2KAX
 
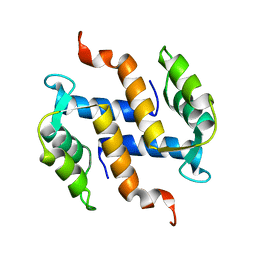 | | Solution structure and dynamics of S100A5 in the apo and Ca2+ -bound states | | Descriptor: | Protein S100-A5 | | Authors: | Bertini, I, Das Gupta, S, Hu, X, Karavelas, T, Luchinat, C, Parigi, G, Yuan, J, Structural Proteomics in Europe (SPINE), Structural Proteomics in Europe 2 (SPINE-2) | | Deposit date: | 2008-11-17 | | Release date: | 2009-06-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of S100A5 in the apo and Ca2+-bound states
J.Biol.Inorg.Chem., 14, 2009
|
|
2LLP
 
 | | Solution structure of a THP type 1 alpha 1 collagen fragment (772-786) | | Descriptor: | Collagen alpha-1(I) chain | | Authors: | Bertini, I, Fragai, M, Luchinat, C, Melikian, M, Toccafondi, M, Lauer, J.L, Fields, G.B, Structural Proteomics in Europe 2 (SPINE-2) | | Deposit date: | 2011-11-15 | | Release date: | 2012-05-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis for matrix metalloproteinase 1-catalyzed collagenolysis.
J.Am.Chem.Soc., 134, 2012
|
|
2OY4
 
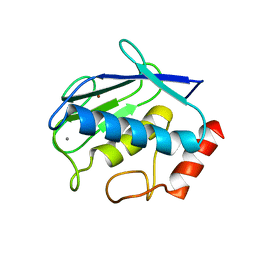 | | Uninhibited human MMP-8 | | Descriptor: | CALCIUM ION, Neutrophil collagenase, ZINC ION | | Authors: | Calderone, V, Bertini, I, Fragai, M, Luchinat, C, Maletta, M, Yeo, K.J. | | Deposit date: | 2007-02-21 | | Release date: | 2007-03-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Snapshots of the reaction mechanism of matrix metalloproteinases.
ANGEW.CHEM.INT.ED.ENGL., 45, 2006
|
|
2JXY
 
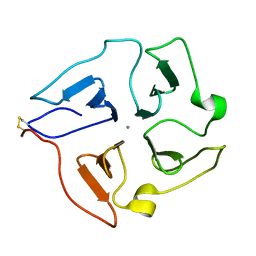 | | Solution structure of the hemopexin-like domain of MMP12 | | Descriptor: | CALCIUM ION, Macrophage metalloelastase | | Authors: | Bertini, I, Calderone, V, Fragai, M, Jaiswal, R, Luchinat, C, Melikian, M. | | Deposit date: | 2007-12-01 | | Release date: | 2008-05-27 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Evidence of reciprocal reorientation of the catalytic and hemopexin-like domains of full-length MMP-12
J.Am.Chem.Soc., 130, 2008
|
|
2OY2
 
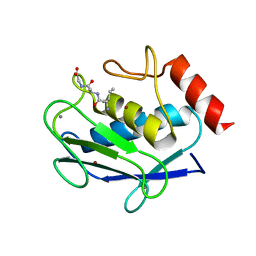 | | Human MMP-8 in complex with peptide IAG | | Descriptor: | CALCIUM ION, ILE-ALA-GLY peptide, Neutrophil collagenase, ... | | Authors: | Calderone, V, Bertini, I, Fragai, M, Luchinat, C, Maletta, M, Yeo, K.J. | | Deposit date: | 2007-02-21 | | Release date: | 2007-03-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Snapshots of the reaction mechanism of matrix metalloproteinases.
ANGEW.CHEM.INT.ED.ENGL., 45, 2006
|
|
1BFY
 
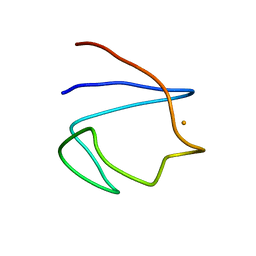 | | SOLUTION STRUCTURE OF REDUCED CLOSTRIDIUM PASTEURIANUM RUBREDOXIN, NMR, 20 STRUCTURES | | Descriptor: | FE (III) ION, RUBREDOXIN | | Authors: | Bertini, I, Kurtz Junior, D.M, Eidsness, M.K, Liu, G, Luchinat, C, Rosato, A, Scott, R.A. | | Deposit date: | 1998-05-23 | | Release date: | 1999-05-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Reduced Clostridium Pasteurianum Rubredoxin
J.Biol.Inorg.Chem., 3, 1998
|
|
2L50
 
