5I21
 
 | |
8P9M
 
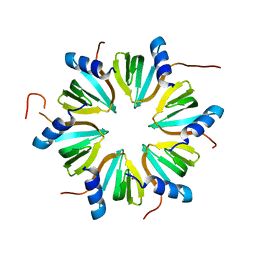 | | Hexameric Hfq from Chromobacterium haemolyticum | | Descriptor: | GLYCEROL, RNA-binding protein Hfq | | Authors: | Nikulin, A.D, Lekontseva, N.V. | | Deposit date: | 2023-06-06 | | Release date: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Structure of the Hfq Protein from Chromobacterium haemolyticum Revealed a New Variant of Regulation of RNA Binding with the Protein
Crystallography Reports, 68, 2023
|
|
8P91
 
 | |
1KUQ
 
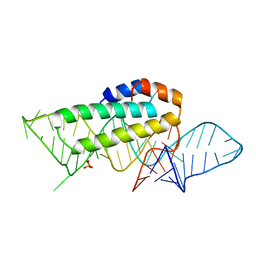 | | CRYSTAL STRUCTURE OF T3C MUTANT S15 RIBOSOMAL PROTEIN IN COMPLEX WITH 16S RRNA | | Descriptor: | 16S RIBOSOMAL RNA FRAGMENT, 30S RIBOSOMAL PROTEIN S15, SULFATE ION | | Authors: | Nikulin, A.D, Tishchenko, S, Revtovich, S, Ehresmann, B, Ehresmann, C, Dumas, P, Garber, M, Nikonov, S, Nevskaya, N. | | Deposit date: | 2002-01-22 | | Release date: | 2003-06-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Role of N-terminal helix in interaction of ribosomal protein S15 with 16S rRNA.
Biochemistry Mosc., 69, 2004
|
|
6TFL
 
 | | Lsm protein (SmAP) from Halobacterium salinarum | | Descriptor: | CALCIUM ION, GLYCEROL, RNA-binding protein Lsm, ... | | Authors: | Nikulin, A.D, Fando, M.S, Lekontseva, N.V. | | Deposit date: | 2019-11-14 | | Release date: | 2020-11-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.397 Å) | | Cite: | Structure and RNA-Binding Properties of Lsm Protein from Halobacterium salinarum.
Biochemistry Mosc., 86, 2021
|
|
3QUI
 
 | | Crystal structure of Pseudomonas aeruginosa Hfq in complex with ADPNP | | Descriptor: | ADENINE, ADENOSINE-5'-DIPHOSPHATE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nikulin, A.D, Murina, V.N, Gabdulkhakov, A.G. | | Deposit date: | 2011-02-24 | | Release date: | 2012-02-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.933 Å) | | Cite: | Hfq binds ribonucleotides in three different RNA-binding sites.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
5MKN
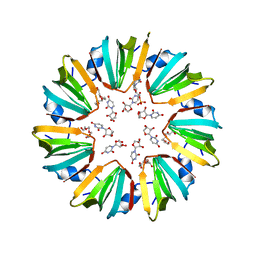 
 | |
6XYJ
 
 | |
2RFV
 
 | | High resolution structure of L-methionine gamma-lyase from Citrobacter freundii | | Descriptor: | CHLORIDE ION, Methionine gamma-lyase | | Authors: | Nikulin, A.D, Revtovich, S.V, Morozova, E.A, Nevskaya, N.A, Nikonov, S.V, Garber, M.B, Demidkina, T.V. | | Deposit date: | 2007-10-02 | | Release date: | 2008-08-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.355 Å) | | Cite: | High-resolution structure of methionine gamma-lyase from Citrobacter freundii.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
4X9D
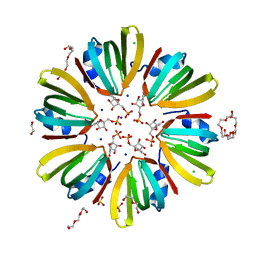 
 | | High-resolution structure of Hfq from Methanococcus jannaschii in complex with UMP | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nikulin, A.D, Tishchenko, S.V, Nikonova, E.Y, Murina, V.N, Mihailina, A.O, Lekontseva, N.V. | | Deposit date: | 2014-12-11 | | Release date: | 2015-12-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Characterization of RNA-binding properties of the archaeal Hfq-like protein from Methanococcus jannaschii.
J. Biomol. Struct. Dyn., 35, 2017
|
|
4X9C
 
