7RKH
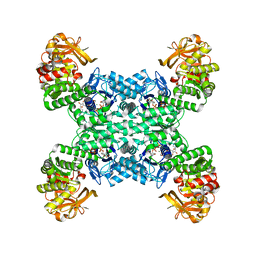 
 | | Yeast CTP Synthase (URA8) tetramer bound to ATP/UTP at neutral pH | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CTP synthase, MAGNESIUM ION, ... | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-22 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (2.8 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RMF
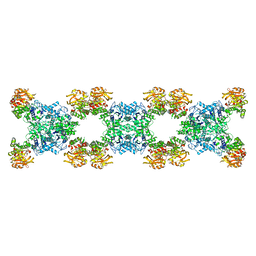 
 | | Substrate-bound Ura7 filament at low pH | | 分子名称: | CTP synthase | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-27 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (7.3 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RMC
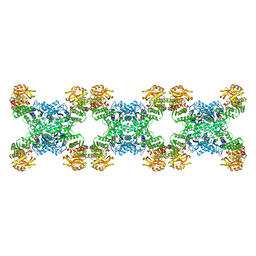 
 | | Yeast CTP Synthase (Ura7) filament bound to CTP at low pH | | 分子名称: | CTP synthase 1, CYTIDINE-5'-TRIPHOSPHATE | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-27 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RMV
 
 | | Yeast CTP Synthase (Ura7) H360R Filament bound to Substrates | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CTP synthase, URIDINE 5'-TRIPHOSPHATE | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-28 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (6.7 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RL5
 
 | | Yeast CTP Synthase (URA8) filament bound to CTP at low pH | | 分子名称: | CTP synthase, CYTIDINE-5'-TRIPHOSPHATE | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-23 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (3.8 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RMK
 
 | | Yeast CTP Synthase (Ura7) Bundle bound to substrates at low pH | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CTP synthase, URIDINE 5'-TRIPHOSPHATE | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-27 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (6.6 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RMO
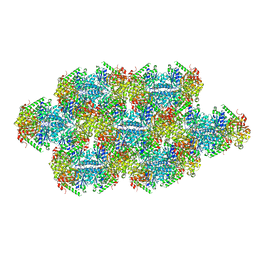 
 | | Yeast CTP Synthase (Ura7) Bundle bound to Products at low pH | | 分子名称: | CTP synthase, CYTIDINE-5'-TRIPHOSPHATE | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-27 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (7 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RNR
 
 | | Yeast CTP Synthase (Ura8) Bundle Bound to Substrates at Low pH | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CTP synthase, MAGNESIUM ION, ... | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-29 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RL0
 | | Yeast CTP Synthase (URA8) Filament bound to ATP/UTP at low pH | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, CTP synthase, MAGNESIUM ION, ... | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-22 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (2.8 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
7RNL
 | | Yeast CTP Synthase (Ura7) H360R Filament bound to Substrates | | 分子名称: | CTP synthase, CYTIDINE-5'-TRIPHOSPHATE, MAGNESIUM ION | | 著者 | Hansen, J.M, Lynch, E.M, Farrell, D.P, DiMaio, F, Quispe, J, Kollman, J.M. | | 登録日 | 2021-07-29 | | 公開日 | 2021-11-24 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation.
Elife, 10, 2021
|
|
1K8A
 
 | | Co-crystal structure of Carbomycin A bound to the 50S ribosomal subunit of Haloarcula marismortui | | 分子名称: | 23S RRNA, 5S RRNA, CADMIUM ION, ... | | 著者 | Hansen, J.L, Ippolito, J.A, Ban, N, Nissen, P, Moore, P.B, Steitz, T. | | 登録日 | 2001-10-23 | | 公開日 | 2002-07-19 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | The structures of four macrolide antibiotics bound to the large ribosomal subunit.
Mol.Cell, 10, 2002
|
|
1KC8
 | | Co-crystal Structure of Blasticidin S Bound to the 50S Ribosomal Subunit | | 分子名称: | 23S RRNA, 5S RRNA, BLASTICIDIN S, ... | | 著者 | Hansen, J.L, Ban, N, Nissen, P, Moore, P.B, Steitz, T.A. | | 登録日 | 2001-11-07 | | 公開日 | 2003-07-22 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3.01 Å) | | 主引用文献 | Structures of Five Antibiotics Bound at the Peptidyl Transferase Center of
the Large Ribosomal Subunit
J.Mol.Biol., 330, 2003
|
|
1K73
 | | Co-crystal Structure of Anisomycin Bound to the 50S Ribosomal Subunit | | 分子名称: | 23S RRNA, 5S RRNA, ANISOMYCIN, ... | | 著者 | Hansen, J, Ban, N, Nissen, P, Moore, P.B, Steitz, T.A. | | 登録日 | 2001-10-18 | | 公開日 | 2003-07-22 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3.01 Å) | | 主引用文献 | Structures of Five Antibiotics Bound at the Peptidyl Transferase Center of
the Large Ribosomal Subunit
J.Mol.Biol., 330, 2003
|
|
1N8R
 | | Structure of large ribosomal subunit in complex with virginiamycin M | | 分子名称: | 23S ribosomal RNA, 50S ribosomal protein L10e, 50S ribosomal protein L13P, ... | | 著者 | Hansen, J.L, Moore, P.B, Steitz, T.A. | | 登録日 | 2002-11-21 | | 公開日 | 2003-07-22 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Structures of Five Antibiotics Bound at the Peptidyl Transferase Center of
the Large Ribosomal Subunit
J.Mol.Biol., 330, 2003
|
|
1NJI
 | | Structure of chloramphenicol bound to the 50S ribosomal subunit | | 分子名称: | 23S ribosomal RNA, 50S ribosomal protein L10e, 50S ribosomal protein L13P, ... | | 著者 | Hansen, J.L, Moore, P.B, Steitz, T.A. | | 登録日 | 2002-12-31 | | 公開日 | 2003-07-22 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Structures of Five Antibiotics Bound at the Peptidyl Transferase Center of
the Large Ribosomal Subunit
J.Mol.Biol., 330, 2003
|
|
1K9M
 
