3TFH
 
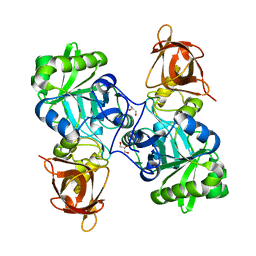 | | DMSP-dependent demethylase from P. ubique - apo | | Descriptor: | GLYCEROL, GcvT-like Aminomethyltransferase protein, SODIUM ION | | Authors: | Schuller, D.J, Reisch, C.R, Moran, M.A, Whitman, W.B, Lanzilotta, W.N. | | Deposit date: | 2011-08-15 | | Release date: | 2011-12-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structures of dimethylsulfoniopropionate-dependent demethylase from the marine organism Pelagabacter ubique.
Protein Sci., 21, 2012
|
|
4KU1
 
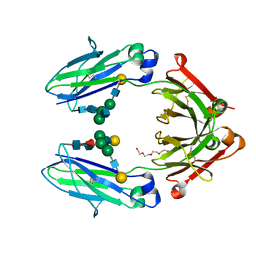 | | Role of the hinge and C-gamma-2/C-gamma-3 interface in immunoglobin G1 Fc domain motions: implications for Fc engineering | | Descriptor: | Ig gamma-1 chain C region, TETRAETHYLENE GLYCOL, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Frank, M, Walker, R, Lanzilotta, W.N, Prestegard, J.H, Barb, A.W. | | Deposit date: | 2013-05-21 | | Release date: | 2014-02-19 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Immunoglobulin g1 fc domain motions: implications for fc engineering.
J.Mol.Biol., 426, 2014
|
|
1YUX
 
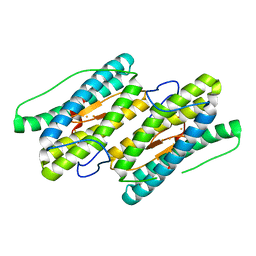 | | Mixed valant state of nigerythrin | | Descriptor: | FE (II) ION, FE (III) ION, Nigerythrin | | Authors: | Iyer, R.B, Silaghi-Dumitrescu, R, Kurtz, D.M, Lanzilotta, W.N. | | Deposit date: | 2005-02-14 | | Release date: | 2005-06-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | High-resolution crystal structures of Desulfovibrio vulgaris (Hildenborough) nigerythrin: facile, redox-dependent iron movement, domain interface variability, and peroxidase activity in the rubrerythrins.
J.Biol.Inorg.Chem., 10, 2005
|
|
1YV1
 
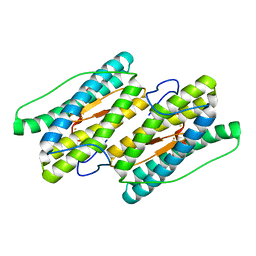 | | Fully reduced state of nigerythrin (all ferrous) | | Descriptor: | FE (II) ION, Nigerythrin | | Authors: | Iyer, R.B, Silaghi-Dumitrescu, R, Kurtz, D.M, Lanzilotta, W.N. | | Deposit date: | 2005-02-14 | | Release date: | 2005-06-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | High-resolution crystal structures of Desulfovibrio vulgaris (Hildenborough) nigerythrin: facile, redox-dependent iron movement, domain interface variability, and peroxidase activity in the rubrerythrins.
J.Biol.Inorg.Chem., 10, 2005
|
|
1YUZ
 
 | | Partially Reduced State of Nigerythrin | | Descriptor: | FE (II) ION, FE (III) ION, Nigerythrin | | Authors: | Iyer, R.B, Silaghi-Dumitrescu, R, Kurtz, D.M, Lanzilotta, W.N. | | Deposit date: | 2005-02-14 | | Release date: | 2005-06-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | High-resolution crystal structures of Desulfovibrio vulgaris (Hildenborough) nigerythrin: facile, redox-dependent iron movement, domain interface variability, and peroxidase activity in the rubrerythrins.
J.Biol.Inorg.Chem., 10, 2005
|
|
4MTJ
 
 | | Structure of the b12-independent glycerol dehydratase with 1,2-propanediol bound | | Descriptor: | B12-independent glycerol dehydratase, S-1,2-PROPANEDIOL | | Authors: | LaMattina, J, Wright, A.V, Demick, J, Soucaille, P, Lanzilotta, W.N. | | Deposit date: | 2013-09-19 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | When Computational Chemistry and Modern Software Get It Right; New Insight Into the Mechanism of a Glycyl Radical Enzyme
To be Published
|
|
