1ADO
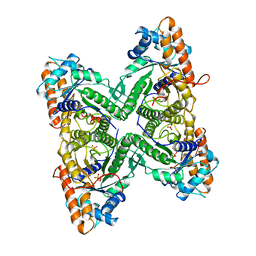 
 | | FRUCTOSE 1,6-BISPHOSPHATE ALDOLASE FROM RABBIT MUSCLE | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, ALDOLASE, SULFATE ION | | Authors: | Blom, N.S, Sygusch, J. | | Deposit date: | 1996-12-02 | | Release date: | 1997-12-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Product binding and role of the C-terminal region in class I D-fructose 1,6-bisphosphate aldolase.
Nat.Struct.Biol., 4, 1997
|
|
1FDJ
 
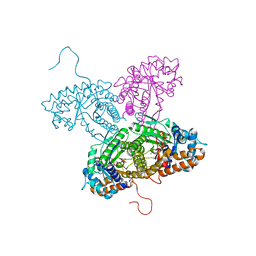 | | FRUCTOSE 1,6-BISPHOSPHATE ALDOLASE FROM RABBIT LIVER | | Descriptor: | 1,3-DIHYDROXYACETONEPHOSPHATE, 1,6-FRUCTOSE DIPHOSPHATE (LINEAR FORM), FRUCTOSE 1,6-BISPHOSPHATE ALDOLASE, ... | | Authors: | Blom, N.S, White, A, Sygusch, J. | | Deposit date: | 2000-07-20 | | Release date: | 2001-07-25 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Reaction intermediates of Rabbit liver D-fructose 1,6-bisphosphate Aldolase
To be Published
|
|
1DOS
 
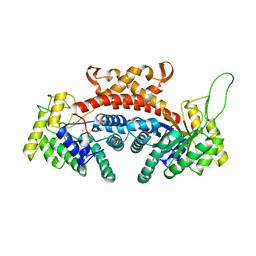 | | STRUCTURE OF FRUCTOSE-BISPHOSPHATE ALDOLASE | | Descriptor: | ALDOLASE CLASS II, AMMONIUM ION, ZINC ION | | Authors: | Blom, N, Tetreault, S, Coulombe, R, Sygusch, J. | | Deposit date: | 1996-06-24 | | Release date: | 1997-07-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Novel active site in Escherichia coli fructose 1,6-bisphosphate aldolase.
Nat.Struct.Biol., 3, 1996
|
|
1FHO
 
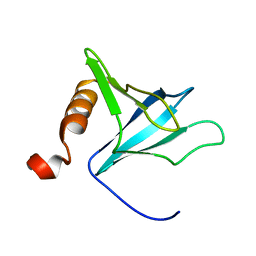 | | Solution Structure of the PH Domain from the C. Elegans Muscle Protein UNC-89 | | Descriptor: | UNC-89 | | Authors: | Blomberg, N, Baraldi, E, Sattler, M, Saraste, M, Nilges, M. | | Deposit date: | 2000-08-02 | | Release date: | 2000-10-04 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of a PH domain from the C. elegans muscle protein UNC-89 suggests a novel function.
Structure Fold.Des., 8, 2000
|
|
1I7G
 | | CRYSTAL STRUCTURE OF THE LIGAND BINDING DOMAIN FROM HUMAN PPAR-ALPHA IN COMPLEX WITH THE AGONIST AZ 242 | | Descriptor: | (2S)-2-ETHOXY-3-[4-(2-{4-[(METHYLSULFONYL)OXY]PHENYL}ETHOXY)PHENYL]PROPANOIC ACID, N,N-BIS(3-D-GLUCONAMIDOPROPYL)DEOXYCHOLAMIDE, PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR ALPHA, ... | | Authors: | Petersen, J.F.W, Cronet, P, Folmer, R, Blomberg, N, Sjoblom, K, Karlsson, U, Lindstedt, E.-L, Bamberg, K. | | Deposit date: | 2001-03-09 | | Release date: | 2002-03-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of the PPARalpha and -gamma ligand binding domain in complex with AZ 242; ligand selectivity and agonist activation in the PPAR family.
Structure, 9, 2001
|
|
1I7I
 
 | | CRYSTAL STRUCTURE OF THE LIGAND BINDING DOMAIN OF HUMAN PPAR-GAMMA IN COMPLEX WITH THE AGONIST AZ 242 | | Descriptor: | (2S)-2-ETHOXY-3-[4-(2-{4-[(METHYLSULFONYL)OXY]PHENYL}ETHOXY)PHENYL]PROPANOIC ACID, PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR GAMMA | | Authors: | Petersen, J.F.W, Cronet, P, Folmer, R, Blomberg, N, Sjoblom, K, Karlsson, U, Lindstedt, E.-L, Bamberg, K. | | Deposit date: | 2001-03-09 | | Release date: | 2002-03-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure of the PPARalpha and -gamma ligand binding domain in complex with AZ 242; ligand selectivity and agonist activation in the PPAR family.
Structure, 9, 2001
|
|
