4BUJ
 
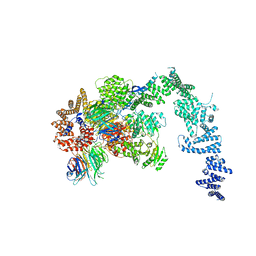 | | Crystal structure of the S. cerevisiae Ski2-3-8 complex | | Descriptor: | ANTIVIRAL HELICASE SKI2, ANTIVIRAL PROTEIN SKI8, SULFATE ION, ... | | Authors: | Halbach, F, Reichelt, P, Rode, M, Conti, E. | | Deposit date: | 2013-06-20 | | Release date: | 2013-08-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | The Yeast Ski Complex: Crystal Structure and RNA Channeling to the Exosome Complex.
Cell(Cambridge,Mass.), 154, 2013
|
|
4A4K
 
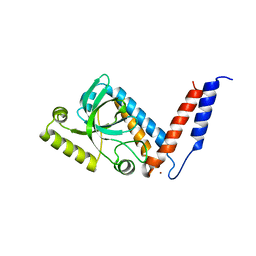 | |
4A4Z
 
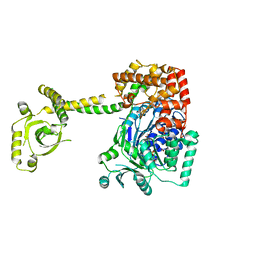 | | CRYSTAL STRUCTURE OF THE S. CEREVISIAE DEXH HELICASE SKI2 BOUND TO AMPPNP | | Descriptor: | 1,2-ETHANEDIOL, ANTIVIRAL HELICASE SKI2, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Halbach, F, Rode, M, Conti, E. | | Deposit date: | 2011-10-20 | | Release date: | 2011-12-21 | | Last modified: | 2019-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Crystal Structure of S. Cerevisiae Ski2, a Dexh Helicase Associated with the Cytoplasmic Functions of the Exosome.
RNA, 18, 2012
|
|
4KHA
 
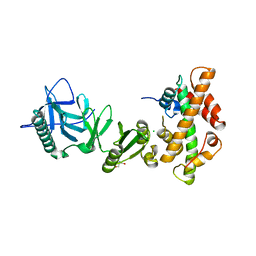 | | Structural basis of histone H2A-H2B recognition by the essential chaperone FACT | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, Histone H2A, ... | | Authors: | Hondele, M, Halbach, F, Hassler, M, Ladurner, A.G. | | Deposit date: | 2013-04-30 | | Release date: | 2013-05-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural basis of histone H2A-H2B recognition by the essential chaperone FACT.
Nature, 499, 2013
|
|
