1ARG
 
 | |
1BQA
 | |
1BQD
 
 | |
2CST
 
 | |
1VDF
 
 | |
1AKC
 
 | |
1AKA
 
 | |
1AKB
 
 | |
4Q7T
 | |
4Q7U
 
 | |
2EBO
 
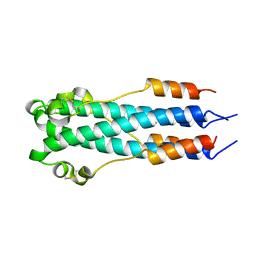 | | CORE STRUCTURE OF GP2 FROM EBOLA VIRUS | | Descriptor: | CHLORIDE ION, EBOLA VIRUS ENVELOPE GLYCOPROTEIN | | Authors: | Malashkevich, V.N, Schneider, B.J, Mcnally, M.L, Milhollen, M.A, Pang, J.X, Kim, P.S. | | Deposit date: | 1998-12-24 | | Release date: | 1999-05-18 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Core structure of the envelope glycoprotein GP2 from Ebola virus at 1.9-A resolution.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1YDY
 
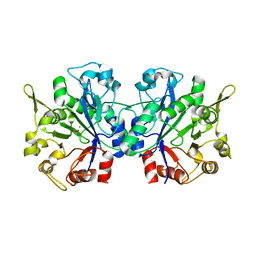 | | Crystal structure of periplasmic glycerophosphodiester phosphodiesterase from Escherichia coli | | Descriptor: | CALCIUM ION, GLYCEROL, Glycerophosphoryl diester phosphodiesterase | | Authors: | Malashkevich, V.N, Fedorov, E, Almo, S.C, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2004-12-27 | | Release date: | 2005-01-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of periplasmic glycerophosphodiester phosphodiesterase from Escherichia coli
To be Published
|
|
1Z3A
 
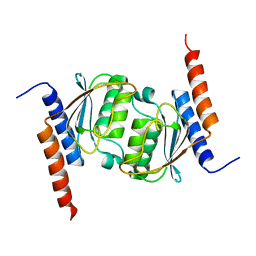 | | Crystal structure of tRNA adenosine deaminase TadA from Escherichia coli | | Descriptor: | ZINC ION, tRNA-specific adenosine deaminase | | Authors: | Malashkevich, V, Kim, J, Lisbin, M, Almo, S.C, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2005-03-10 | | Release date: | 2006-02-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Structural and kinetic characterization of Escherichia coli TadA, the wobble-specific tRNA deaminase.
Biochemistry, 45, 2006
|
|
1AHF
 
 | | ASPARTATE AMINOTRANSFERASE HEXAMUTANT | | Descriptor: | ASPARTATE AMINOTRANSFERASE, INDOLYLPROPIONIC ACID, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-02-22 | | Release date: | 1995-09-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase.
Nat.Struct.Biol., 2, 1995
|
|
1AHG
 
 | | ASPARTATE AMINOTRANSFERASE HEXAMUTANT | | Descriptor: | Aspartate aminotransferase, PYRIDOXAL-5'-PHOSPHATE, TYROSINE | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-02-22 | | Release date: | 1995-09-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase.
Nat.Struct.Biol., 2, 1995
|
|
1AHY
 
 | | ASPARTATE AMINOTRANSFERASE HEXAMUTANT | | Descriptor: | ASPARTATE AMINOTRANSFERASE, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-02-21 | | Release date: | 1995-09-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase.
Nat.Struct.Biol., 2, 1995
|
|
1AHE
 
 | | ASPARTATE AMINOTRANSFERASE HEXAMUTANT | | Descriptor: | ASPARTATE AMINOTRANSFERASE, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-02-22 | | Release date: | 1995-09-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase.
Nat.Struct.Biol., 2, 1995
|
|
1AHX
 
 | | ASPARTATE AMINOTRANSFERASE HEXAMUTANT | | Descriptor: | ASPARTATE AMINOTRANSFERASE, HYDROCINNAMIC ACID, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-02-21 | | Release date: | 1995-09-15 | | Last modified: | 2022-02-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase.
Nat.Struct.Biol., 2, 1995
|
|
1ARI
 