 | | 1.4A crystal structure of Hfq from Methanococcus jannaschii | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nikulin, A.D, Tishchenko, S.V, Nikonova, S.V, Murina, V.N, Mihailina, A.O, Lekontseva, N.V. | | Deposit date: | 2014-12-11 | | Release date: | 2014-12-24 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Characterization of RNA-binding properties of the archaeal Hfq-like protein from Methanococcus jannaschii.
J. Biomol. Struct. Dyn., 35, 2017
|
|
1U1T
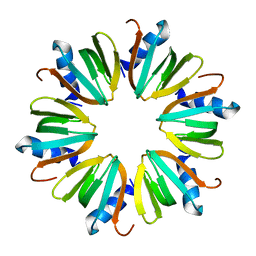 
 | | Hfq protein from Pseudomonas aeruginosa. High-salt crystals | | Descriptor: | Hfq protein | | Authors: | Nikulin, A.D, Stolboushkina, E.A, Perederina, A.A, Vassilieva, I.M, Blaesi, U, Moll, I, Kachalova, G, Yokoyama, S, Vassylyev, D, Garber, M, Nikonov, S.V, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-07-16 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of Pseudomonas aeruginosa Hfq protein.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1U1S
 
 | | Hfq protein from Pseudomonas aeruginosa. Low-salt crystals | | Descriptor: | Hfq protein | | Authors: | Nikulin, A.D, Stolboushkina, E.A, Perederina, I, Vassilieva, I.M, Blaesi, U, Moll, I, Kachalova, G, Vassylyev, D, Yokoyama, S, Garber, M, Nikonov, S.V, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-07-16 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of Pseudomonas aeruginosa Hfq protein.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
5MKI
 
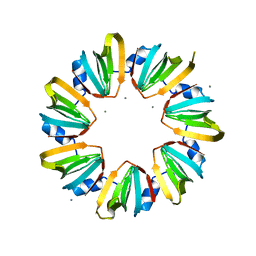 | |
5MKL
 
 | |
4J6W
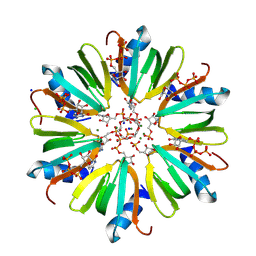 
 | | Crystal structure of HFQ from Pseudomonas aeruginosa in complex with CTP | | Descriptor: | CHLORIDE ION, CYTIDINE-5'-DIPHOSPHATE, CYTIDINE-5'-MONOPHOSPHATE, ... | | Authors: | Nikulin, A.D, Murina, V, Lekontseva, N. | | Deposit date: | 2013-02-12 | | Release date: | 2013-07-31 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Hfq binds ribonucleotides in three different RNA-binding sites.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4J6X
 
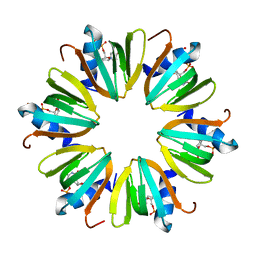 | |
4J6Y
 
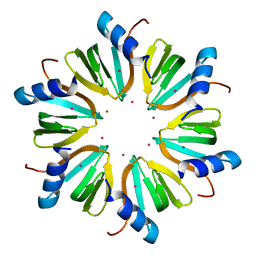 | |
5DY9
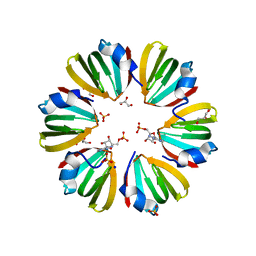 
 | | Y68T Hfq from Methanococcus jannaschii in complex with AMP | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ADENOSINE MONOPHOSPHATE, CHLORIDE ION, ... | | Authors: | Nikulin, A.D, Mikhailina, A.O, Lekontseva, N.V, Balobanov, V.A, Nikonova, E.Y, Tishchenko, S.V. | | Deposit date: | 2015-09-24 | | Release date: | 2016-09-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Characterization of RNA-binding properties of the archaeal Hfq-like protein from Methanococcus jannaschii.
J. Biomol. Struct. Dyn., 35, 2017
|
|
3MKJ
 
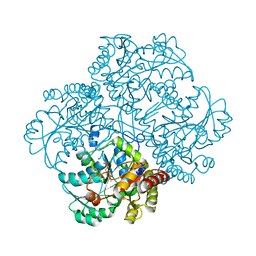 | | Methionine gamma-lyase from Citrobacter freundii with pyridoximine-5'-phosphate | | Descriptor: | Methionine gamma-lyase, [5-hydroxy-4-(iminomethyl)-6-methyl-pyridin-3-yl]methyl dihydrogen phosphate | | Authors: | Revtovich, S.V, Nikulin, A.D, Morozova, E.A, Demidkina, T.V. | | Deposit date: | 2010-04-15 | | Release date: | 2010-08-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Exploring methionine gamma-lyase structure-function relationship via microspectrophotometry and X-ray crystallography
Biochim.Biophys.Acta, 1814, 2011
|
|
3CW2
 
