7AC5
 
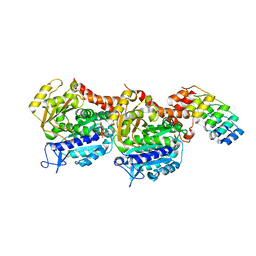 | | Structure of Tubulin Darpin complex 1 collected by rotation serial crystallography on a COC membrane at a synchrotron source | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, Designed Ankyrin Repeat Protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Martiel, I, Olieric, N, Wranik, M, Padeste, C, Karpik, A, Huang, C.Y, Wang, M, Marsh, M. | | Deposit date: | 2020-09-10 | | Release date: | 2021-09-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Versatile microporous polymer-based supports for serial macromolecular crystallography.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
8Q0U
 
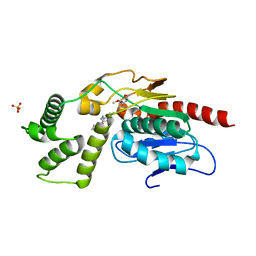 | | Identification and optimisation of novel inhibitors of the Polyketide synthetase 13 thioesterase domain with antitubercular activity | | Descriptor: | Polyketide synthase Pks13, SULFATE ION, ~{N}-[(1~{R})-2-[4-(azetidin-1-ylcarbonyl)phenyl]-1-cyano-ethyl]-3-(3,4-dimethoxyphenyl)-1,2,4-oxadiazole-5-carboxamide | | Authors: | Eadsforth, T.C, Punekar, A.S, Green, S.R, Baragana, B. | | Deposit date: | 2023-07-29 | | Release date: | 2023-11-22 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification and Optimization of Novel Inhibitors of the Polyketide Synthase 13 Thioesterase Domain with Antitubercular Activity.
J.Med.Chem., 66, 2023
|
|
8Q0T
 
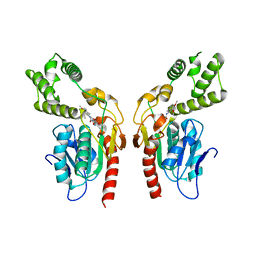 | | Identification and optimisation of novel inhibitors of the Polyketide synthetase 13 thioesterase domain with antitubercular activity | | Descriptor: | 3-(3,4-dimethoxyphenyl)-~{N}-[2-(3,4-dimethoxyphenyl)ethyl]-1,2,4-oxadiazole-5-carboxamide, Polyketide synthase Pks13 | | Authors: | Eadsforth, T.C, Punekar, A.S, Green, S.R, Baragana, B. | | Deposit date: | 2023-07-29 | | Release date: | 2023-11-22 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification and Optimization of Novel Inhibitors of the Polyketide Synthase 13 Thioesterase Domain with Antitubercular Activity.
J.Med.Chem., 66, 2023
|
|
8Q17
 
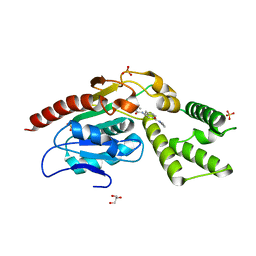 | | Identification and optimisation of novel inhibitors of the Polyketide synthetase 13 thioesterase domain with antitubercular activity | | Descriptor: | (2R)-2-{[(2R)-2-{[(2S)-2-{[(2R)-2-hydroxypropyl]oxy}propyl]oxy}propyl]oxy}propan-1-ol, GLYCEROL, Polyketide synthase Pks13, ... | | Authors: | Eadsforth, T.C, Punekar, A.S, Green, S.R, Baragana, B. | | Deposit date: | 2023-07-30 | | Release date: | 2023-11-22 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Identification and Optimization of Novel Inhibitors of the Polyketide Synthase 13 Thioesterase Domain with Antitubercular Activity.
J.Med.Chem., 66, 2023
|
|
1TU9
 
