3NY8
 
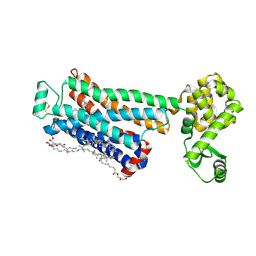 | | Crystal structure of the human beta2 adrenergic receptor in complex with the inverse agonist ICI 118,551 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, (2S,3S)-1-[(7-methyl-2,3-dihydro-1H-inden-4-yl)oxy]-3-[(1-methylethyl)amino]butan-2-ol, Beta-2 adrenergic receptor, ... | | Authors: | Wacker, D, Fenalti, G, Brown, M.A, Katritch, V, Abagyan, R, Cherezov, V, Stevens, R.C, Accelerated Technologies Center for Gene to 3D Structure (ATCG3D), GPCR Network (GPCR) | | Deposit date: | 2010-07-14 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Conserved Binding Mode of Human beta(2) Adrenergic Receptor Inverse Agonists and Antagonist Revealed by X-ray Crystallography
J.Am.Chem.Soc., 132, 2010
|
|
8QAU
 
 | | Outer kinetochore Ndc80-Dam1 alpha/beta-tubulin complex | | Descriptor: | DASH complex subunit DAM1, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Muir, K.W, Barford, D. | | Deposit date: | 2023-08-23 | | Release date: | 2024-04-03 | | Method: | ELECTRON MICROSCOPY (3.54 Å) | | Cite: | Mechanism of outer kinetochore assembly on microtubules and its regulation by mitotic error correction
Biorxiv, 2023
|
|
2V3J
 
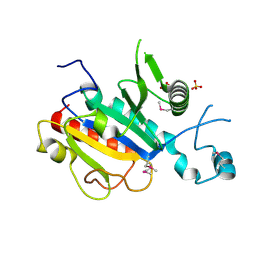 | | The yeast ribosome synthesis factor Emg1 alpha beta knot fold methyltransferase | | Descriptor: | ESSENTIAL FOR MITOTIC GROWTH 1, SULFATE ION | | Authors: | Leulliot, N, Bohnsack, M.T, Graille, M, Tollervey, D, Van Tilbeurgh, H. | | Deposit date: | 2007-06-18 | | Release date: | 2007-12-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Yeast Ribosome Synthesis Factor Emg1 is a Novel Member of the Superfamily of Alpha/Beta Knot Fold Methyltransferases.
Nucleic Acids Res., 36, 2008
|
|
2V7N
 
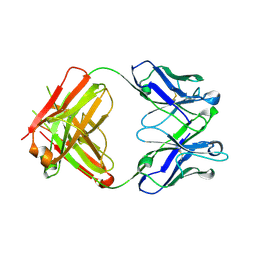 | | Unusual twinning in crystals of the CitS binding antibody Fab fragment f3p4 | | Descriptor: | IMMUNOGLOBULIN HEAVY CHAIN, IMMUNOGLOBULIN LIGHT CHAIN | | Authors: | Frey, D, Huber, T, Plueckthun, A, Gruetter, M.G. | | Deposit date: | 2007-07-31 | | Release date: | 2008-06-17 | | Last modified: | 2019-05-22 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Structure of the Recombinant Antibody Fab Fragment F3P4.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
8PIJ
 
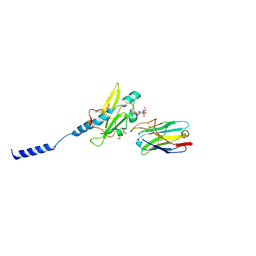 | | Neisseria meningitidis Type IV pilus SB-GATDH variant bound to the C24 nanobody | | Descriptor: | (2~{R})-~{N}-[(2~{R},3~{S},4~{S},5~{R},6~{R})-5-acetamido-2-methyl-4,6-bis(oxidanyl)oxan-3-yl]-2,3-bis(oxidanyl)propanamide, C24 nanobody, Pilin, ... | | Authors: | Fernandez-Martinez, D, Dumenil, G. | | Deposit date: | 2023-06-21 | | Release date: | 2024-04-03 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Cryo-EM structures of type IV pili complexed with nanobodies reveal immune escape mechanisms.
Nat Commun, 15, 2024
|
|
8PJP
 
