1TDR
 
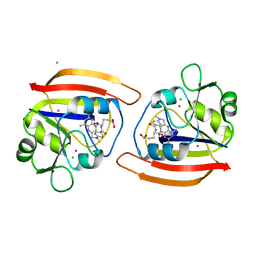 | | EXPRESSION, CHARACTERIZATION, AND CRYSTALLOGRAPHIC ANALYSIS OF TELLUROMETHIONYL DIHYDROFOLATE REDUCTASE | | Descriptor: | CALCIUM ION, CHLORIDE ION, METHOTREXATE, ... | | Authors: | Lewinski, K, Lebioda, L. | | Deposit date: | 1995-04-13 | | Release date: | 1995-07-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Expression, characterization and crystallographic analysis of telluromethionyl dihydrofolate reductase.
Acta Crystallogr.,Sect.D, 51, 1995
|
|
3I7X
 
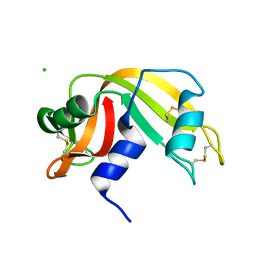 | | High pressure structure of I106A RNase A variant (0.35 GPa) | | Descriptor: | CHLORIDE ION, Ribonuclease pancreatic | | Authors: | Lewinski, K, Kurpiewska, K, Dziubek, K, Katrusiak, A, Font, J, Ribo, M, Vilanova, M. | | Deposit date: | 2009-07-09 | | Release date: | 2009-08-04 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural investigation of ribonuclease A conformational preferences using high pressure protein crystallography
Chem.Phys., 468, 2016
|
|
3I7Y
 
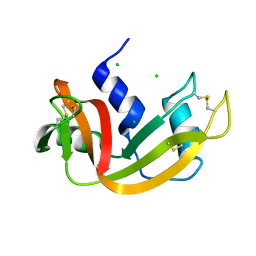 | | High pressure structure of I106A variant of RNase A (0.48 GPa) | | Descriptor: | CHLORIDE ION, Ribonuclease pancreatic | | Authors: | Lewinski, K, Kurpiewska, K, Dziubek, K, Katrusiak, A, Font, J, Ribo, M, Vilanova, M. | | Deposit date: | 2009-07-09 | | Release date: | 2009-08-04 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural investigation of ribonuclease A conformational preferences using high pressure protein crystallography
Chem.Phys., 468, 2016
|
|
3I7W
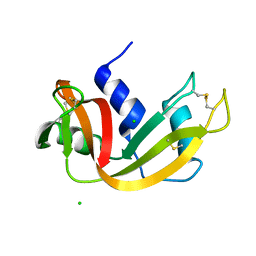 
 | | High pressure structure of wild-type RNase A (0.67 GPa) | | Descriptor: | CHLORIDE ION, Ribonuclease pancreatic | | Authors: | Lewinski, K, Kurpiewska, K, Dziubek, K, Katrusiak, A, Font, J, Ribo, M, Vilanova, M. | | Deposit date: | 2009-07-09 | | Release date: | 2009-08-04 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural investigation of ribonuclease A conformational preferences using high pressure protein crystallography
Chem.Phys., 468, 2016
|
|
7NRZ
 
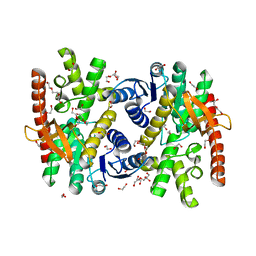 | | Crystal structure of malate dehydrogenase from Trypanosoma cruzi | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Sonani, R.R, Kurpiewska, K, Lewinski, K, Dubin, G. | | Deposit date: | 2021-03-04 | | Release date: | 2022-02-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Distinct sequence and structural feature of trypanosoma malate dehydrogenase.
Biochem.Biophys.Res.Commun., 557, 2021
|
|
6HCE
 
 | |
5VGM
 
 | | Crystal structure of dihydroorotase pyrC from Vibrio cholerae in complex with zinc at 1.95 A resolution. | | Descriptor: | ACETATE ION, CHLORIDE ION, Dihydroorotase, ... | | Authors: | Lipowska, J, Shabalin, I.G, Miks, C.D, Winsor, J, Cooper, D.R, Shuvalova, L, Kwon, K, Lewinski, K, Anderson, W.F, Minor, W, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2017-04-11 | | Release date: | 2017-04-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Pyrimidine biosynthesis in pathogens - Structures and analysis of dihydroorotases from Yersinia pestis and Vibrio cholerae.
Int.J.Biol.Macromol., 136, 2019
|
|
6XVE
 