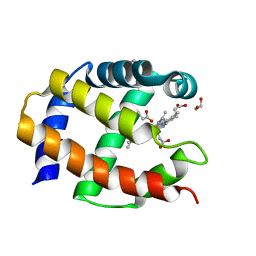 | | Crystal Structure of a Protein PA3967, a Structurally Highly Homologous to a Human Hemoglobin, from Pseudomonas aeruginosa PAO1 | | Descriptor: | 1,2-ETHANEDIOL, PROPANOIC ACID, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Kim, Y, Joachimiak, A, Skarina, T, Egorova, O, Bochkarev, A, Savchenko, A, Edwards, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2004-06-24 | | Release date: | 2004-08-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal Structure of PA3967 from Pseudomonas aeruginosa PAO1, a Hypothetical Protein which is highly homologous to human Hemoglobin in structure.
To be Published
|
|
3K29
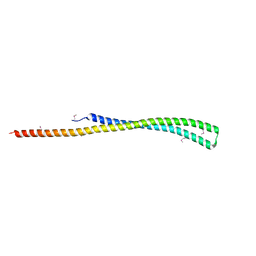 
 | | Structure of a putative YscO homolog CT670 from Chlamydia trachomatis | | Descriptor: | Putative uncharacterized protein | | Authors: | Lam, R, Singer, A, Skarina, T, Onopriyenko, O, Bochkarev, A, Brunzelle, J.S, Edwards, A.M, Anderson, W.F, Chirgadze, N.Y, Savchenko, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2009-09-29 | | Release date: | 2009-10-13 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and protein-protein interaction studies on Chlamydia trachomatis protein CT670 (YscO Homolog).
J.Bacteriol., 192, 2010
|
|
2ITJ
 
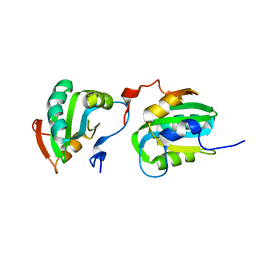 | |
6YVE
 
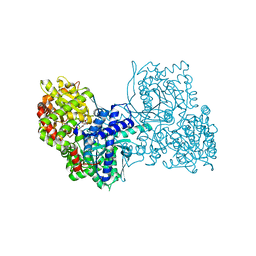 | | Glycogen phosphorylase b in complex with pelargonidin 3-O-beta-D-glucoside | | Descriptor: | DIMETHYL SULFOXIDE, Glycogen phosphorylase, muscle form, ... | | Authors: | Drakou, C.E, Gardeli, C, Tsialtas, I, Alexopoulos, S, Mallouchos, A, Koulas, S, Tsagkarakou, A, Asimakopoulos, D, Leonidas, D.D, Psarra, A.M, Skamnaki, V.T. | | Deposit date: | 2020-04-28 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Affinity Crystallography Reveals Binding of Pomegranate Juice Anthocyanins at the Inhibitor Site of Glycogen Phosphorylase: The Contribution of a Sugar Moiety to Potency and Its Implications to the Binding Mode.
J.Agric.Food Chem., 68, 2020
|
|
1FGU
 
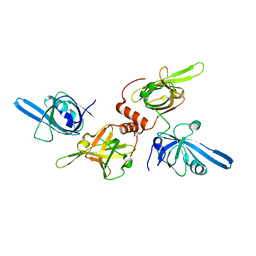 | |
6ON2
 
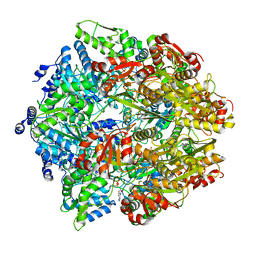 | | Lon Protease from Yersinia pestis with Y2853 substrate | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP-dependent protease La, ... | | Authors: | Shin, M, Asmita, A, Puchades, C, Adjei, E, Wiseman, R.L, Karzai, A.W, Lander, G.C. | | Deposit date: | 2019-04-19 | | Release date: | 2019-05-01 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis for distinct operational modes and protease activation in AAA+ protease Lon.
Sci Adv, 6, 2020
|
|
2ITL
 
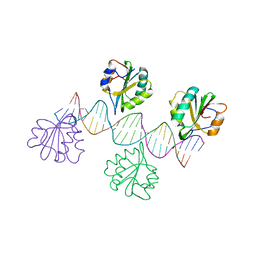 | |
2IPR
 
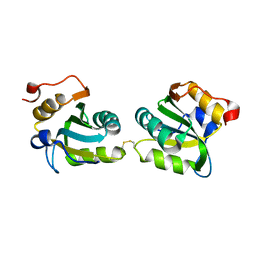 | |
7AC2
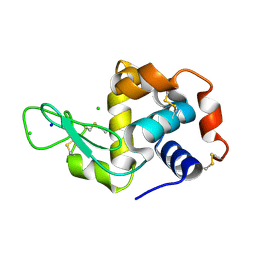 
 | | Structure of Hen Egg White Lysozyme collected by rotation serial crystallography on a COC membrane at a synchrotron source | | Descriptor: | CHLORIDE ION, Lysozyme, SODIUM ION | | Authors: | Martiel, I, Padeste, C, Karpik, A, Huang, C.Y, Vera, L, Wang, M, Marsh, M. | | Deposit date: | 2020-09-09 | | Release date: | 2021-09-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.507 Å) | | Cite: | Versatile microporous polymer-based supports for serial macromolecular crystallography.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
6DK3
 
