1EH8
 | |
1EH7
 | |
1EH6
 
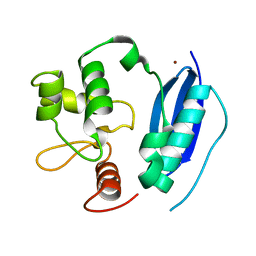 | | HUMAN O6-ALKYLGUANINE-DNA ALKYLTRANSFERASE | | Descriptor: | O6-ALKYLGUANINE-DNA ALKYLTRANSFERASE, ZINC ION | | Authors: | Daniels, D.S, Tainer, J.A. | | Deposit date: | 2000-02-18 | | Release date: | 2000-04-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Active and alkylated human AGT structures: a novel zinc site, inhibitor and extrahelical base binding.
EMBO J., 19, 2000
|
|
1T39
 | | HUMAN O6-ALKYLGUANINE-DNA ALKYLTRANSFERASE COVALENTLY CROSSLINKED TO DNA | | Descriptor: | 5'-D(*GP*CP*CP*AP*TP*GP*(E1X)P*CP*TP*AP*GP*TP*A)-3', 5'-D(*TP*AP*CP*TP*AP*GP*CP*CP*AP*TP*GP*GP*C)-3', Methylated-DNA--protein-cysteine methyltransferase | | Authors: | Daniels, D.S, Woo, T.T, Luu, K.X, Noll, D.M, Clarke, N.D, Pegg, A.E, Tainer, J.A. | | Deposit date: | 2004-04-25 | | Release date: | 2004-07-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | DNA binding and nucleotide flipping by the human DNA repair protein AGT.
Nat.Struct.Mol.Biol., 11, 2004
|
|
1T38
 
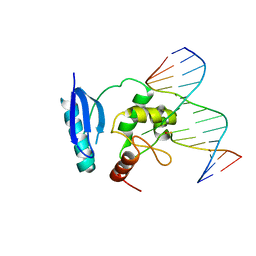 | | HUMAN O6-ALKYLGUANINE-DNA ALKYLTRANSFERASE BOUND TO DNA CONTAINING O6-METHYLGUANINE | | Descriptor: | 5'-D(*GP*CP*CP*AP*TP*GP*(6OG)P*CP*TP*AP*GP*TP*A)-3', 5'-D(*TP*AP*CP*TP*AP*GP*CP*CP*AP*TP*GP*GP*C)-3', Methylated-DNA--protein-cysteine methyltransferase | | Authors: | Daniels, D.S, Woo, T.T, Luu, K.X, Noll, D.M, Clarke, N.D, Pegg, A.E, Tainer, J.A. | | Deposit date: | 2004-04-25 | | Release date: | 2004-07-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | DNA binding and nucleotide flipping by the human DNA repair protein AGT.
Nat.Struct.Mol.Biol., 11, 2004
|
|
5AD2
 
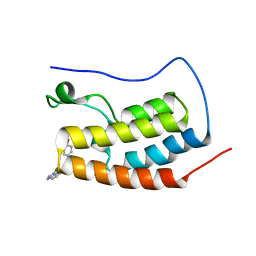 | | Bivalent binding to BET bromodomains | | Descriptor: | (3R)-4-(2-{4-[1-(3-chloro[1,2,4]triazolo[4,3-b]pyridazin-6-yl)-4-piperidinyl]phenoxy}ethyl)-1,3-dimethyl-2-piperazinone, BROMODOMAIN-CONTAINING PROTEIN 4 | | Authors: | Waring, M.J, Chen, H, Rabow, A.A, Walker, G, Bobby, R, Boiko, S, Bradbury, R.H, Callis, R, Dale, I, Daniels, D, Flavell, L, Holdgate, G, Jowitt, T.A, Kikhney, A, McAlister, M, Ogg, D, Patel, J, Petteruti, P, Robb, G.R, Robers, M, Stratton, N, Svergun, D.I, Wang, W, Whittaker, D. | | Deposit date: | 2015-08-19 | | Release date: | 2016-09-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Potent and Selective Bivalent Inhibitors of Bet Bromodomains
Nat.Chem.Biol., 12, 2016
|
|
5AD3
 
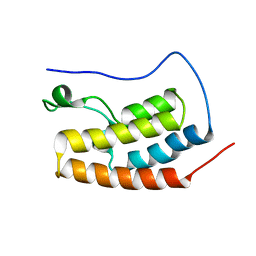 | | Bivalent binding to BET bromodomains | | Descriptor: | 3-methoxy-N-[2-[4-[1-(3-methoxy-[1,2,4]triazolo[4,3-b]pyridazin-6-yl)-4-piperidyl]phenoxy]ethyl]-N-methyl-[1,2,4]triazolo[4,3-b]pyridazin-6-amine, BROMODOMAIN-CONTAINING PROTEIN 4 | | Authors: | Waring, M.J, Chen, H, Rabow, A.A, Walker, G, Bobby, R, Boiko, S, Bradbury, R.H, Callis, R, Dale, I, Daniels, D, Flavell, L, Holdgate, G, Jowitt, T.A, Kikhney, A, McAlister, M, Ogg, D, Patel, J, Petteruti, P, Robb, G.R, Robers, M, Stratton, N, Svergun, D.I, Wang, W, Whittaker, D. | | Deposit date: | 2015-08-19 | | Release date: | 2016-09-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Potent and Selective Bivalent Inhibitors of Bet Bromodomains
Nat.Chem.Biol., 12, 2016
|
|
1M9A
 