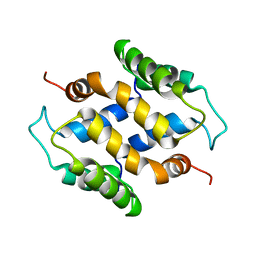 | | Solution structure of apo S100A16 | | Descriptor: | Protein S100-A16 | | Authors: | Babini, E, Bertini, I, Borsi, V, Calderone, V, Hu, X, Luchinat, C, Parigi, G. | | Deposit date: | 2010-10-22 | | Release date: | 2010-11-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of human S100A16, a low-affinity calcium binder.
J.Biol.Inorg.Chem., 16, 2011
|
|
2MKM
 
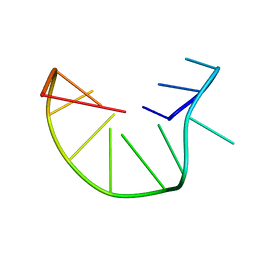 | | G-triplex structure and formation propensity | | Descriptor: | DNA_(5'-D(*GP*GP*TP*TP*GP*GP*TP*GP*TP*GP*G)-3') | | Authors: | Cerofolini, L, Fragai, M, Giachetti, A, Limongelli, V, Luchinat, C, Novellino, E, Parrinello, M, Randazzo, A. | | Deposit date: | 2014-02-10 | | Release date: | 2014-11-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | G-triplex structure and formation propensity.
Nucleic Acids Res., 42, 2014
|
|
2MKO
 
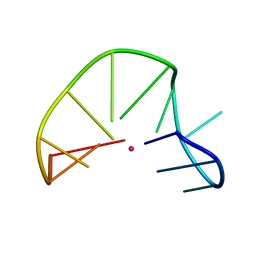 | | G-triplex structure and formation propensity | | Descriptor: | DNA_(5'-D(*GP*GP*TP*TP*GP*GP*TP*GP*TP*GP*G)-3'), POTASSIUM ION | | Authors: | Cerofolini, L, Fragai, M, Giachetti, A, Limongelli, V, Luchinat, C, Novellino, E, Parrinello, M, Randazzo, A. | | Deposit date: | 2014-02-11 | | Release date: | 2014-11-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | G-triplex structure and formation propensity.
Nucleic Acids Res., 42, 2014
|
|
2M0R
 
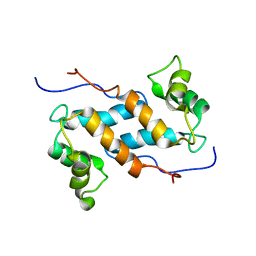 | | Solution structure and dynamics of human S100A14 | | Descriptor: | Protein S100-A14 | | Authors: | Bertini, I, Borsi, V, Cerofolini, L, Das Gupta, S, Fragai, M, Luchinat, C. | | Deposit date: | 2012-11-05 | | Release date: | 2013-01-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of human S100A14.
J.Biol.Inorg.Chem., 18, 2013
|
|
6RLY
 
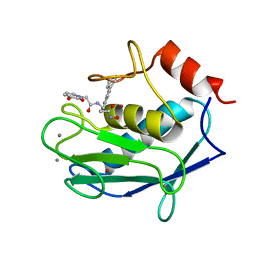 | | Human MMP12 (catalytic domain) in complex with AP316 | | Descriptor: | 2-(3-oxidanyl-2-oxidanylidene-pyridin-1-yl)-~{N}-[2-(4-phenylphenyl)ethyl]ethanamide, ACETOHYDROXAMIC ACID, CALCIUM ION, ... | | Authors: | Calderone, V, Fragai, M, Luchinat, C. | | Deposit date: | 2019-05-03 | | Release date: | 2020-03-11 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Exploration of zinc-binding groups for the design of inhibitors for the oxytocinase subfamily of M1 aminopeptidases.
Bioorg.Med.Chem., 27, 2019
|
|
6RD0
 
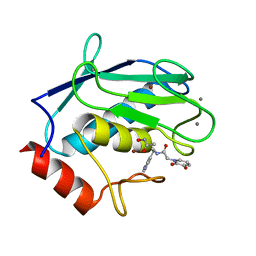 | | Human MMP12 catalytic domain in complex with AP280 | | Descriptor: | ACETOHYDROXAMIC ACID, CALCIUM ION, Macrophage metalloelastase, ... | | Authors: | Calderone, V, Fragai, M, Luchinat, C. | | Deposit date: | 2019-04-12 | | Release date: | 2020-02-19 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Exploration of zinc-binding groups for the design of inhibitors for the oxytocinase subfamily of M1 aminopeptidases.
Bioorg.Med.Chem., 27, 2019
|
|