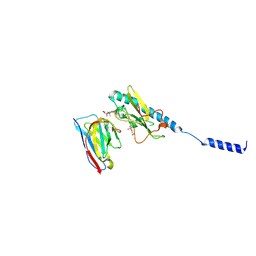 | | Neisseria meningitidis PilE, SB-GATDH variant, bound to the F10 nanobody | | Descriptor: | (2~{R})-~{N}-[(2~{R},3~{S},4~{S},5~{R},6~{R})-5-acetamido-2-methyl-4,6-bis(oxidanyl)oxan-3-yl]-2,3-bis(oxidanyl)propanamide, Nanobody F10, Pilin, ... | | Authors: | Fernandez-Martinez, D, Dumenil, G. | | Deposit date: | 2023-06-23 | | Release date: | 2024-04-03 | | Method: | ELECTRON MICROSCOPY (2.92 Å) | | Cite: | Cryo-EM structures of type IV pili complexed with nanobodies reveal immune escape mechanisms.
Nat Commun, 15, 2024
|
|
1OKY
 
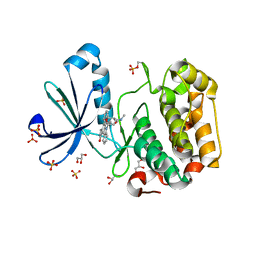 | | Structure of human PDK1 kinase domain in complex with staurosporine | | Descriptor: | 3-PHOSPHOINOSITIDE DEPENDENT PROTEIN KINASE 1, GLYCEROL, STAUROSPORINE, ... | | Authors: | Komander, D, Kular, G.S, Alessi, D.R, Van Aalten, D.M.F. | | Deposit date: | 2003-08-01 | | Release date: | 2004-07-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural Basis for Ucn-01 (7-Hydroxystaurosporine) Specificity and Pdk1 (3-Phosphoinositide-Dependent Protein Kinase-1) Inhibition
Biochem.J., 375, 2003
|
|
1OLL
 
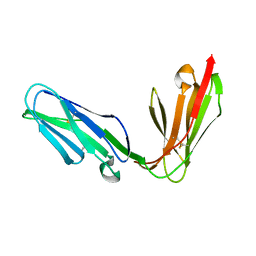 | | Extracellular region of the human receptor NKp46 | | Descriptor: | 1,2-ETHANEDIOL, NK RECEPTOR | | Authors: | Ponassi, M, Cantoni, C, Biassoni, R, Conte, R, Spallarossa, A, Pesce, A, Moretta, A, Moretta, L, Bolognesi, M, Bordo, D. | | Deposit date: | 2003-08-07 | | Release date: | 2003-09-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Structure of the Human Nk Cell Triggering Receptor Nkp46 Ectodomain
Biochem.Biophys.Res.Commun., 309, 2003
|
|
2UYH
 
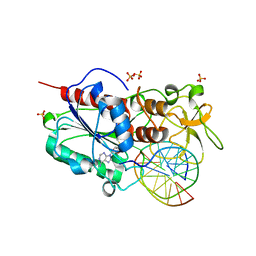 | |
4CZA
 
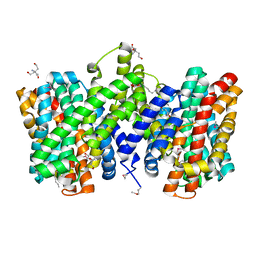 | | Structure of the sodium proton antiporter PaNhaP from Pyrococcus abyssii with bound thallium ion. | | Descriptor: | ACETATE ION, NA+/H+ ANTIPORTER, PUTATIVE, ... | | Authors: | Woehlert, D, Kuhlbrandt, W, Yildiz, O. | | Deposit date: | 2014-04-16 | | Release date: | 2014-12-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure and substrate ion binding in the sodium/proton antiporter PaNhaP.
Elife, 3, 2014
|
|
3O9L
 
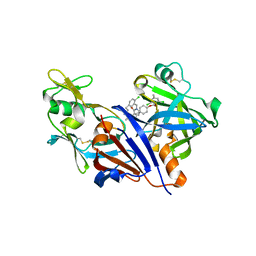 | | Design and optimisation of new piperidines as renin inhibitors | | Descriptor: | (3R,4S)-N-[2-chloro-5-(3-methoxypropyl)benzyl]-N-cyclopropyl-4-{4-[2-(2,6-dichloro-4-methylphenoxy)ethoxy]phenyl}piperidine-3-carboxamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, Renin | | Authors: | Corminboeuf, O, Bezencon, O, Grisostomi, C, Remen, L, Richard-Bildstein, S, Bur, D, Prade, L, Hess, P, Strickner, P, Treiber, A. | | Deposit date: | 2010-08-04 | | Release date: | 2011-03-02 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Design and optimization of new piperidines as renin inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
8QCX
 
