2NQK
 | | MoeA D59N mutant | | Descriptor: | Molybdopterin biosynthesis protein moeA | | Authors: | Nicolas, J, Xiang, S, Schindelin, H, Rajagopalan, K.V. | | Deposit date: | 2006-10-31 | | Release date: | 2007-01-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Mutational Analysis of Escherichia coli MoeA: Two Functional Activities Map to the Active Site Cleft.
Biochemistry, 46, 2007
|
|
2NQM
 
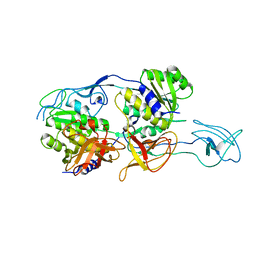 | | MoeA T100A mutant | | Descriptor: | Molybdopterin biosynthesis protein moeA | | Authors: | Nicoas, J, Xiang, S, Schindelin, H, Rajagopalan, K.V. | | Deposit date: | 2006-10-31 | | Release date: | 2007-01-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Mutational Analysis of Escherichia coli MoeA: Two Functional Activities Map to the Active Site Cleft.
Biochemistry, 46, 2007
|
|
1FMA
 
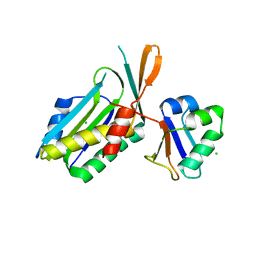 | | MOLYBDOPTERIN SYNTHASE (MOAD/MOAE) | | Descriptor: | CHLORIDE ION, MOLYBDOPTERIN CONVERTING FACTOR, SUBUNIT 1, ... | | Authors: | Rudolph, M.J, Wuebbens, M.M, Rajagolpalan, K.V, Schindelin, H. | | Deposit date: | 2000-08-16 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal structure of molybdopterin synthase and its evolutionary relationship to ubiquitin activation.
Nat.Struct.Biol., 8, 2001
|
|
4GLF
 
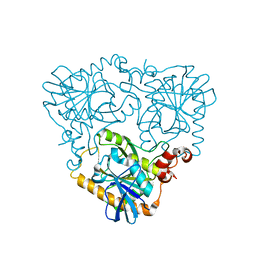 | |
1NVI
 
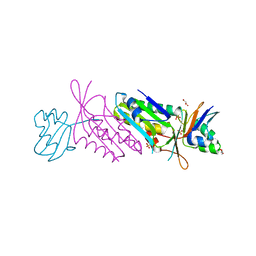 | | Orthorhombic Crystal Form of Molybdopterin Synthase | | Descriptor: | GLYCEROL, Molybdopterin converting factor subunit 1, Molybdopterin converting factor subunit 2, ... | | Authors: | Rudolph, M.J, Wuebbens, M.M, Turque, O, Rajagopalan, K.V, Schindelin, H. | | Deposit date: | 2003-02-03 | | Release date: | 2003-05-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Studies of Molybdopterin Synthase Provide Insights into its Catalytic Mechanism
J.Biol.Chem., 278, 2003
|
|
1MVZ
 
 | | NMR solution structure of a Bowman Birk inhibitor isolated from snail medic seeds (Medicago Scutellata) | | Descriptor: | Bowman-Birk type protease inhibitor, (MSTI) | | Authors: | Catalano, M, Ragona, L, Molinari, H, Tava, A, Zetta, L. | | Deposit date: | 2002-09-27 | | Release date: | 2003-04-22 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Anticarcinogenic Bowman Birk inhibitor isolated from snail medic seeds (Medicago Scutellata): solution structure and analysis of self-association behaviour
Biochemistry, 42, 2003
|
|
4D81
 
 | | Metallosphera sedula Vps4 crystal structure | | Descriptor: | AAA ATPase, central domain protein, ADENOSINE-5'-DIPHOSPHATE | | Authors: | Caillat, C, Macheboeuf, P, Wu, Y, McCarthy, A.A, Boeri-Erba, E, Effantin, G, Gottlinger, H.G, Weissenhorn, W, Renesto, P. | | Deposit date: | 2014-12-02 | | Release date: | 2015-11-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Asymmetric Ring Structure of Vps4 Required for Escrt-III Disassembly.
Nat.Commun., 6, 2015
|
|
1OVN
 
 | | Crystal Structure and Functional Analysis of Drosophila Wind-- a PDI-Related Protein | | Descriptor: | CESIUM ION, Windbeutel | | Authors: | Ma, Q, Guo, C, Barnewitz, K, Sheldrick, G.M, Soling, H.D, Uson, I, Ferrari, D.M. | | Deposit date: | 2003-03-27 | | Release date: | 2004-02-24 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure and functional analysis of Drosophila Wind, a protein-disulfide isomerase-related protein.
J.Biol.Chem., 278, 2003
|
|
4GLJ
 
 | | Crystal structure of methylthioadenosine phosphorylase in complex with rhodamine B | | Descriptor: | CHLORIDE ION, N-[9-(2-carboxyphenyl)-6-(diethylamino)-3H-xanthen-3-ylidene]-N-ethylethanaminium, PHOSPHATE ION, ... | | Authors: | Bujacz, A, Bujacz, G, Cieslinski, H, Bartasun, P. | | Deposit date: | 2012-08-14 | | Release date: | 2013-02-20 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A study on the interaction of rhodamine B with methylthioadenosine phosphorylase protein sourced from an antarctic soil metagenomic library.
Plos One, 8, 2013
|
|
2LFO
 
