280D
 
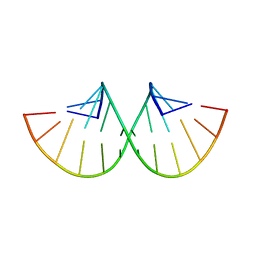 | |
387D
 
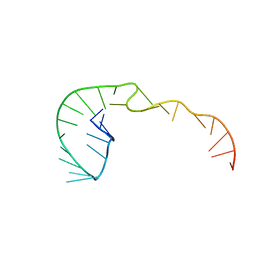 | | RNA Pseudoknot with 3D Domain Swapping | | Descriptor: | RNA Pseudoknot | | Authors: | Lietzke, S.E, Kundrot, C.E, Barnes, C.L. | | Deposit date: | 1998-04-14 | | Release date: | 2003-08-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The Structure of an RNA Pseudoknot Shows 3D Domain Swapping
Structure, Motion, Interaction and Expression of Biological Macromolecules, The Proceedings of the Tenth Conversation held at The University-SUNY, Albany NY, June 17-21, 1997, 10, 1998
|
|
1AIR
 
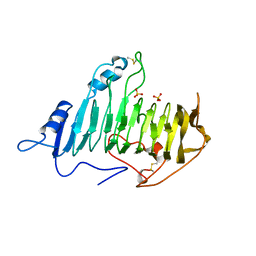 | | PECTATE LYASE C FROM ERWINIA CHRYSANTHEMI (EC16) TO A RESOLUTION OF 2.2 ANGSTROMS WITH 128 WATERS | | Descriptor: | PECTATE LYASE C, SULFATE ION | | Authors: | Lietzke, S.E, Scavetta, R.D, Yoder, M.D, Jurnak, F.A. | | Deposit date: | 1997-04-24 | | Release date: | 1997-06-16 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Refined Three-Dimensional Structure of Pectate Lyase E from Erwinia chrysanthemi at 2.2 A Resolution.
Plant Physiol., 111, 1996
|
|
1PCL
 
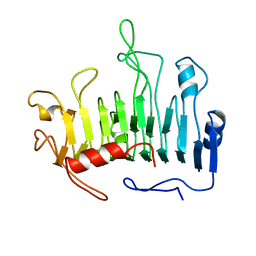 | |
1FGZ
 
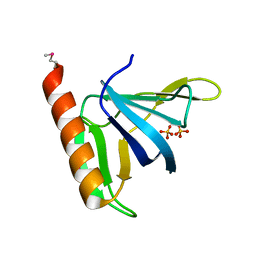 | | GRP1 PH DOMAIN (UNLIGANDED) | | Descriptor: | GRP1, SULFATE ION | | Authors: | Lietzke, S.E, Bose, S, Cronin, T, Klarlund, J, Chawla, A, Czech, M.P, Lambright, D.G. | | Deposit date: | 2000-07-29 | | Release date: | 2000-08-23 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis of 3-phosphoinositide recognition by pleckstrin homology domains.
Mol.Cell, 6, 2000
|
|
1FGY
 
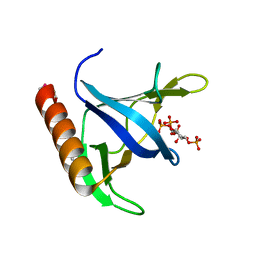 | | GRP1 PH DOMAIN WITH INS(1,3,4,5)P4 | | Descriptor: | GRP1, INOSITOL-(1,3,4,5)-TETRAKISPHOSPHATE | | Authors: | Lietzke, S.E, Bose, S, Cronin, T, Klarlund, J, Chawla, A, Czech, M.P, Lambright, D.G. | | Deposit date: | 2000-07-29 | | Release date: | 2000-08-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis of 3-phosphoinositide recognition by pleckstrin homology domains.
Mol.Cell, 6, 2000
|
|
1HXR
 | | CRYSTAL STRUCTURE OF MSS4 AT 1.65 ANGSTROMS | | Descriptor: | GUANINE NUCLEOTIDE EXCHANGE FACTOR MSS4, ZINC ION | | Authors: | Zhu, Z, Dumas, J.J, Lietzke, S.E, Lambright, D.G. | | Deposit date: | 2001-01-16 | | Release date: | 2001-02-14 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A helical turn motif in Mss4 is a critical determinant of Rab binding and nucleotide release.
Biochemistry, 40, 2001
|
|
