5WQH
 
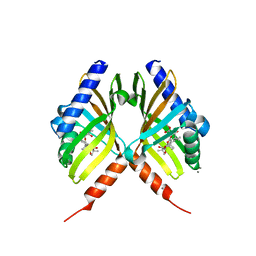 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with substrate analog and endo-terretonin D | | Descriptor: | CALCIUM ION, Isomerase trt14, methyl (2S,4aR,4bS,5S,6aS,10aS,10bS,12aS)-2,4b,7,7,10a,12,12a-heptamethyl-5-oxidanyl-1,4,6,8-tetrakis(oxidanylidene)-4a,5,6a,9,10,10b-hexahydronaphtho[1,2-h]isochromene-2-carboxylate, ... | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
7DRE
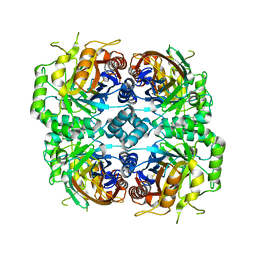 
 | | Cryo-EM structure of DfgA-B at 2.54 angstrom resolution | | Descriptor: | DfgB, Sugar phosphate isomerase/epimerase | | Authors: | Mori, T, Moriya, T, Adachi, N, Senda, T, Abe, I. | | Deposit date: | 2020-12-28 | | Release date: | 2021-12-08 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (2.54 Å) | | Cite: | C-Glycoside metabolism in the gut and in nature: Identification, characterization, structural analyses and distribution of C-C bond-cleaving enzymes.
Nat Commun, 12, 2021
|
|
7DRD
 
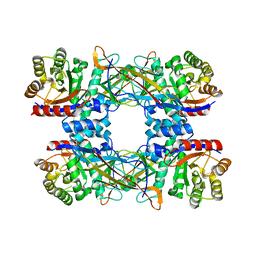 | | Cryo-EM structure of DgpB-C at 2.85 angstrom resolution | | Descriptor: | AP_endonuc_2 domain-containing protein, DgpB | | Authors: | Mori, T, Moriya, T, Adachi, N, Senda, T, Abe, I. | | Deposit date: | 2020-12-28 | | Release date: | 2021-12-08 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (2.85 Å) | | Cite: | C-Glycoside metabolism in the gut and in nature: Identification, characterization, structural analyses and distribution of C-C bond-cleaving enzymes.
Nat Commun, 12, 2021
|
|
8IQR
 
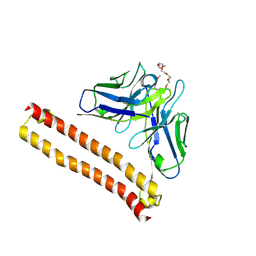 | | Crystal structure of Anti-PEG antibody M9 Fv-clasp fragment with PEG (co-crystallization with PEG550DME) | | Descriptor: | 2,5,8,11,14,17,20,23,26-nonaoxaoctacosane, M9 VH-SARAH, M9 VL-SARAH | | Authors: | Mori, T, Teramoto, T, Liu, Y, Mori, T, Kakuta, Y. | | Deposit date: | 2023-03-17 | | Release date: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Comparation of structures and binding properties between two anti-polyethylene glycol antibodies induced via T cell-independent and T cell-dependent pathway
To Be Published
|
|
8IQP
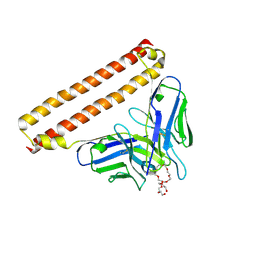 
 | | Crystal structure of Anti-PEG antibody M9 Fv-clasp fragment with PEG (co-crystallization with PEG3350) | | Descriptor: | DODECAETHYLENE GLYCOL, M9 VH-SARAH, M9 VL-SARAH | | Authors: | Mori, T, Teramoto, T, Liu, Y, Mori, T, Kakuta, Y. | | Deposit date: | 2023-03-17 | | Release date: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Comparation of structures and binding properties between two anti-polyethylene glycol antibodies induced via T cell-independent and T cell-dependent pathway
To Be Published
|
|
8IQQ
 