 | |
3PHN
 
 | | Crystal structure of wild-type onconase with resolution 1.46 A | | Descriptor: | ACETATE ION, Protein P-30, SULFATE ION | | Authors: | Kurpiewska, K, Torrent, G, Ribo, M, Vilanova, M, Loch, J, Lewinski, K. | | Deposit date: | 2010-11-04 | | Release date: | 2010-11-17 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Structure of Rana pipiens wild-type onconase at resolution 1.46 A
To be Published
|
|
6CIA
 
 | | Crystal structure of aldo-keto reductase from Klebsiella pneumoniae in complex with NADPH. | | Descriptor: | Aldo/keto reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Lipowska, J, Leung, E.S, Shabalin, I.G, Grabowski, M, Almo, S.C, Satchell, K.J, Joachimiak, A, Lewinski, K, Minor, W, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2018-02-23 | | Release date: | 2018-03-07 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of of aldo-keto reductase from Klebsiella pneumoniae in complex with NADPH.
to be published
|
|
7Q2N
 
 | | Beta-lactoglobulin mutant FAF (I56F/L39A/M107F) in complex with desipramine (FAF-DSM) | | Descriptor: | 1,2-ETHANEDIOL, 3-(10,11-DIHYDRO-5H-DIBENZO[B,F]AZEPIN-5-YL)-N-METHYLPROPAN-1-AMINE, Beta-lactoglobulin, ... | | Authors: | Loch, J.I, Barciszewski, J, Lewinski, K. | | Deposit date: | 2021-10-25 | | Release date: | 2022-05-11 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
7Q2P
 
 | | Beta-lactoglobulin mutant FAW (I56F/L39A/M107W) in complex with desipramine (FAW-DSM#2) | | Descriptor: | 1,2-ETHANEDIOL, 3-(10,11-DIHYDRO-5H-DIBENZO[B,F]AZEPIN-5-YL)-N-METHYLPROPAN-1-AMINE, Beta-lactoglobulin, ... | | Authors: | Loch, J.I, Barciszewski, J, Pokrywka, K, Lewinski, K. | | Deposit date: | 2021-10-25 | | Release date: | 2022-05-11 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
7Q2O
 
 | | Beta-lactoglobulin mutant FAW (I56F/L39A/M107W) in complex with desipramine (FAW-DSM#1) | | Descriptor: | 1,2-ETHANEDIOL, 3-(10,11-DIHYDRO-5H-DIBENZO[B,F]AZEPIN-5-YL)-N-METHYLPROPAN-1-AMINE, Beta-lactoglobulin, ... | | Authors: | Loch, J.I, Barciszewski, J, Lewinski, K. | | Deposit date: | 2021-10-25 | | Release date: | 2022-05-11 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
7Q19
 
 | | Beta-lactoglobulin mutant FAW (I56F/L39A/M107W) in complex with desipramine (FAW-DSM#3) | | Descriptor: | 1,2-ETHANEDIOL, 3-(10,11-DIHYDRO-5H-DIBENZO[B,F]AZEPIN-5-YL)-N-METHYLPROPAN-1-AMINE, Beta-lactoglobulin, ... | | Authors: | Loch, J.I, Barciszewski, J, Pokrywka, K, Lewinski, K. | | Deposit date: | 2021-10-18 | | Release date: | 2022-05-11 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
7Q18
 
 | | Beta-lactoglobulin mutant FAF (I56F/L39A/M107F), unliganded form | | Descriptor: | Beta-lactoglobulin, SULFATE ION | | Authors: | Loch, J.I, Cymborowski, M.T, Minor, W, Lewinski, K. | | Deposit date: | 2021-10-18 | | Release date: | 2022-05-11 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.804 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
7Q17
 
 | | Beta-lactoglobulin mutant FAW (I56F/L39A/M107W), unliganded form | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactoglobulin | | Authors: | Loch, J.I, Barciszewski, J, Lewinski, K. | | Deposit date: | 2021-10-18 | | Release date: | 2022-05-11 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | New ligand-binding sites identified in the crystal structures of [beta]-lactoglobulin complexes with desipramine
Iucrj, 9, 2022
|
|
4DQ3
 
 | | Bovine beta-lactoglobulin complex with oleic acid | | Descriptor: | Beta-lactoglobulin, ETHANOL, GLYCEROL, ... | | Authors: | Loch, J.I, Lewinski, K. | | Deposit date: | 2012-02-15 | | Release date: | 2012-02-29 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binding of 18-carbon unsaturated fatty acids to bovine beta-lactoglobulin--structural and thermodynamic studies.
Int.J.Biol.Macromol., 57, 2013
|
|
4DQ4
 