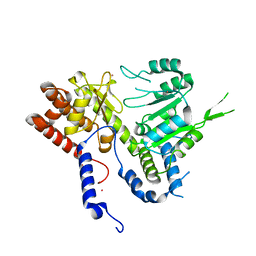 | | HUMAN MITOCHONDRIAL SERINE HYDROXYMETHYLTRANSFERASe 2 | | Descriptor: | Serine hydroxymethyltransferase, mitochondrial, UNKNOWN ATOM OR ION | | Authors: | Dong, A, Wu, H, Zeng, H, Loppnau, P, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Plotnikov, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-05-28 | | Release date: | 2018-06-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | HUMAN MITOCHONDRIAL SERINE HYDROXYMETHYLTRANSFERASe 2
to be published
|
|
5KZS
 
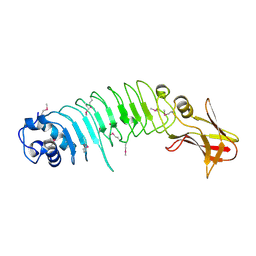 | | Listeria monocytogenes internalin-like protein lmo2027 | | Descriptor: | Putative cell surface protein, similar to internalin proteins | | Authors: | Light, S.H, Nocadello, S, Minasov, G, Cardona-Correa, A, Kwon, K, Faralla, C, Bakardjiev, A, Anderson, W.F, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2016-07-25 | | Release date: | 2017-07-26 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Listeria monocytogenes InlP interacts with afadin and facilitates basement membrane crossing.
Plos Pathog., 14, 2018
|
|
4M8A
 
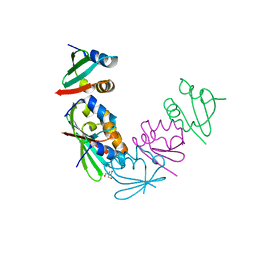 | | Crystal Structure of Thermotoga maritima FtsH Periplasmic Domain | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ATP-dependent zinc metalloprotease FtsH | | Authors: | An, J.Y, Sharif, H, Barrera, F.N, Karabadzhak, A, Kang, G.B, Park, K.J, Sakkiah, S, Lee, K.W, Lee, S, Engelman, D.M, Wang, J, Eom, S.H. | | Deposit date: | 2013-08-13 | | Release date: | 2014-09-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.502 Å) | | Cite: | Structural roles of periplasmic and transmembrane domains of FtsH in ATP-dependent proteolysis
To be Published
|
|
7AC6
 
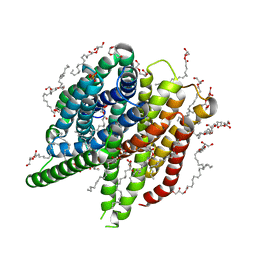 | | Structure of sponge-phase grown PepTst2 collected by rotation serial crystallography on a COC membrane at a synchrotron source | | Descriptor: | (2S)-2,3-DIHYDROXYPROPYL(7Z)-PENTADEC-7-ENOATE, 2-(2-METHOXYETHOXY)ETHANOL, Di-or tripeptide:H+ symporter, ... | | Authors: | Martiel, I, Padeste, C, Karpik, A, Huang, C.Y, Wang, M, Marsh, M. | | Deposit date: | 2020-09-10 | | Release date: | 2021-09-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Versatile microporous polymer-based supports for serial macromolecular crystallography.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
5KGT
 
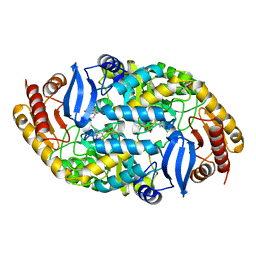 | | Crystal structure of 7,8-diaminopelargonic acid synthase (BioA) from Mycobacterium tuberculosis, complexed with an inhibitor optimized from HTS lead: 1-[4-[4-(3-chlorophenyl)carbonylpiperidin-1-yl]phenyl]ethanone | | Descriptor: | 1-[4-[4-(3-chlorophenyl)carbonylpiperidin-1-yl]phenyl]ethanone, Adenosylmethionine-8-amino-7-oxononanoate aminotransferase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Maize, K.M, Divakaran, A, Aldrich, C.C, Finzel, B.C. | | Deposit date: | 2016-06-13 | | Release date: | 2017-06-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure-Based Optimization of Pyridoxal 5'-Phosphate-Dependent Transaminase Enzyme (BioA) Inhibitors that Target Biotin Biosynthesis in Mycobacterium tuberculosis.
J. Med. Chem., 60, 2017
|
|
5IOX
 
