagi, H, Whisstock, J.C, Norton, R.S. | | Deposit date: | 2015-02-09 | | Release date: | 2015-09-02 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.611 Å) | | Cite: | Structural Basis for Ca2+-mediated Interaction of the Perforin C2 Domain with Lipid Membranes.
J.Biol.Chem., 290, 2015
|
|
4Y1T
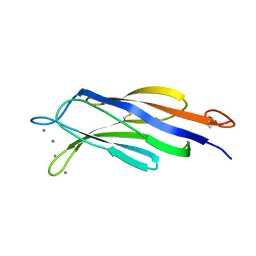 
 | | Structural basis for Ca2+-mediated interaction of the perforin C2 domain with lipid membranes | | Descriptor: | CALCIUM ION, Perforin-1 | | Authors: | Conroy, P.J, Yagi, H, Whisstock, J.C, Norton, R.S. | | Deposit date: | 2015-02-09 | | Release date: | 2015-09-02 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.666 Å) | | Cite: | Structural Basis for Ca2+-mediated Interaction of the Perforin C2 Domain with Lipid Membranes.
J.Biol.Chem., 290, 2015
|
|
1BH1
 
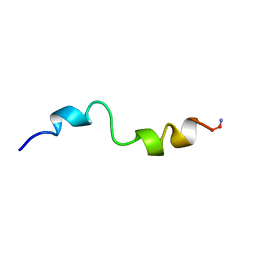 | | STRUCTURAL STUDIES OF D-PRO MELITTIN, NMR, 20 STRUCTURES | | Descriptor: | MELITTIN | | Authors: | Barnham, K.J, Hewish, D, Werkmeister, J, Curtain, C, Kirkpatrick, A, Bartone, N, Liu, S.T, Norton, R, Rivett, D. | | Deposit date: | 1998-06-11 | | Release date: | 1999-01-06 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Structure and activity of D-Pro14 melittin.
J.Protein Chem., 21, 2002
|
|
1RON
 
 | |
1DFZ
 
 | | NMR STRUCTURE OF CONTRYPHAN-SM CYCLIC PEPTIDE (MINOR FORM-TRANS) | | Descriptor: | CONTRYPHAN-SM | | Authors: | Pallaghy, P.K, He, W, Jimenez, E.C, Olivera, B.M, Norton, R.S. | | Deposit date: | 1999-11-22 | | Release date: | 2002-05-01 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structures of the contryphan family of cyclic peptides. Role of electrostatic
interactions in cis-trans isomerism.
Biochemistry, 39, 2000
|
|
6YM1
 
 | | Mycobacterium tuberculosis FtsZ in complex with GDP | | Descriptor: | Cell division protein FtsZ, GUANOSINE-5'-DIPHOSPHATE | | Authors: | Alnami, A.T, Norton, R.S, Pena, H.P, Haider, M, kozielski, F. | | Deposit date: | 2020-04-07 | | Release date: | 2021-04-14 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Conformational Flexibility of A Highly Conserved Helix Controls Cryptic Pocket Formation in FtsZ.
J.Mol.Biol., 433, 2021
|
|
6YM9
 
 | | Mycobacterium tuberculosis FtsZ in complex with GTP-gamma-S | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Cell division protein FtsZ | | Authors: | Alnami, A.T, Norton, R.S, Pena, H.P, Haider, M, kozielski, F. | | Deposit date: | 2020-04-08 | | Release date: | 2021-04-21 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Conformational Flexibility of A Highly Conserved Helix Controls Cryptic Pocket Formation in FtsZ.
J.Mol.Biol., 433, 2021
|
|
6BUC
 
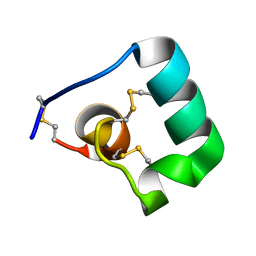 | |
1DFY
 
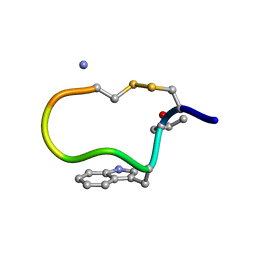 | | NMR STRUCTURE OF CONTRYPHAN-SM CYCLIC PEPTIDE (MAJOR FORM-CIS) | | Descriptor: | CONTRYPHAN-SM | | Authors: | Pallaghy, P.K, He, W, Jimenez, E.C, Olivera, B.M, Norton, R.S. | | Deposit date: | 1999-11-22 | | Release date: | 2002-05-01 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structures of the contryphan family of cyclic peptides. Role of electrostatic interactions in cis-trans isomerism.
Biochemistry, 39, 2000
|
|
1D7T
 