 | | Co-crystal structure of tylosin bound to the 50S ribosomal subunit of Haloarcula marismortui | | 分子名称: | 23S RRNA, 5S RRNA, CADMIUM ION, ... | | 著者 | Hansen, J.L, Ippolito, J.A, Ban, N, Nissen, P, Moore, P.B, Steitz, T.A. | | 登録日 | 2001-10-29 | | 公開日 | 2002-07-19 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | The structures of four macrolide antibiotics bound to the large ribosomal subunit.
Mol.Cell, 10, 2002
|
|
1KD1
 | | Co-crystal Structure of Spiramycin bound to the 50S Ribosomal Subunit of Haloarcula marismortui | | 分子名称: | 23S RRNA, 5S RRNA, CADMIUM ION, ... | | 著者 | Hansen, J.L, Ippolito, J.A, Ban, N, Nissen, P, Moore, P.B, Steitz, T.A. | | 登録日 | 2001-11-12 | | 公開日 | 2002-07-19 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | The structures of four macrolide antibiotics bound to the large ribosomal subunit.
Mol.Cell, 10, 2002
|
|
1M1K
 
 | | Co-crystal structure of azithromycin bound to the 50S ribosomal subunit of Haloarcula marismortui | | 分子名称: | 23S RRNA, 5S RRNA, AZITHROMYCIN, ... | | 著者 | Hansen, J.L, Ippolito, J.A, Ban, N, Nissen, P, Moore, P.B, Steitz, T.A. | | 登録日 | 2002-06-19 | | 公開日 | 2002-07-19 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | The structures of four macrolide antibiotics bound to the large ribosomal subunit.
Mol.Cell, 10, 2002
|
|
1RDR
 
 | |
1Q86
 | | Crystal structure of CCA-Phe-cap-biotin bound simultaneously at half occupancy to both the A-site and P-site of the the 50S ribosomal Subunit. | | 分子名称: | 23S ribosomal rna, 50S ribosomal protein L13P, 50S ribosomal protein L14P, ... | | 著者 | Hansen, J.L, Schmeing, T.M, Moore, P.B, Steitz, T.A. | | 登録日 | 2003-08-20 | | 公開日 | 2003-10-07 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Structural insights into peptide bond formation.
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1Q7Y
 | | Crystal Structure of CCdAP-Puromycin bound at the Peptidyl transferase center of the 50S ribosomal subunit | | 分子名称: | 23S ribosomal rna, 50S ribosomal protein L13P, 50S ribosomal protein L14P, ... | | 著者 | Hansen, J.L, Schmeing, T.M, Moore, P.B, Steitz, T.A. | | 登録日 | 2003-08-20 | | 公開日 | 2003-10-07 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Structural Insights Into Peptide Bond Formation
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1Q82
 | | Crystal Structure of CC-Puromycin bound to the A-site of the 50S ribosomal subunit | | 分子名称: | 23S ribosomal rna, 50S ribosomal protein L13P, 50S ribosomal protein L14P, ... | | 著者 | Hansen, J.L, Schmeing, T.M, Moore, P.B, Steitz, T.A. | | 登録日 | 2003-08-20 | | 公開日 | 2003-10-07 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.98 Å) | | 主引用文献 | Structural Insights Into Peptide Bond Formation
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1Q81
 | | Crystal Structure of minihelix with 3' puromycin bound to A-site of the 50S ribosomal subunit. | | 分子名称: | 23S ribosomal rna, 50S ribosomal protein L13P, 50S ribosomal protein L14P, ... | | 著者 | Hansen, J.L, Schmeing, T.M, Moore, P.B, Steitz, T.A. | | 登録日 | 2003-08-20 | | 公開日 | 2003-10-07 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.95 Å) | | 主引用文献 | Structural Insights into Peptide Bond Formation
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1M90
 | | Co-crystal structure of CCA-Phe-caproic acid-biotin and sparsomycin bound to the 50S ribosomal subunit | | 分子名称: | 23S RRNA, 5S RRNA, 6-AMINOHEXANOIC ACID, ... | | 著者 | Hansen, J.L, Schmeing, T.M, Moore, P.B, Steitz, T.A. | | 登録日 | 2002-07-26 | | 公開日 | 2002-09-06 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structural insights into peptide bond formation.
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
4DTE
 
 | |