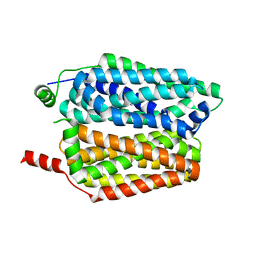 | |
2UYV
 
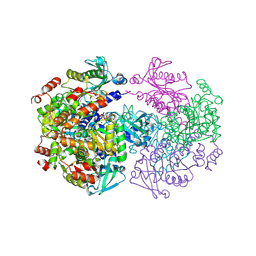 | |
1OHL
 
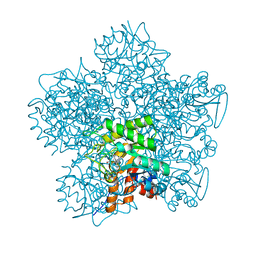 | | YEAST 5-AMINOLAEVULINIC ACID DEHYDRATASE PUTATIVE CYCLIC REACTION INTERMEDIATE COMPLEX | | Descriptor: | 3-[5-(AMINOMETHYL)-4-(CARBOXYMETHYL)-1H-PYRROL-3-YL]PROPANOIC ACID, BETA-MERCAPTOETHANOL, DELTA-AMINOLEVULINIC ACID DEHYDRATASE, ... | | Authors: | Erskine, P.T, Coates, L, Butler, D, Youell, J.H, Brindley, A.A, Wood, S.P, Warren, M.J, Shoolingin-Jordan, P.M, Cooper, J.B. | | Deposit date: | 2003-05-27 | | Release date: | 2003-06-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | X-Ray Structure of a Putative Reaction Intermediateof 5-Aminolaevulinic Acid Dehydratase
Biochem.J., 373, 2003
|
|
3NNJ
 
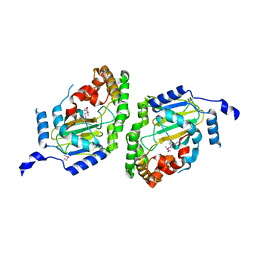 | | Halogenase domain from CurA module (apo Hal) | | Descriptor: | CurA | | Authors: | Khare, D, Smith, J.L. | | Deposit date: | 2010-06-23 | | Release date: | 2010-07-28 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Conformational switch triggered by alpha-ketoglutarate in a halogenase of curacin A biosynthesis
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
8QCS
 
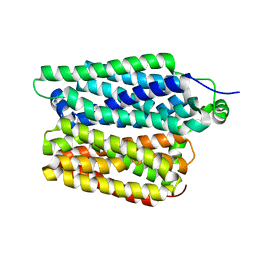 | |
8QCY
 
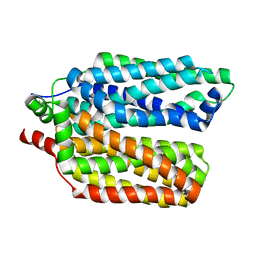 | |
1OKM
 
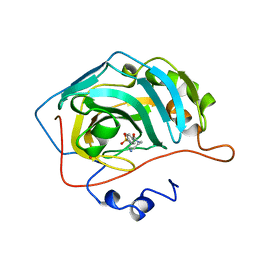 | | CARBONIC ANHYDRASE II COMPLEX WITH THE 1OKM INHIBITOR 4-SULFONAMIDE-[1-(4-AMINOBUTANE)]BENZAMIDE | | Descriptor: | 4-SULFONAMIDE-[1-(4-AMINOBUTANE)]BENZAMIDE, CARBONIC ANHYDRASE II, MERCURY (II) ION, ... | | Authors: | Elbaum, D, Nair, S.K, Patchan, M.W, Thompson, R.B, Christianson, D.W. | | Deposit date: | 1996-06-25 | | Release date: | 1997-01-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-based design of a sulfonamide probe for fluorescence anisotropy detection of zinc with a carbonic anhydrase-based biosensor.
J.Am.Chem.Soc., 118, 1996
|
|
2VHZ
 
