4WP9
 
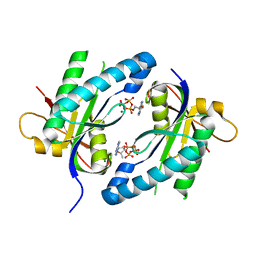 | | Crystal structure of Adenylyl cyclase MA1120 from Mycobacterium Avium bound to 2'5'-DD-3'-ATP, Calcium and Magnesium ion | | Descriptor: | 2',5'-dideoxyadenosine 3'-(tetrahydrogen triphosphate), CALCIUM ION, MAGNESIUM ION, ... | | Authors: | Bharambe, N.G, Barathy, D.V, Suguna, K. | | Deposit date: | 2014-10-17 | | Release date: | 2015-09-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.382 Å) | | Cite: | Autoinhibitory mechanism and activity-related structural changes in a mycobacterial adenylyl cyclase
J.Struct.Biol., 190, 2015
|
|
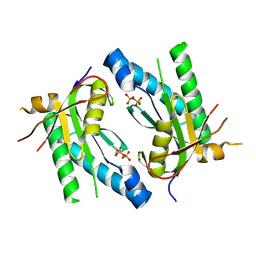
4WPA
 
 | |
8WCR
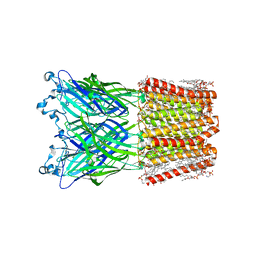 
 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | Authors: | Bharambe, N, Li, Z, Basak, S. | | Deposit date: | 2023-09-13 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
8WCQ
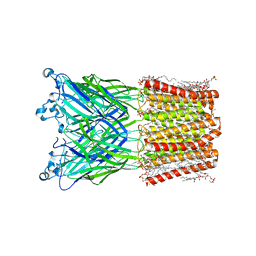 
 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | Authors: | Bharambe, N, Li, Z, Basak, S. | | Deposit date: | 2023-09-13 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (3.35 Å) | | Cite: | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
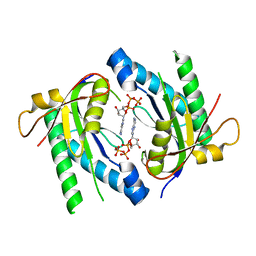
5D0H
 
 | |
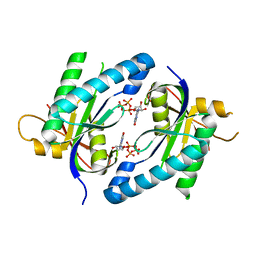
5D0G
 
 | |
5D15
 
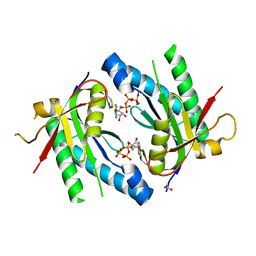 | | Crystal structure of an adenylyl cyclase Ma1120 from Mycobacterium avium in complex with ATP and calcium ion | | Descriptor: | 1,2-ETHANEDIOL, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Bharambe, N.G, Barathy, D.V, Suguna, K. | | Deposit date: | 2015-08-03 | | Release date: | 2016-08-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Substrate specificity determinants of class III nucleotidyl cyclases
Febs J., 283, 2016
|
|
5D0E
 
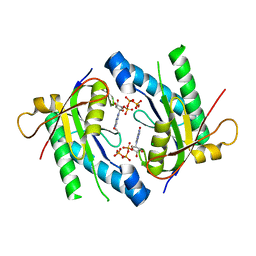 | | Crystal Structure of an adenylyl cyclase Ma1120-Cat in complex with GTP and calcium from Mycobacterium avium | | Descriptor: | CALCIUM ION, CHLORIDE ION, Cyclase, ... | | Authors: | Bharambe, N.G, Barathy, D.V, Suguna, K. | | Deposit date: | 2015-08-03 | | Release date: | 2016-08-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Substrate specificity determinants of class III nucleotidyl cyclases
Febs J., 283, 2016
|
|
8I41
 
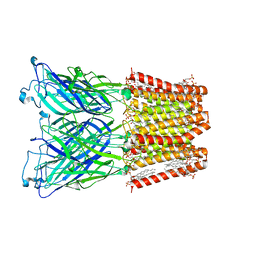 | |
8I42
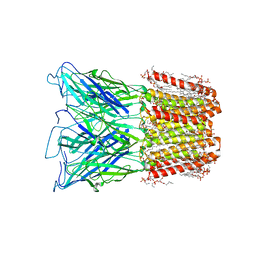 
 | |
8I48
 
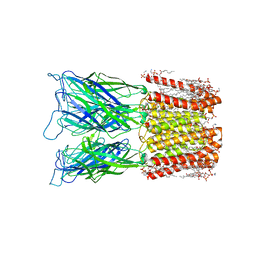 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | Authors: | Bharambe, N, Li, Z, Basak, S. | | Deposit date: | 2023-01-18 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
8I47
 
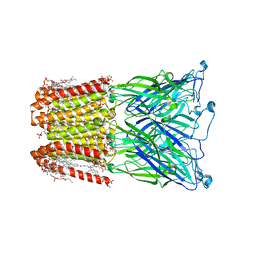 | |
8JJ3
 
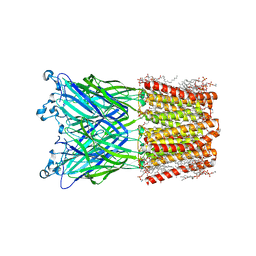 | | Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5 | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CHLORIDE ION, Proton-gated ion channel | | Authors: | Bharambe, N, Li, Z, Basak, S. | | Deposit date: | 2023-05-29 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (2.6476 Å) | | Cite: | Cryo-EM structures of prokaryotic ligand-gated ion channel GLIC provide insights into gating in a lipid environment.
Nat Commun, 15, 2024
|
|
4WP8
 
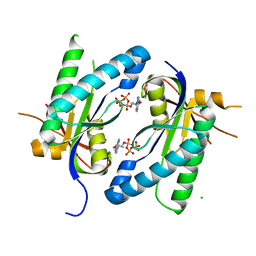 | | Crystal structure of Adenylyl cyclase Ma1120 from Mycobacterium Avium in complex with 2'5'-DD-3'-ATP and Manganese ion | | Descriptor: | 2',5'-dideoxyadenosine 3'-(tetrahydrogen triphosphate), CHLORIDE ION, MANGANESE (II) ION, ... | | Authors: | Barathy, D.V, Bharambe, N.G, Suguna, K. | | Deposit date: | 2014-10-17 | | Release date: | 2015-09-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Autoinhibitory mechanism and activity-related structural changes in a mycobacterial adenylyl cyclase
J.Struct.Biol., 190, 2015
|
|
4WP3
 
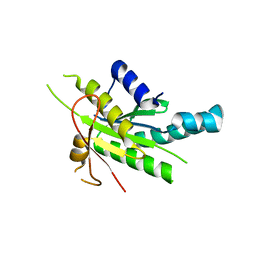 | |