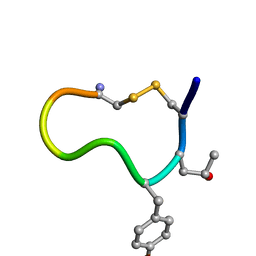 | |
1ROO
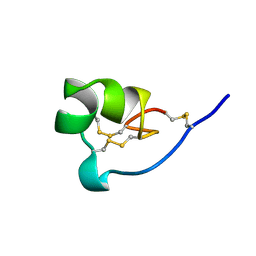 
 | | NMR SOLUTION STRUCTURE OF SHK TOXIN, NMR, 20 STRUCTURES | | Descriptor: | SHK TOXIN | | Authors: | Tudor, J.E, Pallaghy, P.K, Pennington, M.W, Norton, R.S. | | Deposit date: | 1996-01-11 | | Release date: | 1997-01-27 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of ShK toxin, a novel potassium channel inhibitor from a sea anemone.
Nat.Struct.Biol., 3, 1996
|
|
1A3P
 
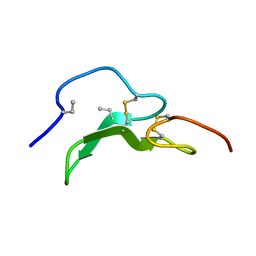 | | ROLE OF THE 6-20 DISULFIDE BRIDGE IN THE STRUCTURE AND ACTIVITY OF EPIDERMAL GROWTH FACTOR, NMR, 20 STRUCTURES | | Descriptor: | EPIDERMAL GROWTH FACTOR | | Authors: | Barnham, K, Torres, A, Alewood, D, Alewood, P, Domagala, T, Nice, E, Norton, R. | | Deposit date: | 1998-01-22 | | Release date: | 1998-07-29 | | Last modified: | 2018-03-14 | | Method: | SOLUTION NMR | | Cite: | Role of the 6-20 disulfide bridge in the structure and activity of epidermal growth factor.
Protein Sci., 7, 1998
|
|
1DSJ
 
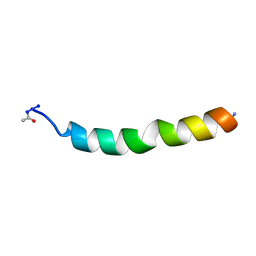 | | NMR SOLUTION STRUCTURE OF VPR50_75, 20 STRUCTURES | | Descriptor: | VPR PROTEIN | | Authors: | Yao, S, Torres, A.M, Azad, A.A, Macreadie, I.G, Norton, R.S. | | Deposit date: | 1997-10-23 | | Release date: | 1998-07-01 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Helical structure of polypeptides from the C-terminal half of HIV-1 VPR.
Protein Pept.Lett., 5, 1998
|
|
1DSK
 
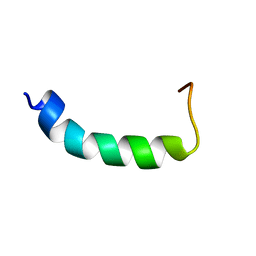 | | NMR SOLUTION STRUCTURE OF VPR59_86, 20 STRUCTURES | | Descriptor: | VPR PROTEIN | | Authors: | Yao, S, Torres, A.M, Azad, A.A, Macreadie, I.G, Norton, R.S. | | Deposit date: | 1997-10-23 | | Release date: | 1998-07-01 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | Solution structure of peptides from HIV-1 Vpr protein that cause membrane permeabilization and growth arrest.
J. Pept. Sci., 4, 1998
|
|
1BEI
 
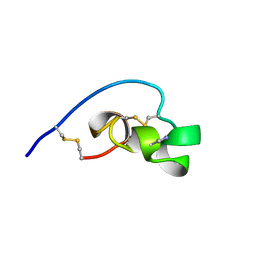 | | Shk-dnp22: A Potent Kv1.3-specific immunosuppressive polypeptide, NMR, 20 structures | | Descriptor: | POTASSIUM CHANNEL TOXIN SHK | | Authors: | Kalman, K, Pennington, M.W, Lanigan, M.D, Nguyen, A, Rauer, H, Mahnir, V, Gutman, G.A, Paschetto, K, Kem, W.R, Grissmer, S, Christian, E.P, Cahalan, M.D, Norton, R.S, Chandy, K.G. | | Deposit date: | 1998-05-14 | | Release date: | 1998-12-02 | | Last modified: | 2022-12-21 | | Method: | SOLUTION NMR | | Cite: | ShK-Dap22, a potent Kv1.3-specific immunosuppressive polypeptide.
J.Biol.Chem., 273, 1998
|
|
1C2U
 