 | | Bovine beta-lactoglobulin complex with linoleic acid | | Descriptor: | Beta-lactoglobulin, ETHANOL, GLYCEROL, ... | | Authors: | Loch, J.I, Lewinski, K. | | Deposit date: | 2012-02-15 | | Release date: | 2012-02-29 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binding of 18-carbon unsaturated fatty acids to bovine beta-lactoglobulin--structural and thermodynamic studies.
Int.J.Biol.Macromol., 57, 2013
|
|
5HTD
 
 | | Recombinant bovine beta-lactoglobulin variant L1A/I2S with endogenous ligand (sBlgB#1) | | Descriptor: | Beta-lactoglobulin, MYRISTIC ACID | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Polit, A, Hawro, B, Lach, A, Ludwin, E, Lewinski, K. | | Deposit date: | 2016-01-26 | | Release date: | 2016-07-20 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Engineered beta-Lactoglobulin Produced in E. coli: Purification, Biophysical and Structural Characterisation.
Mol Biotechnol., 58, 2016
|
|
5HTE
 
 | | Recombinant bovine beta-lactoglobulin variant L1A/I2S (sBlgB#2) | | Descriptor: | Beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Polit, A, Hawro, B, Lach, A, Ludwin, E, Lewinski, K. | | Deposit date: | 2016-01-26 | | Release date: | 2016-07-20 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Engineered beta-Lactoglobulin Produced in E. coli: Purification, Biophysical and Structural Characterisation.
Mol Biotechnol., 58, 2016
|
|
5NUM
 
 | | Engineered beta-lactoglobulin: variant F105L-L39A in complex with chlorpromazine (LG-LA-CLP) | | Descriptor: | 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, Beta-lactoglobulin, PHOSPHATE ION | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Lazinska, I, Szydlowska, J, Lewinski, K. | | Deposit date: | 2017-04-30 | | Release date: | 2018-04-04 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The engineered beta-lactoglobulin with complementarity to the chlorpromazine chiral conformers.
Int. J. Biol. Macromol., 114, 2018
|
|
5NUJ
 
 | | Engineered beta-lactoglobulin: variant I56F-L39A in complex with chlorpromazine (LG-FA-CLP) | | Descriptor: | 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, Beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Lazinska, I, Szydlowska, J, Lewinski, K. | | Deposit date: | 2017-04-30 | | Release date: | 2018-04-04 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The engineered beta-lactoglobulin with complementarity to the chlorpromazine chiral conformers.
Int. J. Biol. Macromol., 114, 2018
|
|
5NUK
 
 | | Engineered beta-lactoglobulin: variant I56F-L39A-M107F in complex with chlorpromazine (LG-FAF-CLP) | | Descriptor: | 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, Beta-lactoglobulin, CHLORIDE ION, ... | | Authors: | Loch, J.I, Bonarek, P, Tworzydlo, M, Lazinska, I, Szydlowska, J, Lewinski, K. | | Deposit date: | 2017-04-30 | | Release date: | 2018-04-04 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The engineered beta-lactoglobulin with complementarity to the chlorpromazine chiral conformers.
Int. J. Biol. Macromol., 114, 2018
|
|
5NUN
 
 | | Engineered beta-lactoglobulin: variant F105L-L39A-M107F in complex with chlorpromazine (LG-LAF-CLP) | | Descriptor: | 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, Beta-lactoglobulin | | Authors: | Loch, J.I, Bonarek, P, Towrzydlo, M, Lazinskia, I, Szydlowska, J, Lewinski, K. | | Deposit date: | 2017-04-30 | | Release date: | 2018-04-04 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The engineered beta-lactoglobulin with complementarity to the chlorpromazine chiral conformers.
Int. J. Biol. Macromol., 114, 2018
|
|
4Y0S
 
 | | Goat beta-lactoglobulin complex with pramocaine (GLG-PRM) | | Descriptor: | Beta-lactoglobulin, Pramocaine, SULFATE ION | | Authors: | Loch, J.I, Bonarek, P, Polit, A, Jablonski, M, Czub, M, Ye, X, Lewinski, K. | | Deposit date: | 2015-02-06 | | Release date: | 2015-07-01 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | beta-Lactoglobulin interactions with local anaesthetic drugs - Crystallographic and calorimetric studies.
Int.J.Biol.Macromol., 80, 2015
|
|
