7UF2
 
 | | RibB from Vibrio cholera bound with D-xylulose-5-phosphate (D-Xy5P) and manganese | | Descriptor: | 1,2-ETHANEDIOL, 3,4-dihydroxy-2-butanone 4-phosphate synthase, 5-O-phosphono-D-xylulose, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF5
 
 | | RibB from Vibrio cholera bound with intermediate 2 in the reaction cycle and the products DHBP and formate | | Descriptor: | (2R)-2-(dihydroxymethyl)-2-hydroxy-3-oxobutyl dihydrogen phosphate, (2R)-2-hydroxy-3-oxobutyl dihydrogen phosphate, 3,4-dihydroxy-2-butanone 4-phosphate synthase, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF3
 
 | | RibB from Vibrio cholera bound with L-xylulose-5-phosphate (L-Xy5P) and manganese | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase, L-xylulose-5-phosphate, MANGANESE (II) ION | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UF4
 
 | | RibB from Vibrio cholera bound with intermediate 1 of the reaction cycle and D-ribulose-5-phosphate (D-Ru5P) | | Descriptor: | (2R)-2-hydroxy-3,4-dioxopentyl dihydrogen phosphate, 1,2-ETHANEDIOL, 3,4-dihydroxy-2-butanone 4-phosphate synthase, ... | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
7UEZ
 
 | | Apo RibB from Vibrio cholera | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase | | Authors: | Kenjic, N, Meneely, K.M, Lamb, A.L. | | Deposit date: | 2022-03-22 | | Release date: | 2022-07-20 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Evidence for the Chemical Mechanism of RibB (3,4-Dihydroxy-2-butanone 4-phosphate Synthase) of Riboflavin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
6PBT
 
 | | Pseudopaline Dehydrogenase with (R)-Pseudopaline Soaked 2 hours | | Descriptor: | 1,2-ETHANEDIOL, N-[(3S)-3-amino-3-carboxypropyl]-L-histidine, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | McFarlane, J.S, Lamb, A.L. | | Deposit date: | 2019-06-14 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Staphylopine and pseudopaline dehydrogenase from bacterial pathogens catalyze reversible reactions and produce stereospecific metallophores.
J.Biol.Chem., 294, 2019
|
|
6PBN
 
 | | Pseudopaline Dehydrogenase with (R)-Pseudopaline Soaked 1 hour | | Descriptor: | 1,2-ETHANEDIOL, 2-OXOGLUTARIC ACID, N-[(3S)-3-amino-3-carboxypropyl]-L-histidine, ... | | Authors: | McFarlane, J.S, Lamb, A.L. | | Deposit date: | 2019-06-14 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Staphylopine and pseudopaline dehydrogenase from bacterial pathogens catalyze reversible reactions and produce stereospecific metallophores.
J.Biol.Chem., 294, 2019
|
|
6PBM
 
 | | Pseudopaline Dehydrogenase with NADP+ bound | | Descriptor: | 1,2-ETHANEDIOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Pseudopaline Dehydrogenase | | Authors: | McFarlane, J.S, Lamb, A.L. | | Deposit date: | 2019-06-14 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Staphylopine and pseudopaline dehydrogenase from bacterial pathogens catalyze reversible reactions and produce stereospecific metallophores.
J.Biol.Chem., 294, 2019
|
|
6PBP
 
 | | Pseudopaline Dehydrogenase with (S)-Pseudopaline Soaked 1 hour | | Descriptor: | 1,2-ETHANEDIOL, N-[(1S)-1-carboxy-3-{[(1S)-1-carboxy-2-(1H-imidazol-5-yl)ethyl]amino}propyl]-L-glutamic acid, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | McFarlane, J.S, Lamb, A.L. | | Deposit date: | 2019-06-14 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Staphylopine and pseudopaline dehydrogenase from bacterial pathogens catalyze reversible reactions and produce stereospecific metallophores.
J.Biol.Chem., 294, 2019
|
|
3HGW
 
 | | Apo Structure of Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase I87T mutant | | Descriptor: | CALCIUM ION, Salicylate biosynthesis protein pchB | | Authors: | Luo, Q, Lamb, A.L. | | Deposit date: | 2009-05-14 | | Release date: | 2009-06-30 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure-function analyses of isochorismate-pyruvate lyase from Pseudomonas aeruginosa suggest differing catalytic mechanisms for the two pericyclic reactions of this bifunctional enzyme
Biochemistry, 48, 2009
|
|
3HGX
 