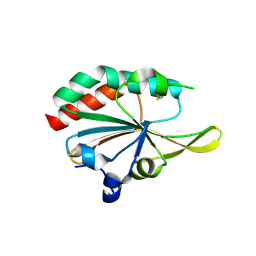 | | Xanthomonas campestris Peroxiredoxin Q - Structure LUss | | Descriptor: | Bacterioferritin comigratory protein | | Authors: | Perkins, A, Parsonage, D, Nelson, K.J, Poole, L.B, Karplus, A. | | Deposit date: | 2016-03-09 | | Release date: | 2016-09-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Peroxiredoxin Catalysis at Atomic Resolution.
Structure, 24, 2016
|
|
2B3G
 
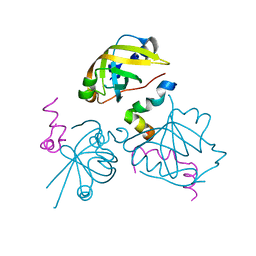 | | p53N (fragment 33-60) bound to RPA70N | | Descriptor: | Cellular tumor antigen p53, Replication protein A 70 kDa DNA-binding subunit | | Authors: | Bochkareva, E, Kaustov, L, Ayed, A, Yi, G.S, Lu, Y, Pineda-Lucena, A, Liao, J.C, Okorokov, A.L, Milner, J, Arrowsmith, C.H, Bochkarev, A. | | Deposit date: | 2005-09-20 | | Release date: | 2005-10-11 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Single-stranded DNA mimicry in the p53 transactivation domain interaction with replication protein A.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
2QPZ
 
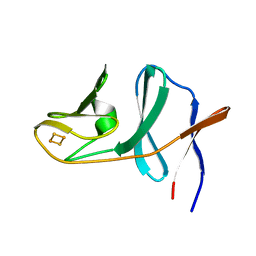 | | Naphthalene 1,2-dioxygenase Rieske ferredoxin | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, Naphthalene 1,2-dioxygenase system ferredoxin subunit | | Authors: | Brown, E.N, Ramaswamy, S, Karlsson, A, Friemann, R, Parales, J.V, Parales, R, Gibson, D.T. | | Deposit date: | 2007-07-25 | | Release date: | 2008-09-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Determining Rieske cluster reduction potentials.
J.Biol.Inorg.Chem., 13, 2008
|
|
5IMV
 
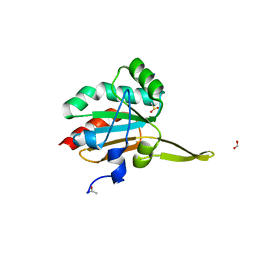 | | Xanthomonas campestris Peroxiredoxin Q - Structure F6 | | Descriptor: | Bacterioferritin comigratory protein, FORMIC ACID, SODIUM ION | | Authors: | Perkins, A, Parsonage, D, Nelson, K.J, Poole, L.B, Karplus, A. | | Deposit date: | 2016-03-07 | | Release date: | 2016-09-21 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Peroxiredoxin Catalysis at Atomic Resolution.
Structure, 24, 2016
|
|
5IMZ
 
 | | Xanthomonas campestris Peroxiredoxin Q - Structure F7 | | Descriptor: | Bacterioferritin comigratory protein, CHLORIDE ION, FORMIC ACID, ... | | Authors: | Perkins, A, Parsonage, D, Nelson, K.J, Poole, L.B, Karplus, A. | | Deposit date: | 2016-03-07 | | Release date: | 2016-09-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Peroxiredoxin Catalysis at Atomic Resolution.
Structure, 24, 2016
|
|
5IO2
 
 | | Xanthomonas campestris Peroxiredoxin Q - C48S mutant | | Descriptor: | Bacterioferritin comigratory protein, PHOSPHATE ION, SODIUM ION | | Authors: | Perkins, A, Parsonage, D, Nelson, K.J, Poole, L.B, Karplus, A. | | Deposit date: | 2016-03-08 | | Release date: | 2016-09-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Peroxiredoxin Catalysis at Atomic Resolution.
Structure, 24, 2016
|
|
5IMA
 
 | | Xanthomonas campestris Peroxiredoxin Q - Structure F2 | | Descriptor: | Bacterioferritin comigratory protein, FORMIC ACID, SODIUM ION | | Authors: | Perkins, A, Parsonage, D, Nelson, K.J, Poole, L.B, Karplus, A. | | Deposit date: | 2016-03-06 | | Release date: | 2016-09-21 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Peroxiredoxin Catalysis at Atomic Resolution.
Structure, 24, 2016
|
|
