6O86
 
 | |
5GL3
 
 | | Crystal structure of TON_0340 in complex with Mg | | Descriptor: | MAGNESIUM ION, Uncharacterized protein | | Authors: | Lee, S.G, Sohn, Y.S, Oh, B.H. | | Deposit date: | 2016-07-07 | | Release date: | 2016-12-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Identification of a Highly Conserved Hypothetical Protein TON_0340 as a Probable Manganese-Dependent Phosphatase.
PLoS ONE, 11, 2016
|
|
5GL4
 
 | | Crystal structure of TON_0340 in complex with Mn | | Descriptor: | MANGANESE (II) ION, Uncharacterized protein | | Authors: | Lee, S.G, Sohn, Y.S, Oh, B.H. | | Deposit date: | 2016-07-08 | | Release date: | 2016-12-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of a Highly Conserved Hypothetical Protein TON_0340 as a Probable Manganese-Dependent Phosphatase.
PLoS ONE, 11, 2016
|
|
5GKX
 
 | | Crystal structure of TON_0340, apo form | | Descriptor: | PHOSPHATE ION, Uncharacterized protein | | Authors: | Lee, S.G, Sohn, Y.S, Oh, B.H. | | Deposit date: | 2016-07-07 | | Release date: | 2016-12-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Identification of a Highly Conserved Hypothetical Protein TON_0340 as a Probable Manganese-Dependent Phosphatase.
PLoS ONE, 11, 2016
|
|
5GL2
 
 | | Crystal structure of TON_0340 in complex with Ca | | Descriptor: | CALCIUM ION, Uncharacterized protein | | Authors: | Lee, S.G, Sohn, Y.S, Oh, B.H. | | Deposit date: | 2016-07-07 | | Release date: | 2016-12-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Identification of a Highly Conserved Hypothetical Protein TON_0340 as a Probable Manganese-Dependent Phosphatase.
PLoS ONE, 11, 2016
|
|
6E1J
 
 | |
4DWZ
 
 | | Crystal Structure of Ton_0340 | | Descriptor: | Hypothetical protein TON_0340, ZINC ION | | Authors: | Lee, S.G, Lee, K.H, Cha, S.S, Oh, B.H. | | Deposit date: | 2012-02-27 | | Release date: | 2013-01-16 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Experimental phasing using zinc anomalous scattering
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4FC5
 
 | | Crystal Structure of Ton_0340 | | Descriptor: | Putative uncharacterized protein, ZINC ION | | Authors: | Lee, S.G, Lee, K.H, An, Y.J, Cha, S.S, Oh, B.H. | | Deposit date: | 2012-05-24 | | Release date: | 2012-09-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Experimental phasing using zinc anomalous scattering
Acta Crystallogr.,Sect.D, 68, 2012
|
|
6WLF
 
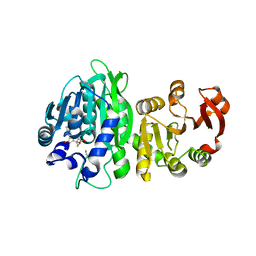 | |
5YI5
 
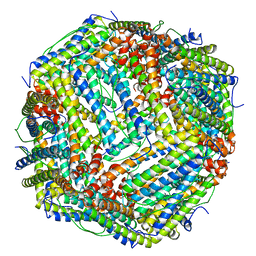 | | human ferritin mutant - E-helix deletion | | Descriptor: | Ferritin heavy chain | | Authors: | Lee, S.G, Yoon, H.R, Ahn, B.J, Jeong, H, Hyun, J, Jung, Y, Kim, H. | | Deposit date: | 2017-10-02 | | Release date: | 2018-02-21 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Four-fold Channel-Nicked Human Ferritin Nanocages for Active Drug Loading and pH-Responsive Drug Release
Angew. Chem. Int. Ed. Engl., 57, 2018
|
|
6X9L
 