 | |
3HHP
 
 | |
6NN4
 | | The structure of human liver pyruvate kinase, hLPYK-D499N, in complex with Fru-1,6-BP | | Descriptor: | 1,2-ETHANEDIOL, 1,6-di-O-phosphono-beta-D-fructofuranose, PHOSPHOENOLPYRUVATE, ... | | Authors: | McFarlane, J.S, Ronnebaum, T.A, Meneely, K.M, Fenton, A.W, Lamb, A.L. | | Deposit date: | 2019-01-14 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Changes in the allosteric site of human liver pyruvate kinase upon activator binding include the breakage of an intersubunit cation-pi bond.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
6NN5
 
 | | The structure of human liver pyruvate kinase, hLPYK-W527H | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, Pyruvate kinase PKLR | | Authors: | McFarlane, J.S, Ronnebaum, T.A, Meneely, K.M, Fenton, A.W, Lamb, A.L. | | Deposit date: | 2019-01-14 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.256 Å) | | Cite: | Changes in the allosteric site of human liver pyruvate kinase upon activator binding include the breakage of an intersubunit cation-pi bond.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
6NN7
 
 | | The structure of human liver pyruvate kinase, hLPYK-GGG | | Descriptor: | 1,2-ETHANEDIOL, CITRATE ANION, GLYCEROL, ... | | Authors: | McFarlane, J.S, Ronnebaum, T.A, Meneely, K.M, Fenton, A.W, Lamb, A.L. | | Deposit date: | 2019-01-14 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Changes in the allosteric site of human liver pyruvate kinase upon activator binding include the breakage of an intersubunit cation-pi bond.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
6NN8
 
 | | The structure of human liver pyruvate kinase, hLPYK-S531E | | Descriptor: | 1,2-ETHANEDIOL, Pyruvate kinase PKLR | | Authors: | McFarlane, J.S, Ronnebaum, T.A, Meneely, K.M, Fenton, A.W, Lamb, A.L. | | Deposit date: | 2019-01-14 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.416 Å) | | Cite: | Changes in the allosteric site of human liver pyruvate kinase upon activator binding include the breakage of an intersubunit cation-pi bond.
Acta Crystallogr.,Sect.F, 75, 2019
|
|
3REM
 
 | | Structure of the Isochorismate-Pyruvate Lyase from Pseudomonas aerugionsa with Bound Salicylate and Pyruvate | | Descriptor: | 2-HYDROXYBENZOIC ACID, PYRUVIC ACID, Salicylate biosynthesis protein pchB | | Authors: | Olucha, J, Ouellette, A.N, Luo, Q, Lamb, A.L. | | Deposit date: | 2011-04-04 | | Release date: | 2011-07-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | pH Dependence of Catalysis by Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase: Implications for Transition State Stabilization and the Role of Lysine 42.
Biochemistry, 50, 2011
|
|
3RET
 
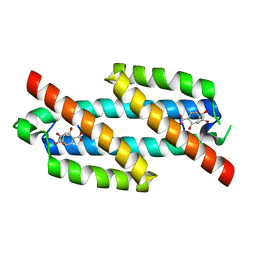 | | Salicylate and Pyruvate Bound Structure of the Isochorismate-Pyruvate Lyase K42E Mutant from Pseudomonas aerugionsa | | Descriptor: | 2-HYDROXYBENZOIC ACID, PYRUVIC ACID, Salicylate biosynthesis protein pchB | | Authors: | Olucha, J, Ouellette, A.N, Luo, Q, Lamb, A.L. | | Deposit date: | 2011-04-05 | | Release date: | 2011-07-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | pH Dependence of Catalysis by Pseudomonas aeruginosa Isochorismate-Pyruvate Lyase: Implications for Transition State Stabilization and the Role of Lysine 42.
Biochemistry, 50, 2011
|
|
6U4S
 
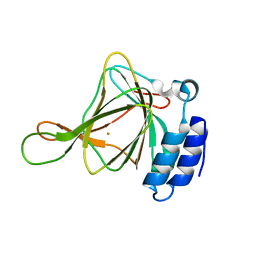 | | wild type cysteine dioxygenase | | Descriptor: | Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Meneely, K.M, Chilton, A.S, Forbes, D.L, Ellis, H.R, Lamb, A.L. | | Deposit date: | 2019-08-26 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | The 3-His Metal Coordination Site Promotes the Coupling of Oxygen Activation to Cysteine Oxidation in Cysteine Dioxygenase.
Biochemistry, 59, 2020
|
|
6U4L
 
