5OOU
 
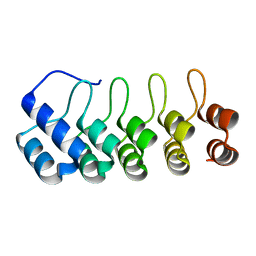 | | Designed Ankyrin Repeat Protein (DARPin) YTRL-1 selected by directed evolution against Lysozyme | | Descriptor: | DARPin YTRL-1 | | Authors: | Fischer, G, Hogan, B.J, Houlihan, G, Edmond, S, Huovinen, T.T.K, Hollfelder, F, Hyvonen, M. | | Deposit date: | 2017-08-08 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.104 Å) | | Cite: | Designed Ankyrin Repeat Protein (DARPin) YTRL-1 selected by directed evolution against Lysozyme
To be published
|
|
5OP1
 
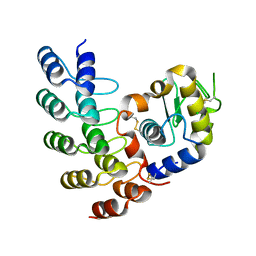 | | Designed Ankyrin Repeat Protein (DARPin) A4 in complex with Lysozyme | | Descriptor: | CHLORIDE ION, DARPin A4, Lysozyme C | | Authors: | Fischer, G, Hogan, B.J, Houlihan, G, Edmond, S, Huovinen, T.T.K, Hollfelder, F, Hyvonen, M. | | Deposit date: | 2017-08-09 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.284 Å) | | Cite: | Designed Ankyrin Repeat Protein (DARPin) A4 in complex with Lysozyme
To be published
|
|
5QUE
 
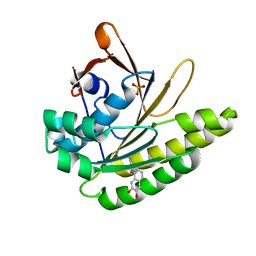 | |
5QUF
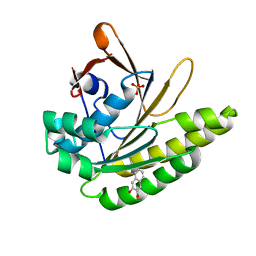 
 | |
5QUB
 
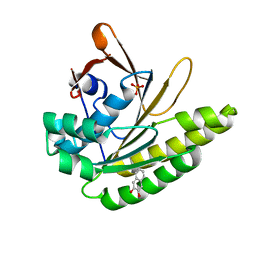 | |
5QUC
 
 | |
5QUH
 
 | |
5QUD
 
 | |
5QUG
 
 | |
5QUO
 
 | |
5QUI
 
 | |
5QUQ
 
 | |
5QUP
 
 | |
5QUJ
 
 | |
5QUK
 
 | | Structure of unliganded HumRadA1.2 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUN
 
 | | Structure of unliganded HumRadA1.6 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUM
 
 | | Structure of unliganded HumRadA1.4 | | Descriptor: | RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUL
 
 | | Structure of unliganded HumRadA1.3 | | Descriptor: | PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.278 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUT
 
 | |
5QUS
 
 | | HumRadA1 soaked with 5 mM indazole in 10% DMSO | | Descriptor: | 1H-indazole, DIMETHYL SULFOXIDE, PHOSPHATE ION, ... | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.247 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUU
 
 | | HumRadA1 as soaking control with 10% DMSO | | Descriptor: | DIMETHYL SULFOXIDE, PHOSPHATE ION, RadA | | Authors: | Marsh, M, Hyvonen, M. | | Deposit date: | 2020-01-27 | | Release date: | 2021-03-03 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.248 Å) | | Cite: | Optimisation of crystal forms for structure-guided drug discovery
To be published
|
|
5QUR
 
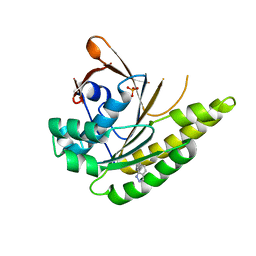 | |
5OOV
 
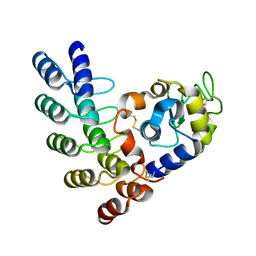 | | Designed Ankyrin Repeat Protein (DARPin) ETVD-1 in complex with Lysozyme | | Descriptor: | DARPin ETVD-1, Lysozyme C | | Authors: | Houlihan, G, Fischer, G, Hogan, B.J, Edmond, S, Huovinen, T.T.K, Hollfelder, F, Hyvonen, M. | | Deposit date: | 2017-08-08 | | Release date: | 2018-08-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.365 Å) | | Cite: | Designed Ankyrin Repeat Protein (DARPin) ETVD-1 in complex with Lysozyme
To be published
|
|
5OOS
 
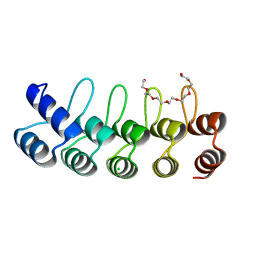 | | Designed Ankyrin Repeat Protein (DARPin) YTRL-2 selected by directed evolution against Lysozyme | | Descriptor: | CHLORIDE ION, DARPin YTRL-2, NONAETHYLENE GLYCOL | | Authors: | Hogan, B.J, Fischer, G, Houlihan, G, Edmond, S, Huovinen, T.T.K, Hollfelder, F, Hyvonen, M. | | Deposit date: | 2017-08-08 | | Release date: | 2018-08-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Designed Ankyrin Repeat Protein (DARPin) YTRL-2 selected by directed evolution against Lysozyme
To be published
|
|
4E9D
 
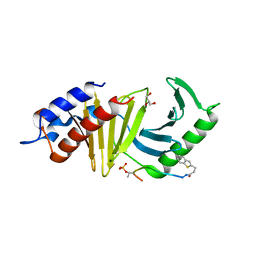 | | The structure of the polo-box domain (PBD) of polo-like kinase 1 (Plk1) in complex with 3-(1-benzothiophen-2-yl)propanoyl-derivatized DPPLHSpTA peptide | | Descriptor: | 3-(1-benzothiophen-2-yl)propanoyl-derivatized DPPLHSpTA peptide, GLYCEROL, Serine/threonine-protein kinase PLK1 | | Authors: | Sledz, P, Hyvonen, M, Lang, S, Stubbs, C.J, Abell, C. | | Deposit date: | 2012-03-21 | | Release date: | 2012-10-10 | | Last modified: | 2012-12-12 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | High-throughput interrogation of ligand binding mode using a fluorescence-based assay.
Angew.Chem.Int.Ed.Engl., 51, 2012
|
|