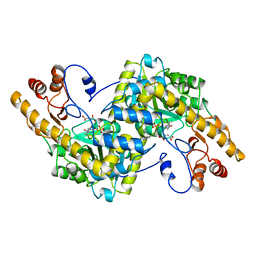 | | Aspartate aminotransferase, W140H mutant, maleate complex | | Descriptor: | ASPARTATE AMINOTRANSFERASE, MALEIC ACID, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-08-23 | | Release date: | 1995-11-14 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Substitution of apolar residues in the active site of aspartate aminotransferase by histidine. Effects on reaction and substrate specificity.
Eur.J.Biochem., 227, 1995
|
|
1ARH
 
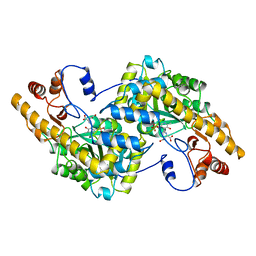 | | ASPARTATE AMINOTRANSFERASE, Y225R/R386A MUTANT | | Descriptor: | 2-[(3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYLENE)-AMINO]-SUCCINIC ACID, ASPARTATE AMINOTRANSFERASE | | Authors: | Malashkevich, V.N, Jansonius, J.N. | | Deposit date: | 1995-08-23 | | Release date: | 1995-11-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Changing the reaction specificity of a pyridoxal-5'-phosphate-dependent enzyme.
Eur.J.Biochem., 232, 1995
|
|
1YVT
 
 | | The high salt (phosphate) crystal structure of CO Hemoglobin E (Glu26Lys) at physiological pH (pH 7.35) | | Descriptor: | CARBON MONOXIDE, GLYCEROL, Hemoglobin alpha chain, ... | | Authors: | Malashkevich, V.N, Balazs, T.C, Almo, S.C, Hirsch, R.E. | | Deposit date: | 2005-02-16 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The high salt (phosphate) crystal structure of CO Hemoglobin E (Glu26Lys) at physiological pH (pH 7.35)
To be Published
|
|
1YVQ
 | | The low salt (PEG) crystal structure of CO Hemoglobin E (betaE26K) approaching physiological pH (pH 7.5) | | Descriptor: | CARBON MONOXIDE, Hemoglobin alpha chain, Hemoglobin beta chain, ... | | Authors: | Malashkevich, V.N, Balazs, T.C, Almo, S.C, Hirsch, R.E. | | Deposit date: | 2005-02-16 | | Release date: | 2006-02-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structures of CO Hemoglobin E (betaE26K) approaching physiological pH (pH 7.5)
To be Published
|
|
1ZZM
 
 | | Crystal structure of YJJV, TATD Homolog from Escherichia coli k12, at 1.8 A resolution | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, ZINC ION, putative deoxyribonuclease yjjV | | Authors: | Malashkevich, V.N, Xiang, D.F, Raushel, F.M, Almo, S.C, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2005-06-14 | | Release date: | 2005-06-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of YJJV, TATD homolog from Escherichia coli K12, at 1.8 A resolution
To be Published
|
|
1D7S
 
 | | CRYSTAL STRUCTURE OF THE COMPLEX OF 2,2-DIALKYLGLYCINE DECARBOXYLASE WITH DCS | | Descriptor: | D-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-N,O-CYCLOSERYLAMIDE, POTASSIUM ION, PROTEIN (2,2-DIALKYLGLYCINE DECARBOXYLASE (PYRUVATE)), ... | | Authors: | Malashkevich, V.N, Toney, M.D, Strop, P, Keller, J, Jansonius, J.N. | | Deposit date: | 1999-10-19 | | Release date: | 1999-11-19 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structures of dialkylglycine decarboxylase inhibitor complexes.
J.Mol.Biol., 294, 1999
|
|
1D7V
 
 | | CRYSTAL STRUCTURE OF THE COMPLEX OF 2,2-DIALKYLGLYCINE DECARBOXYLASE WITH NMA | | Descriptor: | N-[3-HYDROXY-2-METHYL-5-PHOSPHONOOXYMETHYL-PYRIDIN-4-YLMETHYL]-2-METHYLALANINE, POTASSIUM ION, PROTEIN (2,2-DIALKYLGLYCINE DECARBOXYLASE (PYRUVATE)), ... | | Authors: | Malashkevich, V.N, Toney, M.D, Strop, P, Keller, J, Jansonius, J.N. | | Deposit date: | 1999-10-19 | | Release date: | 1999-11-19 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structures of dialkylglycine decarboxylase inhibitor complexes.
J.Mol.Biol., 294, 1999
|
|