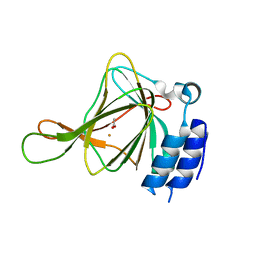 | | cysteine dioxygenase variant - C93E | | Descriptor: | ACETATE ION, Cysteine dioxygenase type 1, FE (III) ION | | Authors: | Meneely, K.M, Chilton, A.S, Forbes, D.L, Ellis, H.R, Lamb, A.L. | | Deposit date: | 2019-08-26 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.911 Å) | | Cite: | The 3-His Metal Coordination Site Promotes the Coupling of Oxygen Activation to Cysteine Oxidation in Cysteine Dioxygenase.
Biochemistry, 59, 2020
|
|
6C4N
 
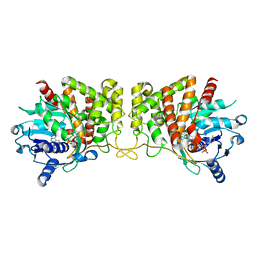 | | Pseudopaline dehydrogenase (PaODH) - NADP+ bound | | Descriptor: | 1,2-ETHANEDIOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Pseudopaline dehydrogenase | | Authors: | McFarlane, J.S, Davis, C.L, Lamb, A.L. | | Deposit date: | 2018-01-12 | | Release date: | 2018-04-11 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Staphylopine, pseudopaline, and yersinopine dehydrogenases: A structural and kinetic analysis of a new functional class of opine dehydrogenase.
J. Biol. Chem., 293, 2018
|
|
6C4L
 
 | | Yersinopine dehydrogenase (YpODH) - Apo | | Descriptor: | Yersinopine dehydrogenase | | Authors: | McFarlane, J.S, Davis, C.L, Lamb, A.L. | | Deposit date: | 2018-01-12 | | Release date: | 2018-04-11 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Staphylopine, pseudopaline, and yersinopine dehydrogenases: A structural and kinetic analysis of a new functional class of opine dehydrogenase.
J. Biol. Chem., 293, 2018
|
|
6C4M
 
 | | Yersinopine dehydrogenase (YpODH) - NADP+ bound | | Descriptor: | NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Yersinopine dehydrogenase | | Authors: | McFarlane, J.S, Davis, C.L, Lamb, A.L. | | Deposit date: | 2018-01-12 | | Release date: | 2018-04-11 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Staphylopine, pseudopaline, and yersinopine dehydrogenases: A structural and kinetic analysis of a new functional class of opine dehydrogenase.
J. Biol. Chem., 293, 2018
|
|
6C4R
 
 | | Staphylopine dehydrogenase (SaODH) - Apo | | Descriptor: | GLYCEROL, SULFATE ION, Staphylopine dehydrogenase | | Authors: | McFarlane, J.S, Davis, C.L, Lamb, A.L. | | Deposit date: | 2018-01-12 | | Release date: | 2018-04-11 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.288 Å) | | Cite: | Staphylopine, pseudopaline, and yersinopine dehydrogenases: A structural and kinetic analysis of a new functional class of opine dehydrogenase.
J. Biol. Chem., 293, 2018
|
|
6CUL
 
 | | PvdF of pyoverdin biosynthesis is a structurally unique N10-formyltetrahydrofolate-dependent formyltransferase | | Descriptor: | CITRIC ACID, N-(4-{[(2-amino-4-oxo-1,4-dihydroquinazolin-6-yl)methyl]amino}benzene-1-carbonyl)-D-glutamic acid, Pyoverdine synthetase F | | Authors: | Kenjic, N, Hoag, M.R, Moraski, G.C, Caperelli, C.A, Moran, G.R, Lamb, A.L. | | Deposit date: | 2018-03-26 | | Release date: | 2019-02-06 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | PvdF of pyoverdin biosynthesis is a structurally unique N10-formyltetrahydrofolate-dependent formyltransferase.
Arch. Biochem. Biophys., 664, 2019
|
|
