4OW1
 
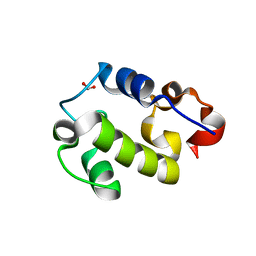 | | Crystal Structure of Resuscitation Promoting Factor C | | Descriptor: | 1,2-ETHANEDIOL, Resuscitation-promoting factor RpfC | | Authors: | Chauviac, F.X, Quay, D.H.X, Cohen-Gonsaud, M, Keep, N.H. | | Deposit date: | 2014-01-29 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The RpfC (Rv1884) atomic structure shows high structural conservation within the resuscitation-promoting factor catalytic domain.
Acta Crystallogr.,Sect.F, 70, 2014
|
|
2YMV
 
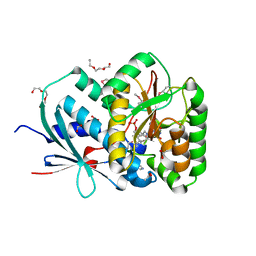 | | Structure of Reduced M Smegmatis 5246, a homologue of M.Tuberculosis Acg | | Descriptor: | 1-DEOXY-1-(7,8-DIMETHYL-2,4-DIOXO-3,4-DIHYDRO-2H-BENZO[G]PTERIDIN-1-ID-10(5H)-YL)-5-O-PHOSPHONATO-D-RIBITOL, ACETATE ION, ACG NITROREDUCTASE, ... | | Authors: | Chauviac, F.-X, Bommer, M, Dobbek, H, Keep, N.H. | | Deposit date: | 2012-10-10 | | Release date: | 2012-11-21 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure of Reduced Msacg, a Putative Nitroreductase from Mycobacterium Smegmatis and a Close Homologue of Mycobacterium Tuberculosis Acg.
J.Biol.Chem., 287, 2012
|
|
3CKX
 
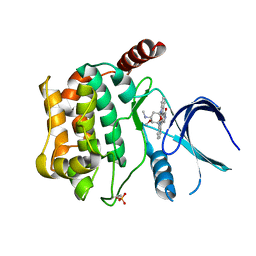 | | Crystal structure of sterile 20-like kinase 3 (MST3, STK24) in complex with staurosporine | | Descriptor: | STAUROSPORINE, Serine/threonine-protein kinase 24 | | Authors: | Antonysamy, S.S, Burley, S.K, Buchanan, S, Chau, F, Feil, I, Wu, L, Sauder, J.M, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-03-17 | | Release date: | 2008-04-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of sterile 20-like kinase 3 (MST3, STK24) in complex with staurosporine.
To be Published
|
|
3CKW
 
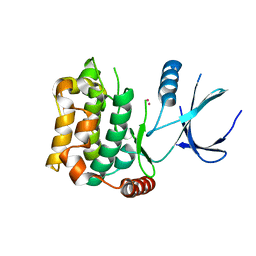 | | Crystal structure of sterile 20-like kinase 3 (MST3, STK24) | | Descriptor: | MERCURY (II) ION, Serine/threonine-protein kinase 24 | | Authors: | Antonysamy, S.S, Burley, S.K, Buchanan, S, Chau, F, Feil, I, Wu, L, Sauder, J.M, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-03-17 | | Release date: | 2008-04-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Crystal structure of sterile 20-like kinase 3 (MST3, STK24).
To be Published
|
|
3K8Z
 
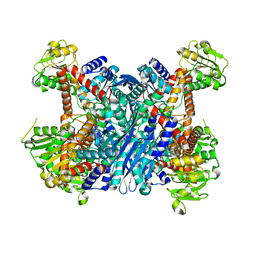 | | Crystal Structure of Gudb1 a decryptified secondary glutamate dehydrogenase from B. subtilis | | Descriptor: | NAD-specific glutamate dehydrogenase | | Authors: | Gunka, K, Newman, J.A, Commichau, F.M, Herzberg, C, Rodrigues, C, Hewitt, L, Lewis, R.J, Stulke, J. | | Deposit date: | 2009-10-15 | | Release date: | 2010-06-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Functional dissection of a trigger enzyme: mutations of the bacillus subtilis glutamate dehydrogenase RocG that affect differentially its catalytic activity and regulatory properties
J.Mol.Biol., 400, 2010
|
|
4Z45
 