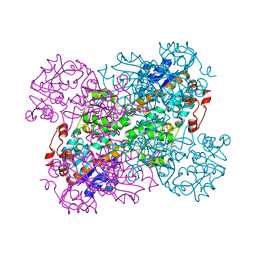 | |
2VHW
 
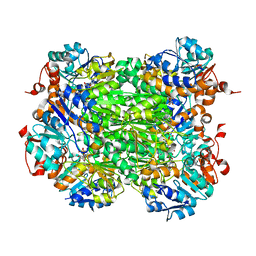 | |
8Q7V
 
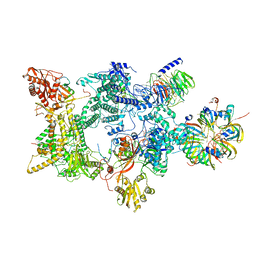 | | Structure of the recycling U5 snRNP bound to chaperones CD2BP2 and TSSC4 (State 1) | | Descriptor: | 116 kDa U5 small nuclear ribonucleoprotein component, CD2 antigen cytoplasmic tail-binding protein 2, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Riabov Bassat, D, Plaschka, C, Vorlaender, M.K. | | Deposit date: | 2023-08-17 | | Release date: | 2024-04-17 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural basis of human U5 snRNP late biogenesis and recycling.
Nat.Struct.Mol.Biol., 31, 2024
|
|
1OGK
 
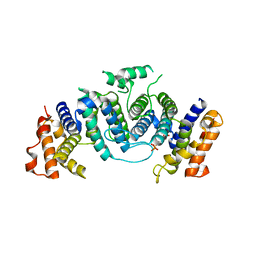 | | The crystal structure of Trypanosoma cruzi dUTPase in complex with dUDP | | Descriptor: | DEOXYURIDINE TRIPHOSPHATASE, DEOXYURIDINE-5'-DIPHOSPHATE | | Authors: | Harkiolaki, M, Dodson, E.J, Bernier-Villamor, V, Turkenburg, J.P, Gonzalez-Pacanowska, D, Wilson, K.S. | | Deposit date: | 2003-05-07 | | Release date: | 2004-01-22 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | The Crystal Structure of Trypanosoma Cruzi Dutpase Reveals a Novel Dutp/Dudp Binding Fold
Structure, 12, 2004
|
|
8Q7E
 
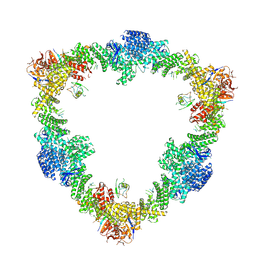 | |
1OGL
 
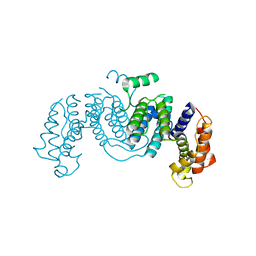 | | The crystal structure of native Trypanosoma cruzi dUTPase | | Descriptor: | DEOXYURIDINE TRIPHOSPHATASE | | Authors: | Harkiolaki, M, Dodson, E.J, Bernier-Villamor, V, Turkenburg, J.P, Gonzalez-Pacanowska, D, Wilson, K.S. | | Deposit date: | 2003-05-07 | | Release date: | 2004-01-22 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Crystal Structure of Trypanosoma Cruzi Dutpase Reveals a Novel Dutp/Dudp Binding Fold
Structure, 12, 2004
|
|
1OHD
 
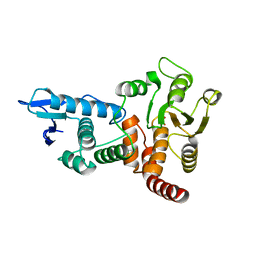 | | structure of cdc14 in complex with tungstate | | Descriptor: | CDC14B2 PHOSPHATASE, TUNGSTATE(VI)ION | | Authors: | Gray, C.H, Good, V.M, Tonks, N.K, Barford, D. | | Deposit date: | 2003-05-24 | | Release date: | 2003-07-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Structure of the Cell Cycle Protein Cdc14 Reveals a Proline-Directed Protein Phosphatase
Embo J., 22, 2003
|
|