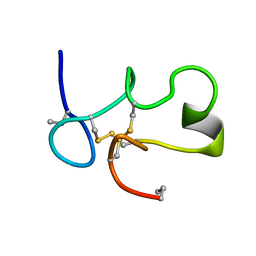 | | SOLUTION STRUCTURE OF [ABU3,35]SHK12-28,17-32 | | Descriptor: | SYNTHETIC PEPTIDE ANALOGUE OF SHK TOXIN | | Authors: | Pennington, M.W, Lanigan, M.D, Kalman, K, Manhir, V.M, Rauer, H, McVaugh, C.T, Behm, D, Donaldson, D, Chandy, K.G, Kem, W.R, Norton, R.S. | | Deposit date: | 1999-07-27 | | Release date: | 1999-11-10 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Role of disulfide bonds in the structure and potassium channel blocking activity of ShK toxin.
Biochemistry, 38, 1999
|
|
1QDP
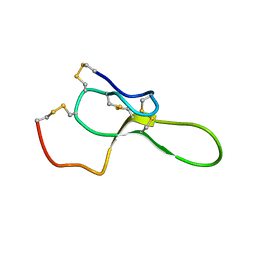 
 | | SOLUTION STRUCTURE OF ROBUSTOXIN, THE LETHAL NEUROTOXIN FROM THE FUNNEL WEB SPIDER ATRAX ROBUSTUS, NMR, 20 STRUCTURES | | Descriptor: | ROBUSTOXIN | | Authors: | Pallaghy, P.K, Alewood, D, Alewood, P.F, Norton, R.S. | | Deposit date: | 1997-10-09 | | Release date: | 1998-01-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution structure of robustoxin, the lethal neurotoxin from the funnel-web spider Atrax robustus.
FEBS Lett., 419, 1997
|
|
1S6W
 
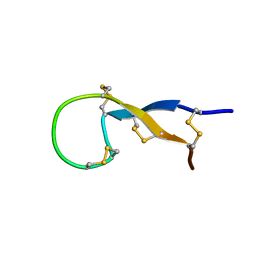 | | Solution Structure of hybrid white striped bass hepcidin | | Descriptor: | Hepcidin | | Authors: | Babon, J.J, Singh, S, Pennington, M.W, Norton, R.S, Westerman, M.E. | | Deposit date: | 2004-01-28 | | Release date: | 2004-12-14 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Bass hepcidin synthesis, solution structure, antimicrobial activities and synergism, and in vivo hepatic response to bacterial infections.
J.Biol.Chem., 280, 2005
|
|
2JRY
 
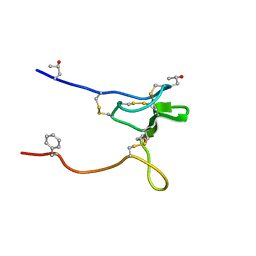 | | Structure and Sodium Channel Activity of an Excitatory I1-Superfamily Conotoxin | | Descriptor: | I-superfamily conotoxin r11a | | Authors: | Buczek, O, Wei, D, Babon, J, Yang, X, Fiedler, B, Chen, P, Yoshikami, D, Olivera, B, Bulaj, G, Norton, R. | | Deposit date: | 2007-06-29 | | Release date: | 2007-10-23 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure and sodium channel activity of an excitatory I1-superfamily conotoxin.
Biochemistry, 46, 2007
|
|
1Q2J
 
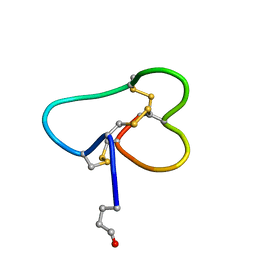 | | Structural basis for tetrodotoxin-resistant sodium channel binding by mu-conotoxin SmIIIA | | Descriptor: | Mu-conotoxin SmIIIA | | Authors: | Keizer, D.W, West, P.J, Lee, E.F, Olivera, B.M, Bulaj, G, Yoshikami, D, Norton, R.S. | | Deposit date: | 2003-07-24 | | Release date: | 2004-02-24 | | Last modified: | 2020-06-24 | | Method: | SOLUTION NMR | | Cite: | Structural basis for tetrodotoxin-resistant sodium channel binding by mu-conotoxin SmIIIA.
J.Biol.Chem., 278, 2003
|
|
1QFA
 
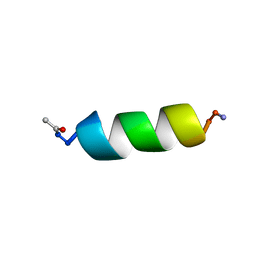 | | STRUCTURE OF A NEUROPEPTIDE Y Y2 AGONIST | | Descriptor: | PROTEIN (NEUROPEPTIDE Y) | | Authors: | Barnham, K.J, Catalfamo, F, Pallaghy, P.K, Howlett, G.J, Norton, R.S. | | Deposit date: | 1999-04-08 | | Release date: | 2000-04-08 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Helical structure and self-association in a 13 residue neuropeptide Y Y2 receptor agonist: relationship to biological activity.
Biochim.Biophys.Acta, 1435, 1999
|
|
