8EVD
 | |
8F6U
 | |
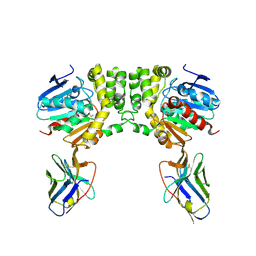
8F6V
 
 | |
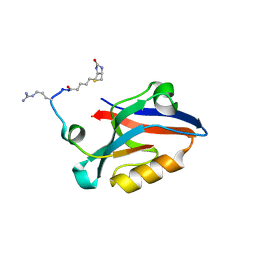
4Q6S
 
 | |
4Q5P
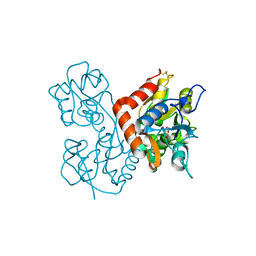
 | | Lysine-Ligated Yeast Iso-1 Cytochrome C | | Descriptor: | Cytochrome c iso-1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Amacher, J.F, Zhu, M.Q, Zhong, F, Pletneva, E.V, Madden, D.R. | | Deposit date: | 2014-04-17 | | Release date: | 2015-04-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | A Compact Structure of Cytochrome c Trapped in a Lysine-Ligated State: Loop Refolding and Functional Implications of a Conformational Switch.
J.Am.Chem.Soc., 137, 2015
|
|
3KDA
 
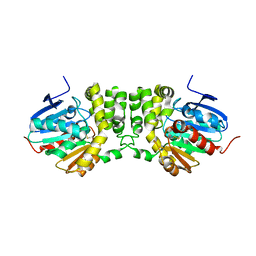 | |
3KEI
 
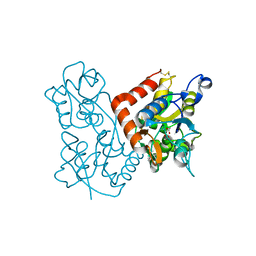 | |
3KFM
 
 | |
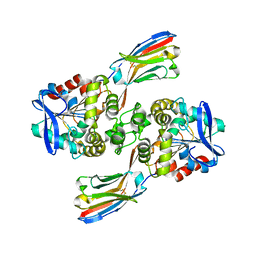
8ELN
 
 | |
4E34
 
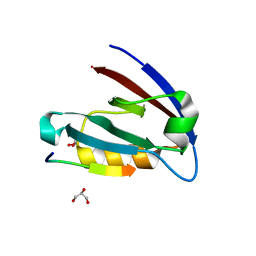 | | Crystal structure of CFTR Associated Ligand (CAL) PDZ domain bound to iCAL36 (ANSRWPTSII) peptide | | Descriptor: | GLYCEROL, Golgi-associated PDZ and coiled-coil motif-containing protein, decameric peptide, ... | | Authors: | Amacher, J.F, Beck, T, Madden, D.R. | | Deposit date: | 2012-03-09 | | Release date: | 2012-12-26 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.395 Å) | | Cite: | Stereochemical Determinants of C-terminal Specificity in PDZ Peptide-binding Domains: A NOVEL CONTRIBUTION OF THE CARBOXYLATE-BINDING LOOP.
J.Biol.Chem., 288, 2013
|
|
4E3B
 | |
4EHB
 
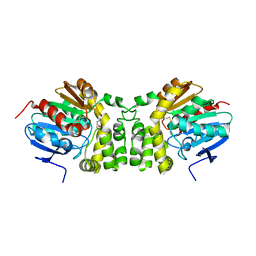 | |
4EUS
 
 | |
4DMK
 
 | |
4DMF
 
 | |
4DMH
 
 | |
4DMC
 
 | |
4DNO
 
 | | Crystal structure of the CFTR inhibitory factor Cif with the E153Q mutation adducted with the 1,2-epoxyhexane hydrolysis intermediate | | Descriptor: | (2R)-hexane-1,2-diol, Putative hydrolase | | Authors: | Bahl, C.D, Patankar, Y.R, Madden, D.R. | | Deposit date: | 2012-02-08 | | Release date: | 2013-08-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Visualizing the Mechanism of Epoxide Hydrolysis by the Bacterial Virulence Enzyme Cif.
Biochemistry, 55, 2016
|
|
4DNF
 
 | |
4DM7
 
 | |
4DLN
 
 | |
4E35
 
 | |
2J80
 
 | | Structure of Citrate-bound Periplasmic Domain of Sensor Histidine Kinase CitA | | Descriptor: | CITRATE ANION, GLYCEROL, SENSOR KINASE CITA, ... | | Authors: | Sevvana, M, Vijayan, V, Zweckstetter, M, Reinelt, S, Madden, D.R, Sheldrick, G.M, Bott, M, Griesinger, C, Becker, S. | | Deposit date: | 2006-10-18 | | Release date: | 2007-10-23 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A Ligand-Induced Switch in the Periplasmic Domain of Sensor Histidine Kinase Cita.
J.Mol.Biol., 377, 2008
|
|
2LOB
 
 | | PDZ Domain of CAL (Cystic Fibrosis Transmembrane Regulator-Associated Ligand) | | Descriptor: | Cystic fibrosis transmembrane conductance regulator, Golgi-associated PDZ and coiled-coil motif-containing protein | | Authors: | Piserchio, A, Fellows, A, Madden, D.R, Mierke, D.F. | | Deposit date: | 2012-01-20 | | Release date: | 2012-06-13 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Association of the cystic fibrosis transmembrane regulator with CAL: structural features and molecular dynamics.
Biochemistry, 44, 2005
|
|
6V84
 
 | |