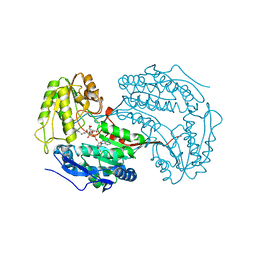 | |
5XB1
 
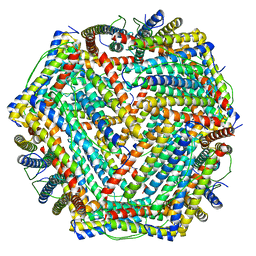 | | human ferritin mutant - E-helix deletion | | Descriptor: | Ferritin heavy chain | | Authors: | Lee, S.G, Ahn, B.J, Jeong, H, Kim, H, Hyun, J, Jung, Y. | | Deposit date: | 2017-03-15 | | Release date: | 2018-02-21 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Four-fold Channel-Nicked Human Ferritin Nanocages for Active Drug Loading and pH-Responsive Drug Release
Angew. Chem. Int. Ed. Engl., 57, 2018
|
|
8VGA
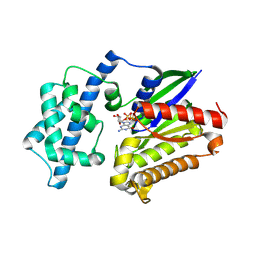 
 | |
8VGB
 
 | |
6O88
 
 | | Crystal Structure of UDP-dependent glucosyltransferases (UGT) from Stevia rebaudiana in complex with UDP and rebaudioside A | | Descriptor: | (8alpha,9beta,10alpha,13alpha)-13-{[alpha-L-allopyranosyl-(1->2)-[beta-D-mannopyranosyl-(1->3)]-beta-D-allopyranosyl]oxy}kauran-18-oic acid, UDP-glycosyltransferase 76G1, URIDINE-5'-DIPHOSPHATE | | Authors: | Lee, S.G, Jez, J.M. | | Deposit date: | 2019-03-09 | | Release date: | 2019-06-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Molecular basis for branched steviol glucoside biosynthesis.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6O87
 
 | |
4FGZ
 
 | | Crystal Structure of Phosphoethanolamine Methyltransferase from Plasmodium falciparum in Complex with Amodiaquine | | Descriptor: | 4-[(7-CHLOROQUINOLIN-4-YL)AMINO]-2-[(DIETHYLAMINO)METHYL]PHENOL, PHOSPHATE ION, Phosphoethanolamine N-methyltransferase, ... | | Authors: | Lee, S.G, Alpert, T.D, Jez, J.M. | | Deposit date: | 2012-06-05 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.994 Å) | | Cite: | Crystal structure of phosphoethanolamine methyltransferase from Plasmodium falciparum in complex with amodiaquine.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
4KRI
 
 | |
4KRG
 
 | |
4KRH
 
 | |
4OMZ
 
 | |
4OMY
 
 | |
4ON0
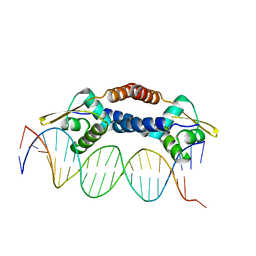 
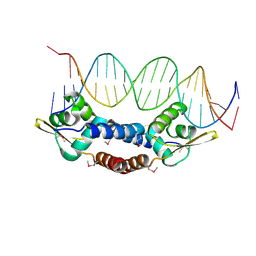
 | | Crystal Structure of NolR from Sinorhizobium fredii in complex with oligo AA DNA | | Descriptor: | DNA (5 -D(*TP*AP*AP*TP*CP*TP*CP*TP*TP*GP*GP*GP*AP*CP*TP*AP*CP*AP*AP*TP*TP*A)-3 ), DNA (5 -D(*TP*AP*TP*TP*AP*GP*AP*GP*AP*AP*CP*CP*CP*TP*GP*AP*TP*GP*TP*TP*AP*A)-3 ), NolR | | Authors: | Lee, S.G, Krishnan, H.B, Jez, J.M. | | Deposit date: | 2014-01-28 | | Release date: | 2014-04-16 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis for regulation of rhizobial nodulation and symbiosis gene expression by the regulatory protein NolR.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
5WP4
 
 | |
6MH4
 
 | |