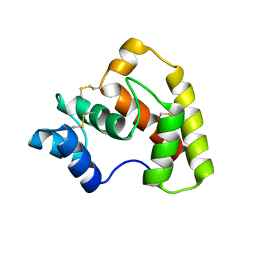 | | Structure of OBP3 from the currant-lettuce aphid Nasonovia ribisnigri | | Descriptor: | Odorant-binding protein NribOBP3 | | Authors: | Northey, T, Venthur, H, De Biasio, F, Chauviac, F.-X, Cole, A.R, Field, L.M, Zhou, J.-J, Keep, N.H. | | Deposit date: | 2015-04-01 | | Release date: | 2016-04-13 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Crystal Structures and Binding Dynamics of Odorant-Binding Protein 3 from two aphid species Megoura viciae and Nasonovia ribisnigri.
Sci Rep, 6, 2016
|
|
4Z39
 
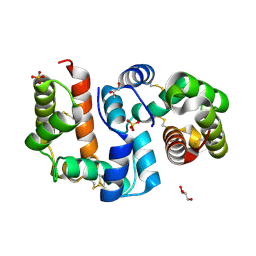 | | Structure of OBP3 from the vetch aphid Megoura viciae | | Descriptor: | GLYCEROL, Odorant-binding protein, SULFATE ION | | Authors: | Northey, T, Venthur, H, De Biasio, F, Chauviac, F.-X, Cole, A.R, Field, L.M, Zhou, J.-J, Keep, N.H. | | Deposit date: | 2015-03-31 | | Release date: | 2016-04-13 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structures and Binding Dynamics of Odorant-Binding Protein 3 from two aphid species Megoura viciae and Nasonovia ribisnigri.
Sci Rep, 6, 2016
|
|
3K92
 
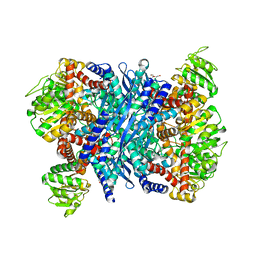 | | Crystal structure of a E93K mutant of the majour Bacillus subtilis glutamate dehydrogenase RocG | | Descriptor: | DI(HYDROXYETHYL)ETHER, NAD-specific glutamate dehydrogenase | | Authors: | Gunka, K, Newman, J.A, Commichau, F.M, Herzberg, C, Rodrigues, C, Hewitt, L, Lewis, R.J, Stulke, J. | | Deposit date: | 2009-10-15 | | Release date: | 2010-06-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Functional dissection of a trigger enzyme: mutations of the bacillus subtilis glutamate dehydrogenase RocG that affect differentially its catalytic activity and regulatory properties
J.Mol.Biol., 400, 2010
|
|
2VI7
 
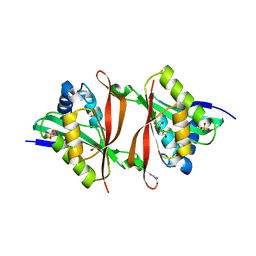 | | Structure of a Putative Acetyltransferase (PA1377)from Pseudomonas aeruginosa | | Descriptor: | ACETYLTRANSFERASE PA1377, AZIDE ION, GLYCEROL, ... | | Authors: | Davies, A.M, Tata, R, Chauviac, F.X, Sutton, B.J, Brown, P.R. | | Deposit date: | 2007-11-28 | | Release date: | 2008-05-20 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of a Putative Acetyltransferase (Pa1377) from Pseudomonas Aeruginosa.
Acta Crystallogr.,Sect.F, 64, 2008
|
|
2A2F
 
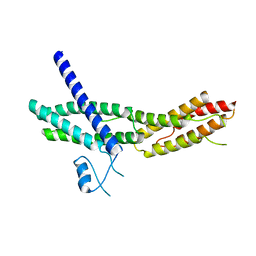 | | Crystal Structure of Sec15 C-terminal domain | | Descriptor: | Exocyst complex component Sec15 | | Authors: | Wu, S, Mehta, S.Q, Pichaud, F, Bellen, H.J, Quiocho, F.A. | | Deposit date: | 2005-06-22 | | Release date: | 2005-09-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Sec15 interacts with Rab11 via a novel domain and affects Rab11 localization in vivo.
Nat.Struct.Mol.Biol., 12, 2005
|
|
2WCI
 
 | | Structure of E. coli monothiol glutaredoxin GRX4 homodimer | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, GLUTAREDOXIN-4, GLUTATHIONE, ... | | Authors: | Iwema, T, Picchiocci, A, Traore, D.A.K, Ferrer, J.-L, Chauvat, F, Jacquamet, L. | | Deposit date: | 2009-03-12 | | Release date: | 2009-06-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis for Delivery of the Intact [Fe2S2] Cluster by Monothiol Glutaredoxin.
Biochemistry, 48, 2009
|
|
