3PPH
 
 | | Crystal structure of the Candida albicans methionine synthase by surface entropy reduction, threonine variant | | Descriptor: | 5-methyltetrahydropteroyltriglutamate--homocysteine methyltransferase | | Authors: | Ubhi, D, Kavanagh, K, Monzingo, A.F, Robertus, J.D. | | Deposit date: | 2010-11-24 | | Release date: | 2011-10-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of Candida albicans methionine synthase determined by employing surface residue mutagenesis.
Arch.Biochem.Biophys., 513, 2011
|
|
4HV3
 
 | | Structure of Ricin A chain bound with N-(N-(pterin-7-yl)carbonyl-L-serinyl)-L-tryptophan | | Descriptor: | (2S)-2-[[(2S)-2-[(2-azanyl-4-oxidanylidene-1H-pteridin-7-yl)carbonylamino]-3-oxidanyl-propanoyl]amino]-3-(1H-indol-3-yl)propanoic acid, MALONIC ACID, Ricin, ... | | Authors: | Robertus, J.D, Manzano, L.A, Jasheway, K.R, Monzingo, A.F, Saito, R, Pruet, J.M, Wiget, P.A, Anslyn, E.V. | | Deposit date: | 2012-11-05 | | Release date: | 2012-12-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Peptide-conjugated pterins as inhibitors of ricin toxin A.
J.Med.Chem., 56, 2013
|
|
3PPG
 
 | | Crystal structure of the Candida albicans methionine synthase by surface entropy reduction, alanine variant with zinc | | Descriptor: | 5-methyltetrahydropteroyltriglutamate--homocysteine methyltransferase, ZINC ION | | Authors: | Ubhi, D, Kavanagh, K, Monzingo, A.F, Robertus, J.D. | | Deposit date: | 2010-11-24 | | Release date: | 2011-10-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure of Candida albicans methionine synthase determined by employing surface residue mutagenesis.
Arch.Biochem.Biophys., 513, 2011
|
|
1OBT
 
 | | STRUCTURE OF RICIN A CHAIN MUTANT, COMPLEX WITH AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, RICIN A CHAIN | | Authors: | Day, P.J, Ernst, S.R, Frankel, A.E, Monzingo, A.F, Pascal, J.M, Svinth, M, Robertus, J.D. | | Deposit date: | 1996-06-22 | | Release date: | 1997-06-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure and activity of an active site substitution of ricin A chain.
Biochemistry, 35, 1996
|
|
1OBS
 
 | | STRUCTURE OF RICIN A CHAIN MUTANT | | Descriptor: | RICIN A CHAIN | | Authors: | Day, P.J, Ernst, S.R, Frankel, A.E, Monzingo, A.F, Pascal, J.M, Svinth, M, Robertus, J.D. | | Deposit date: | 1996-06-25 | | Release date: | 1997-06-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure and activity of an active site substitution of ricin A chain.
Biochemistry, 35, 1996
|
|
3EE8
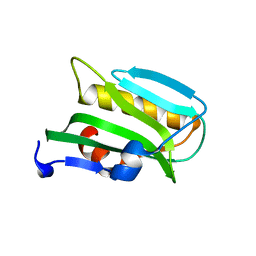 
 | |
3EE9
 
 | | Structure of NS1 effector domain | | Descriptor: | Non-structural protein 1, SULFATE ION | | Authors: | Xia, S, Monzingo, A.F, Robertus, J.D. | | Deposit date: | 2008-09-04 | | Release date: | 2009-01-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structure of NS1A effector domain from the influenza A/Udorn/72 virus.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3EJ5
 
 | |
3ET9
 
 | | Crystal structure of the engineered neutralizing antibody 1H | | Descriptor: | Antibody 1H light chain and antibody 1H heavy chain linked with a synthetic (GGGGS)4 linker | | Authors: | Leysath, C.E, Monzingo, A.F, Barnett, J, Iverson, B.L, Georgiou, G, Robertus, J.D. | | Deposit date: | 2008-10-07 | | Release date: | 2009-04-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the engineered neutralizing antibody m18 complexed to domain 4 of the anthrax protective antigen.
J.Mol.Biol., 387, 2009
|
|
1P6D
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS IN COMPLEX WITH (3S)-3,4,DI-N-HEXANOYLOXYBUTYL-1-PHOSPHOCHOLINE | | Descriptor: | (3S)-3,4-DI-N-HEXANOYLOXYBUTYL-1-PHOSPHOCHOLINE, PHOSPHOLIPASE C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-29 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
1P5X
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS | | Descriptor: | Phospholipase C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-28 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
1P6E
 
 | | STRUCTURE OF THE D55N MUTANT OF PHOSPHOLIPASE C FROM BACILLUS CEREUS IN COMPLEX WITH 1,2-DI-N-PENTANOYL-SN-GLYCERO-3-DITHIOPHOSPHOCHOLINE | | Descriptor: | 1,2-DI-N-PENTANOYL-SN-GLYCERO-3-DITHIOPHOSPHOCHOLINE, Phospholipase C, ZINC ION | | Authors: | Antikainen, N.M, Monzingo, A.F, Franklin, C.L, Robertus, J.D, Martin, S.F. | | Deposit date: | 2003-04-29 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base.
Arch.Biochem.Biophys., 417, 2003
|
|
4ESI
 