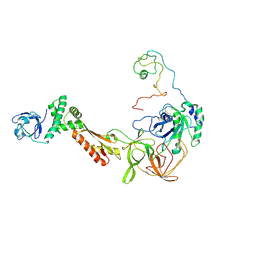 | | Crystal structure of the intact archaeal translation initiation factor 2 from Sulfolobus solfataricus . | | Descriptor: | Translation initiation factor 2 subunit alpha, Translation initiation factor 2 subunit beta, Translation initiation factor 2 subunit gamma | | Authors: | Stolboushkina, E.A, Nikonov, S.V, Nikulin, A.D, Blaesi, U, Manstein, D.J, Fedorov, R.V, Garber, M.B, Nikonov, O.S. | | Deposit date: | 2008-04-21 | | Release date: | 2009-01-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the intact archaeal translation initiation factor 2 demonstrates very high conformational flexibility in the alpha- and beta-subunits.
J.Mol.Biol., 382, 2008
|
|
1Y4I
 
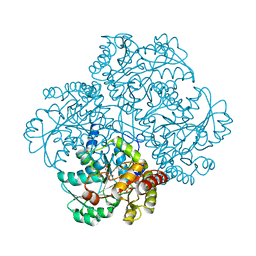 | | Crystal structure of Citrobacter Freundii L-methionine-lyase | | Descriptor: | SULFATE ION, methionine gamma-lyase | | Authors: | Revtovich, S.V, Mamaeva, D.V, Morozova, E.A, Nikulin, A.D, Nikonov, S.V, Garber, M.B, Demidkina, T.V. | | Deposit date: | 2004-12-01 | | Release date: | 2005-06-14 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of Citrobacter freundii L-methionine gamma-lyase.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
5K30
 
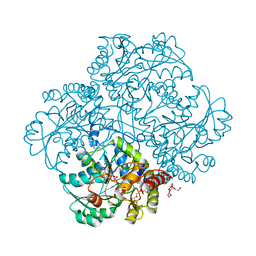 | | Crystal structure of methionine gamma-lyase from Citrobacter freundii modified by S-Ethyl-L-cysteine sulfoxide | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, DI(HYDROXYETHYL)ETHER, Methionine gamma-lyase, ... | | Authors: | Revtovich, S.V, Nikulin, A.D, Morozova, E.A, Demidkina, T.V. | | Deposit date: | 2016-05-19 | | Release date: | 2017-07-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Sulfoxides of sulfur-containing amino acids are suicide substrates of Citrobacter freundii methionine gamma-lyase. Structural bases of the enzyme inactivation.
Biochimie, 168, 2020
|
|
6EGR
 
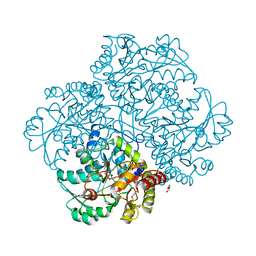 | | Crystal structure of Citrobacter freundii methionine gamma-lyase with V358Y replacement | | Descriptor: | DI(HYDROXYETHYL)ETHER, Methionine gamma-lyase, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Revtovich, S.V, Demitri, N, Raboni, S, Nikulin, A.D, Morozova, E.A, Demidkina, T.V, Storici, P, Mozzarelli, A. | | Deposit date: | 2017-09-12 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Engineering methionine gamma-lyase from Citrobacter freundii for anticancer activity.
Biochim Biophys Acta Proteins Proteom, 1866, 2018
|
|
5M3Z
 
 | | Crystal structure of Citrobacter freundii methionine gamma-lyase with C115H replacement in the complex with L-norleucine | | Descriptor: | 2-[O-PHOSPHONOPYRIDOXYL]-AMINO-HEXANOIC ACID, Methionine gamma-lyase, NORLEUCINE, ... | | Authors: | Revtovich, S.V, Nikulin, A.D, Anufrieva, N.V, Morozova, E.A, Demidkina, T.V. | | Deposit date: | 2016-10-17 | | Release date: | 2017-06-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal structure of mutant form Cys115His of Citrobacter freundii methionine gamma-lyase complexed with l-norleucine.
Biochim. Biophys. Acta, 1865, 2017
|
|