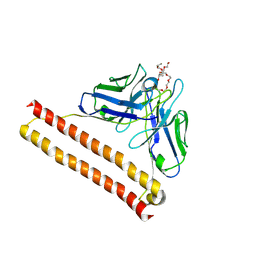 | | Crystal structure of Anti-PEG antibody M9 Fv-clasp fragment with PEG (co-crystallization with PEG2000MME) | | Descriptor: | DODECAETHYLENE GLYCOL, M9 VH-SARAH, M9 VL-SARAH | | Authors: | Mori, T, Teramoto, T, Liu, Y, Mori, T, Kakuta, Y. | | Deposit date: | 2023-03-17 | | Release date: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Comparation of structures and binding properties between two anti-polyethylene glycol antibodies induced via T cell-independent and T cell-dependent pathway
To Be Published
|
|
8IQS
 
 | | Crystal structure of Anti-PEG antibody M11 Fv-clasp fragment with PEG (co-crystallization with PEG3350) | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, M11 VH-SARAH, M11 VL-SARAH, ... | | Authors: | Mori, T, Teramoto, T, Liu, Y, Mori, T, Kakuta, Y. | | Deposit date: | 2023-03-17 | | Release date: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Comparation of structures and binding properties between two anti-polyethylene glycol antibodies induced via T cell-independent and T cell-dependent pathway
To Be Published
|
|
7YN3
 
 | | Crystal structure of CcbD complex with CcbZ carrier protein domain | | Descriptor: | 1,1'-ethane-1,2-diyldipyrrolidine-2,5-dione, 4'-PHOSPHOPANTETHEINE, CcbD, ... | | Authors: | Mori, T, Lyu, S, Abe, I. | | Deposit date: | 2022-07-29 | | Release date: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of CcbD complex with carrier protein
To Be Published
|
|
7YN1
 
 | |
7YN2
 
 | | Crystal structure of CcbD with methylthiolincosamide | | Descriptor: | (2R,3R,4S,5R,6R)-2-[(1R,2R)-1-azanyl-2-oxidanyl-propyl]-6-methylsulfanyl-oxane-3,4,5-triol, CcbD | | Authors: | Mori, T, Lyu, S, Abe, I. | | Deposit date: | 2022-07-29 | | Release date: | 2023-08-02 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of CcbD with methylthiolincosamide
To Be Published
|
|
8K3H
 
 | |
8K3I
 
 | |
5GK2
 
 | |
5GK0
 
 | |
5GK1
 
 | | Crystal structure of the ketosynthase StlD complexed with substrate | | Descriptor: | 3-OXO-5-METHYLHEXANOIC ACID, Ketosynthase StlD | | Authors: | Mori, T, Saito, Y, Morita, H, Abe, I. | | Deposit date: | 2016-07-03 | | Release date: | 2017-07-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural Insight into the Enzymatic Formation of Bacterial Stilbene.
Cell Chem Biol, 23, 2016
|
|
7BVR
 
 | | DgpB-DgpC complex apo | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, AP_endonuc_2 domain-containing protein, DgpB, ... | | Authors: | Mori, T, He, H, Abe, I. | | Deposit date: | 2020-04-11 | | Release date: | 2021-04-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | C-Glycoside metabolism in the gut and in nature: Identification, characterization, structural analyses and distribution of C-C bond-cleaving enzymes.
Nat Commun, 12, 2021
|
|
7BVS
 
 | | DfgA-DfgB complex apo | | Descriptor: | DfgB, GLYCEROL, MANGANESE (II) ION, ... | | Authors: | Mori, T, He, H, Abe, I. | | Deposit date: | 2020-04-11 | | Release date: | 2021-04-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | C-Glycoside metabolism in the gut and in nature: Identification, characterization, structural analyses and distribution of C-C bond-cleaving enzymes.
Nat Commun, 12, 2021
|
|
6TF8
 
 | |
4YZJ
 
 | |
8Y9P
 
 | |
4YLA
 
 | | Crystal structure of the indole prenyltransferase MpnD complexed with indolactam V and DMSPP | | Descriptor: | (2S,5S)-5-(hydroxymethyl)-1-methyl-2-(propan-2-yl)-1,2,4,5,6,8-hexahydro-3H-[1,4]diazonino[7,6,5-cd]indol-3-one, Aromatic prenyltransferase, DIMETHYLALLYL S-THIOLODIPHOSPHATE, ... | | Authors: | Mori, T, Morita, H, Abe, I. | | Deposit date: | 2015-03-05 | | Release date: | 2016-03-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases.
Nat Commun, 7, 2016
|
|
4YL7
 
 | |
4YZL
 
 | | Crystal structure of the indole prenyltransferase TleC complexed with indolactam V and DMSPP | | Descriptor: | (2S,5S)-5-(hydroxymethyl)-1-methyl-2-(propan-2-yl)-1,2,4,5,6,8-hexahydro-3H-[1,4]diazonino[7,6,5-cd]indol-3-one, DIMETHYLALLYL S-THIOLODIPHOSPHATE, Tryptophan dimethylallyltransferase | | Authors: | Mori, T, Morita, H, Abe, I. | | Deposit date: | 2015-03-25 | | Release date: | 2016-03-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.096 Å) | | Cite: | Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases.
Nat Commun, 7, 2016
|
|
4YZK
 
 | |
7WIJ
 
 | |
