5H1Y
 
 | |
5EFX
 
 | | Crystal structure of Rho GTPase regulator | | 分子名称: | Rho guanine nucleotide exchange factor 2 | | 著者 | Jiang, Y, Ouyang, S, Liu, Z.J. | | 登録日 | 2015-10-26 | | 公開日 | 2016-06-29 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.451 Å) | | 主引用文献 | Crystal structure of hGEF-H1 PH domain provides insight into incapability in phosphoinositide binding
Biochem.Biophys.Res.Commun., 471, 2016
|
|
5BOB
 
 | | Crystal Structure of the Meningitis Pathogen Streptococcus suis adhesion Fhb | | 分子名称: | GLYCEROL, Translation initiation factor 2 (IF-2 GTPase) | | 著者 | Jiang, Y, Zhang, C, Yu, Y. | | 登録日 | 2015-05-27 | | 公開日 | 2015-11-18 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Expression, purification, crystallization and structure determination of the N terminal domain of Fhb, a factor H binding protein from Streptococcus suis.
Biochem.Biophys.Res.Commun., 466, 2015
|
|
3K04
 
 | | Crystal Structure of CNG mimicking NaK mutant, NaK-DTPP, Na+ complex | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, Potassium channel protein NaK, SODIUM ION | | 著者 | Jiang, Y, Derebe, M.G. | | 登録日 | 2009-09-24 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.58 Å) | | 主引用文献 | Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3K0D
 
 | | Crystal Structure of CNG mimicking NaK mutant, NaK-ETPP, K+ complex | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, POTASSIUM ION, Potassium channel protein NaK | | 著者 | Jiang, Y, Derebe, M.G. | | 登録日 | 2009-09-24 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3K06
 
 | | Crystal Structure of CNG mimicking NaK mutant, NaK-NTPP, K+ complex | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, POTASSIUM ION, Potassium channel protein NaK | | 著者 | Jiang, Y, Derebe, M.G. | | 登録日 | 2009-09-24 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.58 Å) | | 主引用文献 | Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3K0G
 
 | | Crystal Structure of CNG mimicking NaK mutant, NaK-ETPP, Na+ complex | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, Potassium channel protein NaK, SODIUM ION | | 著者 | Jiang, Y, Derebe, M.G. | | 登録日 | 2009-09-24 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3K08
 
 | | Crystal Structure of CNG mimicking NaK mutant, NaK-NTPP, Na+ complex | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, Potassium channel protein NaK, SODIUM ION | | 著者 | Jiang, Y, Derebe, M.G. | | 登録日 | 2009-09-24 | | 公開日 | 2011-01-12 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (1.62 Å) | | 主引用文献 | Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1LNQ
 
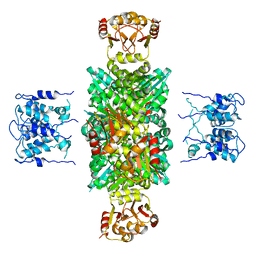 | | CRYSTAL STRUCTURE OF MTHK AT 3.3 A | | 分子名称: | CALCIUM ION, POTASSIUM CHANNEL RELATED PROTEIN | | 著者 | Jiang, Y, Lee, A, Chen, J, Cadene, M, Chait, B.T, Mackinnon, R. | | 登録日 | 2002-05-03 | | 公開日 | 2002-06-19 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | CRYSTAL STRUCTURE AND MECHANISM OF A CALCIUM-GATED POTASSIUM CHANNEL
Nature, 417, 2002
|
|
6AH8
 
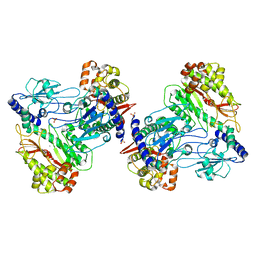 | |
1ORQ
 
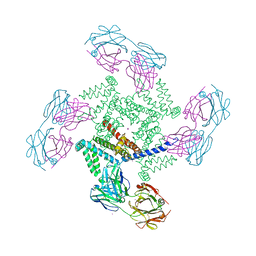 | | X-ray structure of a voltage-dependent potassium channel in complex with an Fab | | 分子名称: | 6E1 Fab heavy chain, 6E1 Fab light chain, CADMIUM ION, ... | | 著者 | Jiang, Y, Lee, A, Chen, J, Ruta, V, Cadene, M, Chait, B.T, MacKinnon, R. | | 登録日 | 2003-03-14 | | 公開日 | 2003-05-06 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | X-ray structure of a voltage-dependent K+ channel
Nature, 423, 2003
|
|
1ORS
 
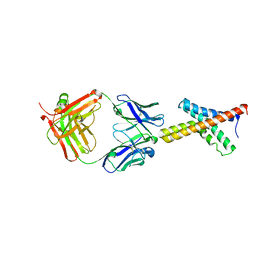 | | X-ray structure of the KvAP potassium channel voltage sensor in complex with an Fab | | 分子名称: | 33H1 Fab heavy chain, 33H1 Fab light chain, potassium channel | | 著者 | Jiang, Y, Lee, A, Chen, J, Ruta, V, Cadene, M, Chait, B.T, MacKinnon, R. | | 登録日 | 2003-03-14 | | 公開日 | 2003-05-06 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | X-ray structure of a voltage-dependent K+ channel
Nature, 423, 2003
|
|
2L5F
 
 | |
6O6J
 
 | |
6O7A
 
 | |
6PPT
 
 | |
6O7C
 
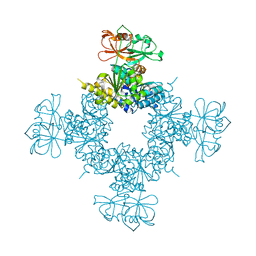 | |
6PQ2
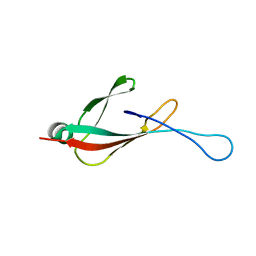 
 | |
6PQM
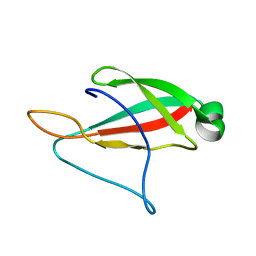 
 | |
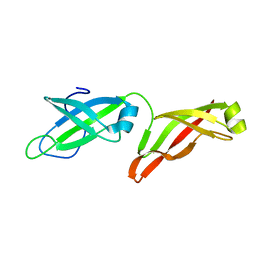
6PRQ
 
 | |
6PRJ
 
 | |
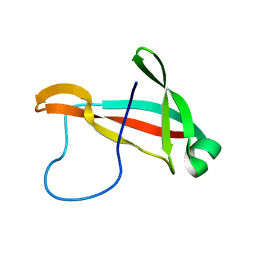
6PRI
 
 | |
6PQE
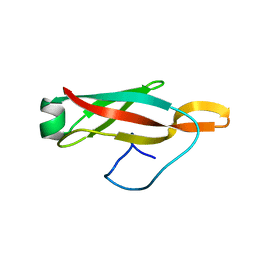 
 | |
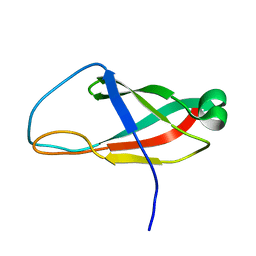
6PRP
 
 | |
6PSI
 
 | |
