4TS6
 | |
3I4O
 
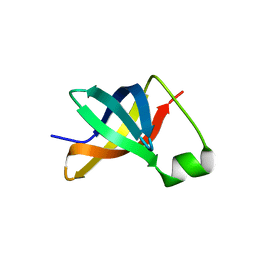 | |
6ZZG
 
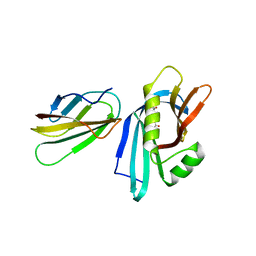 | | MB_CRS6-1 bound to CrSAS-6_N | | Descriptor: | ACETATE ION, Centriole protein, MB_CRS6-15 | | Authors: | Hatzopoulos, G.N, Kukenshoner, T, Banterle, N, Favez, T, Fluckiger, I, Hantschel, O, Gonczy, P. | | Deposit date: | 2020-08-04 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.93 Å) | | Cite: | Tuning SAS-6 architecture with monobodies impairs distinct steps of centriole assembly.
Nat Commun, 12, 2021
|
|
6ZZD
 
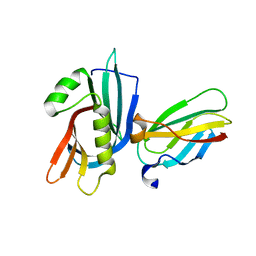 | | MB_CRS6-13 bound to CrSAS-6_N | | Descriptor: | Centriole protein, MB_CRS6-13 | | Authors: | Hatzopoulos, G.N, Kukenshoner, T, Banterle, N, Favez, T, Fluckiger, I, Hantschel, O, Gonczy, P. | | Deposit date: | 2020-08-04 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Tuning SAS-6 architecture with monobodies impairs distinct steps of centriole assembly.
Nat Commun, 12, 2021
|
|
6ZZC
 
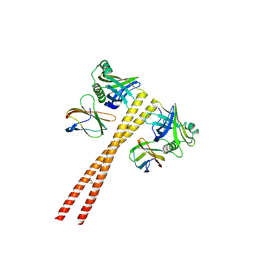 | | MB_CRS6-1 bound to CrSAS-6_6HR | | Descriptor: | Centriole protein, DODECAETHYLENE GLYCOL, MB_CrS6-1 | | Authors: | Hatzopoulos, G.N, Kukenshoner, T, Banterle, N, Favez, T, Fluckiger, I, Hantschel, O, Gonczy, P. | | Deposit date: | 2020-08-04 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.93 Å) | | Cite: | Tuning SAS-6 architecture with monobodies impairs distinct steps of centriole assembly.
Nat Commun, 12, 2021
|
|
6ZZ8
 
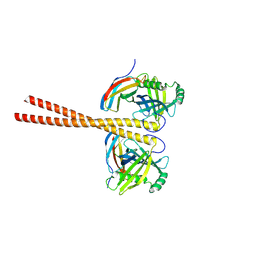 | | MB_CRS6-15 bound to CrSAS-6_6HR | | Descriptor: | Centriole protein, Protein B | | Authors: | Hatzopoulos, G.N, Kukenshoner, T, Banterle, N, Favez, T, Fluckiger, I, Hantschel, O, Gonczy, P. | | Deposit date: | 2020-08-04 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.73 Å) | | Cite: | Tuning SAS-6 architecture with monobodies impairs distinct steps of centriole assembly.
Nat Commun, 12, 2021
|
|
4LD3
 
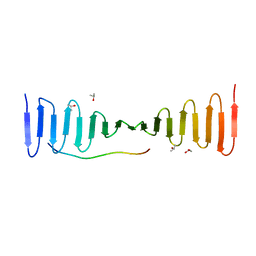 | |
4LD1
 | |
4LZF
 
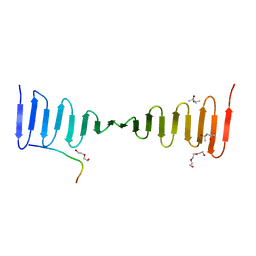 | | A novel domain in the microcephaly protein CPAP suggests a role in centriole architecture | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Centrosomal P4.1-associated protein, SCL-interrupting locus protein homolog, ... | | Authors: | Hatzopoulos, G.N, Erat, M.C, Cutts, E, Rogala, K, Slatter, L, Stansfeld, P.J, Vakonakis, I. | | Deposit date: | 2013-07-31 | | Release date: | 2013-09-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Structural analysis of the G-box domain of the microcephaly protein CPAP suggests a role in centriole architecture.
Structure, 21, 2013
|
|
1Z6S
 
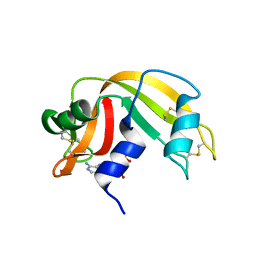 | | Ribonuclease A- AMP complex | | Descriptor: | ADENOSINE MONOPHOSPHATE, Ribonuclease pancreatic | | Authors: | Hatzopoulos, G.N, Leonidas, D.D, Kardakaris, R, Kobe, J, Oikonomakos, N.G. | | Deposit date: | 2005-03-23 | | Release date: | 2005-08-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The binding of IMP to Ribonuclease A
Febs J., 272, 2005
|
|
1Z6D
 
 | | Ribonuclease A- IMP complex | | Descriptor: | INOSINIC ACID, Ribonuclease pancreatic | | Authors: | Hatzopoulos, G.N, Leonidas, D.D, Kardakaris, R, Kobe, J, Oikonomakos, N.G. | | Deposit date: | 2005-03-22 | | Release date: | 2005-08-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | The binding of IMP to Ribonuclease A
Febs J., 272, 2005
|
|
4YNH
 
 | |
4YV4
 
 | |