 | | Structure of ricin A chain bound with N-((1H-1,2,3-triazol-4-yl)methyl-2-amino-4-oxo-3,4-dihydropteridine-7-carboxamide | | Descriptor: | 2-amino-4-oxo-N-(1H-1,2,3-triazol-5-ylmethyl)-1,4-dihydropteridine-7-carboxamide, Ricin | | Authors: | Jasheway, K.R, Pruet, J.M, Ryoto, S, Manzano, L.A, Wiget, P.A, Kamat, I, Anslyn, E.V, Monzingo, A.F, Robertus, J.D. | | Deposit date: | 2012-04-23 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Optimized 5-membered heterocycle-linked pterins for the inhibition of Ricin Toxin A.
ACS Med Chem Lett, 3, 2012
|
|
1HWO
 
 | | EBULIN COMPLEXED WITH LACTOSE, TRIGONAL CRYSTAL FORM | | Descriptor: | EBULIN, beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Pascal, J.M, Day, P.J, Monzingo, A.F, Ernst, S.R, Robertus, J.D. | | Deposit date: | 2001-01-09 | | Release date: | 2001-01-24 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | 2.8-A crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l.
Proteins, 43, 2001
|
|
1IBT
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE AT-170 C | | Descriptor: | HISTIDINE DECARBOXYLASE ALPHA CHAIN, HISTIDINE DECARBOXYLASE BETA CHAIN | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1J1R
 
 | | Structure of Pokeweed Antiviral Protein from Seeds (PAP-S1) Complexed with Adenine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ADENINE, Antiviral Protein S | | Authors: | Watanabe, K, Sato, E, Honjo, E, Motoshima, H, Kurokawa, H, Mikami, B, Monzingo, A.F, Robertus, J.D, Fujii, H, Hidaka, A. | | Deposit date: | 2002-12-14 | | Release date: | 2004-02-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of Pokweed Antiviral Protein from Seeds (PAP-S1) at 1.8 Angstrom Resolution
To be published
|
|
1J1S
 
 | | Pokeweed Antiviral Protein from Seeds (PAP-S1) Complexed with Formycin | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Antiviral Protein S, FORMYCIN-5'-MONOPHOSPHATE | | Authors: | Watanabe, K, Sato, E, Honjo, E, Motoshima, H, Kurokawa, H, Mikami, B, Monzingo, A.F, Robertus, J.D, Fujii, H, Hidaka, A. | | Deposit date: | 2002-12-14 | | Release date: | 2004-02-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of Pokweed Antiviral Protein from Seeds (PAP-S1) at 1.8 Angstrom Resolution
To be published
|
|
1IBV
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT-170 C | | Descriptor: | HISTIDINE DECARBOXYLASE BETA CHAIN, HISTIDINE-METHYL-ESTER, Histidine decarboxylase alpha chain | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1LL7
 
 | |
1LL4
 
 | | STRUCTURE OF C. IMMITIS CHITINASE 1 COMPLEXED WITH ALLOSAMIDIN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-allopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-allopyranose, ALLOSAMIZOLINE, CHITINASE 1 | | Authors: | Bortone, K, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2002-04-26 | | Release date: | 2002-09-25 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | THE STRUCTURE OF AN ALLOSAMIDIN COMPLEX WITH THE Coccidioides IMMITIS CHITINASE DEFINES A ROLE FOR A SECOND ACID RESIDUE IN SUBSTRATE-ASSISTED MECHANISM
J.Mol.Biol., 320, 2002
|
|
1IBW
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT 25 C | | Descriptor: | HISTIDINE DECARBOXYLASE BETA CHAIN, HISTIDINE-METHYL-ESTER, Histidine decarboxylase alpha chain | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1IBU
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE AT 25 C | | Descriptor: | HISTIDINE DECARBOXYLASE ALPHA CHAIN, HISTIDINE DECARBOXYLASE BETA CHAIN | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1LL6
 
 | |
1J1Q
 
 | | Structure of Pokeweed Antiviral Protein from Seeds (PAP-S1) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Antiviral protein S | | Authors: | Watanabe, K, Sato, E, Honjo, E, Motoshima, H, Kurokawa, H, Mikami, B, Monzingo, A.F, Robertus, J.D, Fujii, H, Hidaka, A. | | Deposit date: | 2002-12-14 | | Release date: | 2004-02-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of Pokweed Antiviral Protein from Seeds (PAP-S1) at 1.8 Angstrom Resolution
To be published
|
|
4MFA
 
 | | Structure of human DNA polymerase beta complexed with nicked DNA containing a mismatched template O6MG and incoming TTP | | Descriptor: | DNA polymerase beta, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Koag, M.C, Min, K, Monzingo, A.F, Lee, S. | | Deposit date: | 2013-08-27 | | Release date: | 2014-08-27 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Structures of human DNA polymerase beta inserting bases opposite templating O6MG
To be Published
|
|
