8TXS
 
 | |
8TV4
 
 | | NMR structure of temporin L in solution | | Descriptor: | Temporin-1Tl peptide | | Authors: | McShan, A.C, Jia, R, Halim, M.A. | | Deposit date: | 2023-08-17 | | Release date: | 2023-09-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Antiviral peptides inhibiting the main protease of SARS-CoV-2 investigated by computational screening and in vitro protease assay.
J.Pept.Sci., 30, 2024
|
|
8T62
 
 | |
8T63
 
 | |
8T61
 | |
8TB1
 
 | |
7RNO
 
 | | Model of the Ac-6-FP/hpMR1/bB2m/TAPBPR complex from integrated docking, NMR and restrained MD | | Descriptor: | Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, N-(6-formyl-4-oxo-3,4-dihydropteridin-2-yl)acetamide, ... | | Authors: | McShan, A.C, Sgourakis, N.G. | | Deposit date: | 2021-07-29 | | Release date: | 2022-05-11 | | Last modified: | 2022-08-10 | | Method: | SOLUTION NMR | | Cite: | TAPBPR employs a ligand-independent docking mechanism to chaperone MR1 molecules.
Nat.Chem.Biol., 18, 2022
|
|
9BVU
 | | NMR structure of TLP-2 in solution | | Descriptor: | Temporin-1Tl | | Authors: | Jia, R, McShan, A.C. | | Deposit date: | 2024-05-20 | | Release date: | 2024-06-05 | | Method: | SOLUTION NMR | | Cite: | Design and Development of Temporin L Analogues to Inhibit the Main Protease of SARS-CoV-2
To Be Published
|
|
9BVV
 
 | | NMR structure of TLP-3 in solution | | Descriptor: | Temporin-1Tl | | Authors: | Jia, R, McShan, A.C. | | Deposit date: | 2024-05-20 | | Release date: | 2024-06-05 | | Method: | SOLUTION NMR | | Cite: | Design and Development of Temporin L Analogues to Inhibit the Main Protease of SARS-CoV-2
To Be Published
|
|
9BVS
 
 | | NMR structure of TLP-1 in solution | | Descriptor: | Temporin-1Tl | | Authors: | Jia, R, McShan, A.C. | | Deposit date: | 2024-05-20 | | Release date: | 2024-06-05 | | Method: | SOLUTION NMR | | Cite: | Design and Development of Temporin L Analogues to Inhibit the Main Protease of SARS-CoV-2
To Be Published
|
|
6MPP
 
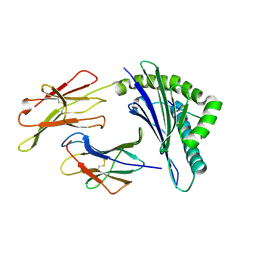 | |
6NPR
 
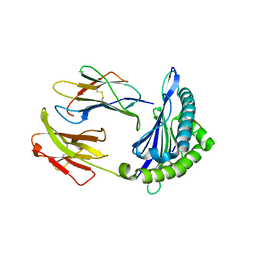 | | Crystal structure of H-2Dd with C84-C139 disulfide in complex with gp120 derived peptide P18-I10 | | Descriptor: | ARG-GLY-PRO-GLY-ARG-ALA-PHE-VAL-THR-ILE, Beta-2-microglobulin, H-2 class I histocompatibility antigen, ... | | Authors: | Toor, J, McShan, A.C, Tripathi, S.M, Sgourakis, N.G. | | Deposit date: | 2019-01-18 | | Release date: | 2020-01-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Molecular determinants of chaperone interactions on MHC-I for folding and antigen repertoire selection.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6OSW
 
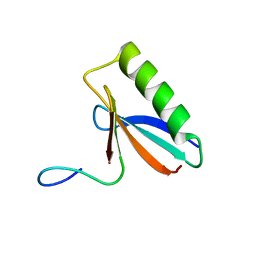 | | An order-to-disorder structural switch activates the FoxM1 transcription factor | | Descriptor: | Forkhead box M1 | | Authors: | Marceau, A.H, Rubin, S.M, Nerli, S, McShane, A.C, Sgourakis, N.G. | | Deposit date: | 2019-05-02 | | Release date: | 2019-05-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | An order-to-disorder structural switch activates the FoxM1 transcription factor.
Elife, 8, 2019
|
|