 | | Crystal structure of the 26 kDa glutathione S-transferase from Schistosoma japonicum complexed with S-hexylglutathione | | Descriptor: | Glutathione S-Transferase 26 kDa, S-HEXYLGLUTATHIONE | | Authors: | Cardoso, R.M.F, Daniels, D.S, Bruns, C.M, Tainer, J.A. | | Deposit date: | 2002-07-28 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Characterization of the electrophile
binding site and substrate binding
mode of the 26-kDa glutathione
S-transferase from Schistosoma
japonicum
PROTEINS: STRUCT.,FUNCT.,GENET., 51, 2003
|
|
1M9B
 
 | | Crystal structure of the 26 kDa glutathione S-transferase from Schistosoma japonicum complexed with gamma-glutamyl[S-(2-iodobenzyl)cysteinyl]glycine | | Descriptor: | GAMMA-GLUTAMYL[S-(2-IODOBENZYL)CYSTEINYL]GLYCINE, Glutathione S-Transferase 26 kDa | | Authors: | Cardoso, R.M.F, Daniels, D.S, Bruns, C.M, Tainer, J.A. | | Deposit date: | 2002-07-28 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Characterization of the electrophile
binding site and substrate binding
mode of the 26-kDa glutathione
S-transferase from Schistosoma
japonicum
PROTEINS: STRUCT.,FUNCT.,GENET., 51, 2003
|
|
1M99
 
 | | Crystal structure of the 26 kDa glutathione S-transferase from Schistosoma japonicum complexed with glutathione sulfonic acid | | Descriptor: | GLUTATHIONE SULFONIC ACID, Glutathione S-Transferase 26kDa | | Authors: | Cardoso, R.M.F, Daniels, D.S, Bruns, C.M, Tainer, J.A. | | Deposit date: | 2002-07-28 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Characterization of the electrophile
binding site and substrate binding
mode of the 26-kDa glutathione
S-transferase from Schistosoma
japonicum
PROTEINS: STRUCT.,FUNCT.,GENET., 51, 2003
|
|
1PJS
 
 | | The co-crystal structure of CysG, the multifunctional methyltransferase/dehydrogenase/ferrochelatase for siroheme synthesis, in complex with it NAD cofactor | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, PHOSPHATE ION, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Stroupe, M.E, Leech, H.K, Daniels, D.S, Warren, M.J, Getzoff, E.D. | | Deposit date: | 2003-06-03 | | Release date: | 2003-12-02 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | CysG structure reveals tetrapyrrole-binding features and novel regulation of siroheme biosynthesis.
Nat.Struct.Biol., 10, 2003
|
|
1PJQ
 
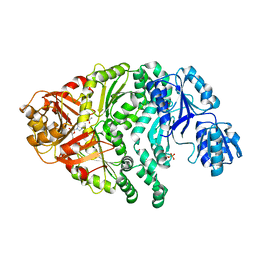 | | Structure and function of CysG, the multifunctional methyltransferase/dehydrogenase/ferrochelatase for siroheme synthesis | | Descriptor: | ACETATE ION, S-ADENOSYL-L-HOMOCYSTEINE, Siroheme synthase, ... | | Authors: | Stroupe, M.E, Leech, H.K, Daniels, D.S, Warren, M.J, Getzoff, E.D. | | Deposit date: | 2003-06-03 | | Release date: | 2003-12-02 | | Last modified: | 2014-11-12 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | CysG structure reveals tetrapyrrole-binding features and novel regulation of siroheme biosynthesis.
Nat.Struct.Biol., 10, 2003
|
|
1PJT
 
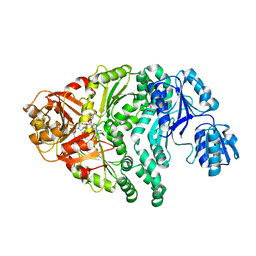 | | The structure of the Ser128Ala point-mutant variant of CysG, the multifunctional methyltransferase/dehydrogenase/ferrochelatase for siroheme synthesis | | Descriptor: | PHOSPHATE ION, S-ADENOSYL-L-HOMOCYSTEINE, Siroheme synthase | | Authors: | Stroupe, M.E, Leech, H.K, Daniels, D.S, Warren, M.J, Getzoff, E.D. | | Deposit date: | 2003-06-03 | | Release date: | 2003-12-02 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | CysG structure reveals tetrapyrrole-binding features and novel regulation of siroheme biosynthesis.
Nat.Struct.Biol., 10, 2003
|
|
3R5I
 
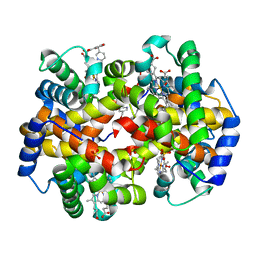 | | Crystal structure of liganded Hemoglobin complexed with a potent Antisickling agent, INN-312 | | Descriptor: | 5-methoxy-2-(pyridin-3-ylmethoxy)benzaldehyde, Hemoglobin subunit alpha, Hemoglobin subunit beta, ... | | Authors: | Safo, M.K, Musayev, F.N, Safo, R.P, Daniels, D, Eseonu, D.N, Parra, J. | | Deposit date: | 2011-03-18 | | Release date: | 2011-04-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and in Vitro Chracterization of Pyridyl Derivatives of Benzaldehydes : Highly Potent Antisickling Agents
To be Published
|
|