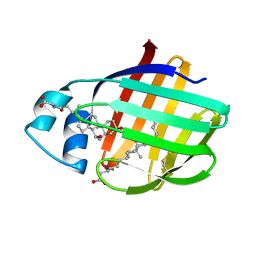 | | NMR structure of cl-BABP/SS complexed with glycochenodeoxycholic and glycocholic acids | | Descriptor: | Fatty acid-binding protein, liver, GLYCOCHENODEOXYCHOLIC ACID, ... | | Authors: | Tomaselli, S, Cogliati, C, Pagano, K, Zetta, L, Zanzoni, S, Assfalg, M, Molinari, H, Ragona, L. | | Deposit date: | 2011-07-07 | | Release date: | 2012-07-11 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | A disulfide bridge allows for site-selective binding in liver bile acid binding protein thereby stabilising the orientation of key amino acid side chains.
Chemistry, 18, 2012
|
|
1QEZ
 
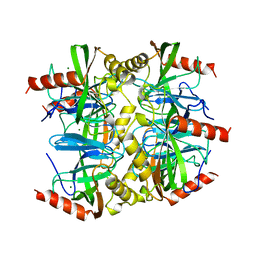 | | SULFOLOBUS ACIDOCALDARIUS INORGANIC PYROPHOSPHATASE: AN ARCHAEL PYROPHOSPHATASE. | | Descriptor: | MAGNESIUM ION, PROTEIN (INORGANIC PYROPHOSPHATASE) | | Authors: | Leppanen, V.-M, Nummelin, H, Hansen, T, Lahti, R, Schafer, G, Goldman, A. | | Deposit date: | 1999-04-06 | | Release date: | 1999-04-14 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Sulfolobus acidocaldarius inorganic pyrophosphatase: structure, thermostability, and effect of metal ion in an archael pyrophosphatase.
Protein Sci., 8, 1999
|
|
1DI7
 
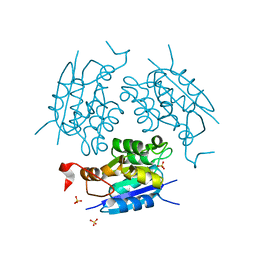 | | 1.60 ANGSTROM CRYSTAL STRUCTURE OF THE MOLYBDENUM COFACTOR BIOSYNTHESIS PROTEIN MOGA FROM ESCHERICHIA COLI | | Descriptor: | MOLYBDENUM COFACTOR BIOSYNTHETIC ENZYME, SULFATE ION | | Authors: | Liu, M.T.W, Wuebbens, M.M, Rajagopalan, K.V, Schindelin, H. | | Deposit date: | 1999-11-29 | | Release date: | 2000-01-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of the gephyrin-related molybdenum cofactor biosynthesis protein MogA from Escherichia coli.
J.Biol.Chem., 275, 2000
|
|
1DI6
 
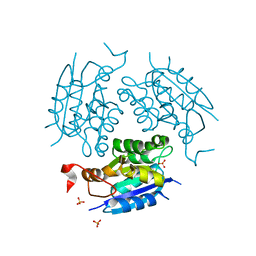 | | 1.45 A CRYSTAL STRUCTURE OF THE MOLYBDENUMM COFACTOR BIOSYNTHESIS PROTEIN MOGA FROM ESCHERICHIA COLI | | Descriptor: | MOLYBDENUM COFACTOR BIOSYNTHETIC ENZYME, SULFATE ION | | Authors: | Liu, M.T.W, Wuebbens, M.M, Rajagopalan, K.V, Schindelin, H. | | Deposit date: | 1999-11-29 | | Release date: | 2000-01-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal structure of the gephyrin-related molybdenum cofactor biosynthesis protein MogA from Escherichia coli.
J.Biol.Chem., 275, 2000
|
|
1DMS
 
 | | STRUCTURE OF DMSO REDUCTASE | | Descriptor: | 2-AMINO-5,6-DIMERCAPTO-7-METHYL-3,7,8A,9-TETRAHYDRO-8-OXA-1,3,9,10-TETRAAZA-ANTHRACEN-4-ONE GUANOSINE DINUCLEOTIDE, DMSO REDUCTASE, MOLYBDENUM (IV)OXIDE | | Authors: | Schneider, F, Loewe, J, Huber, R, Schindelin, H, Kisker, C, Knaeblein, J. | | Deposit date: | 1996-09-03 | | Release date: | 1998-07-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystal structure of dimethyl sulfoxide reductase from Rhodobacter capsulatus at 1.88 A resolution.
J.Mol.Biol., 263, 1996
|
|
2NAY
 
 | |
2PKO
 
 | |
