1YP4
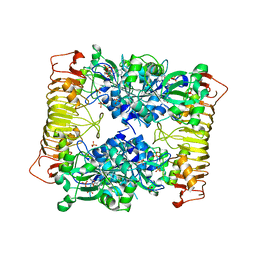 
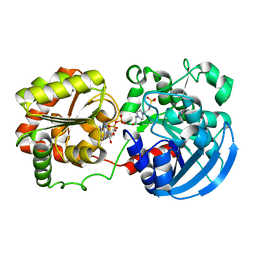
 | | Crystal structure of potato tuber ADP-glucose pyrophosphorylase in complex with ADP-glucose | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE-GLUCOSE, Glucose-1-phosphate adenylyltransferase small subunit, ... | | Authors: | Jin, X, Ballicora, M.A, Preiss, J, Geiger, J.H. | | Deposit date: | 2005-01-29 | | Release date: | 2005-03-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of potato tuber ADP-glucose pyrophosphorylase.
Embo J., 24, 2005
|
|
3CR6
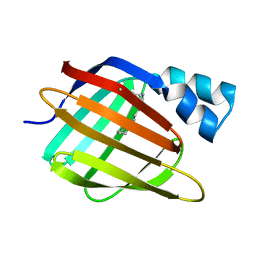 
 | |
7LHJ
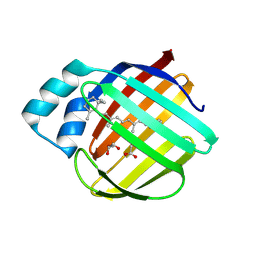 
 | |
3NZ4
 
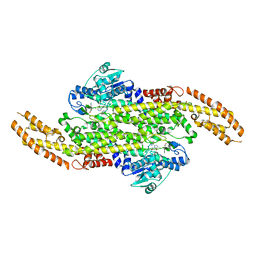 | | Crystal Structure of a Taxus Phenylalanine Aminomutase | | Descriptor: | PHENYLETHYLENECARBOXYLIC ACID, Phenylalanine ammonia-lyase | | Authors: | Feng, L, Geiger, J.H. | | Deposit date: | 2010-07-16 | | Release date: | 2011-03-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Mechanistic, mutational, and structural evaluation of a taxus phenylalanine aminomutase.
Biochemistry, 50, 2011
|
|
4ZH6
 
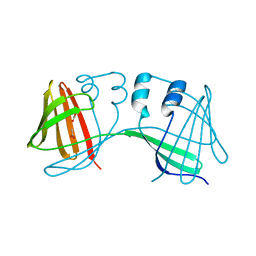 | |
4ZH9
 
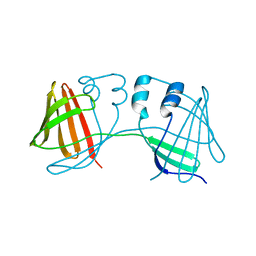 | |
7LHN
 | |
7LHM
 | |
4ZR2
 
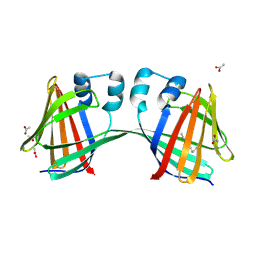 | |
7LHO
 | |
7MS1
 
 | |
7MS0
 
 | | Crystal structure of native Cg10062 | | Descriptor: | 4-oxalocrotonate tautomerase | | Authors: | Nayebi, G.H, Geiger, J.H, Draths, K. | | Deposit date: | 2021-05-10 | | Release date: | 2022-02-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Cg10062 Catalysis Forges a Link between Acetylenecarboxylic Acid and Bacterial Metabolism.
Biochemistry, 60, 2021
|
|
7MS8
 
 | |
7MS3
 
 | |
3F8A
 
 | | Crystal Structure of the R132K:R111L:L121E:R59W Mutant of Cellular Retinoic Acid-Binding Protein Type II Complexed with C15-aldehyde (a retinal analog) at 1.95 Angstrom resolution. | | Descriptor: | 1,3,3-trimethyl-2-[(1E,3E)-3-methylpenta-1,3-dien-1-yl]cyclohexene, 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Cellular retinoic acid-binding protein 2 | | Authors: | Jia, X, Geiger, J.H. | | Deposit date: | 2008-11-12 | | Release date: | 2009-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Probing Wavelength Regulation with an Engineered Rhodopsin Mimic and a C15-Retinal Analogue
Chempluschem, 77, 2012
|
|
3FA6
 | |
3F9D
 
 | |
3CWK
 
 | |
3D1J
 
 | |
3CX4
 
 | |
3COP
 
 | |
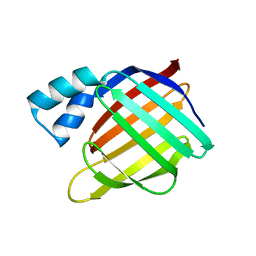
3D96
 
 | |
3D95
 | |
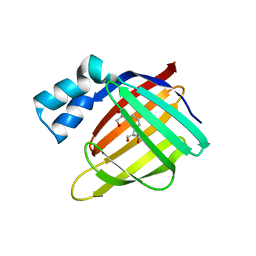
3D97
 
 | |
4GKC
 | |
