9BUN
 
 | |
5KPY
 
 | | Structure of a 5-hydroxytryptophan aptamer | | Descriptor: | 5-hydroxy-L-tryptophan, 5-hydroxytryptophan RNA aptamer, IRIDIUM HEXAMMINE ION, ... | | Authors: | Batey, R.T, Porter, E, Merck, M. | | Deposit date: | 2016-07-05 | | Release date: | 2017-01-11 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Recurrent RNA motifs as scaffolds for genetically encodable small-molecule biosensors.
Nat. Chem. Biol., 13, 2017
|
|
3LQX
 
 | | SRP ribonucleoprotein core complexed with cobalt hexammine | | Descriptor: | CHLORIDE ION, COBALT HEXAMMINE(III), POTASSIUM ION, ... | | Authors: | Batey, R.T. | | Deposit date: | 2010-02-10 | | Release date: | 2010-03-02 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Structural and Energetic Analysis of Metal Ions Essential to SRP Signal Recognition Domain Assembly
Biochemistry, 41, 2002
|
|
3D0X
 
 | |
1HQ1
 
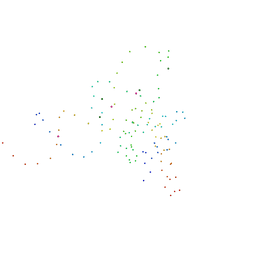 | | STRUCTURAL AND ENERGETIC ANALYSIS OF RNA RECOGNITION BY A UNIVERSALLY CONSERVED PROTEIN FROM THE SIGNAL RECOGNITION PARTICLE | | Descriptor: | 4.5S RNA DOMAIN IV, MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Batey, R.T, Sagar, M.B, Doudna, J.A. | | Deposit date: | 2000-12-13 | | Release date: | 2001-01-03 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Structural and energetic analysis of RNA recognition by a universally conserved protein from the signal recognition particle.
J.Mol.Biol., 307, 2001
|
|
1DUL
 
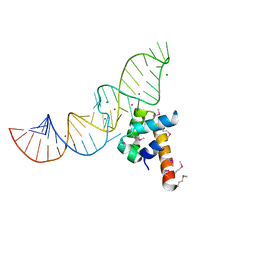 | | STRUCTURE OF THE RIBONUCLEOPROTEIN CORE OF THE E. COLI SIGNAL RECOGNITION PARTICLE | | Descriptor: | 4.5 S RNA DOMAIN IV, MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Batey, R.T, Rambo, R.P, Lucast, L, Rha, B, Doudna, J.A. | | Deposit date: | 2000-01-17 | | Release date: | 2000-02-28 | | Last modified: | 2020-10-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the ribonucleoprotein core of the signal recognition particle.
Science, 287, 2000
|
|
4GMA
 
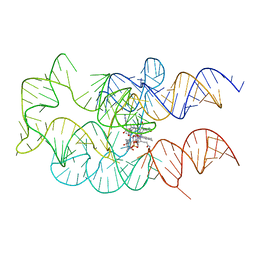 | | Crystal structure of the adenosylcobalamin riboswitch | | Descriptor: | Adenosylcobalamin, Adenosylcobalamin riboswitch | | Authors: | Reyes, F.E, Johnson, J.E, Polaski, J.T, Batey, R.T. | | Deposit date: | 2012-08-15 | | Release date: | 2012-10-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.94 Å) | | Cite: | B12 cofactors directly stabilize an mRNA regulatory switch.
Nature, 492, 2012
|
|
6DN1
 
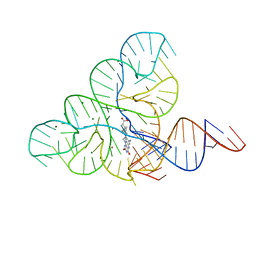 | | CRYSTAL STRUCTURE OF THE FMN RIBOSWITCH BOUND TO BRX1151 SPLIT RNA | | Descriptor: | 10-(6-carboxyhexyl)-8-(cyclopentylamino)-2,4-dihydroxy-7-methylbenzo[g]pteridin-10-ium, MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Vicens, Q, Mondragon, E, Reyes, F.E, Berman, J, Kaur, H, Kells, K, Wickens, P, Wilson, J, Gadwood, R, Schostarez, H, Suto, R.K, Coish, P, Blount, K.F, Batey, R.T. | | Deposit date: | 2018-06-05 | | Release date: | 2018-09-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | Structure-Activity Relationship of Flavin Analogues That Target the Flavin Mononucleotide Riboswitch.
ACS Chem. Biol., 13, 2018
|
|
2G9C
 
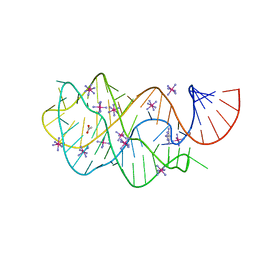 | | Modified pyrimidines Specifically bind the purine riboswitch | | Descriptor: | ACETATE ION, COBALT HEXAMMINE(III), PYRIMIDINE-2,4,6-TRIAMINE, ... | | Authors: | Gilbert, S.D, Mediatore, S.J, Batey, R.T. | | Deposit date: | 2006-03-06 | | Release date: | 2006-11-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Modified pyrimidines specifically bind the purine riboswitch.
J.Am.Chem.Soc., 128, 2006
|
|
2GIS
 
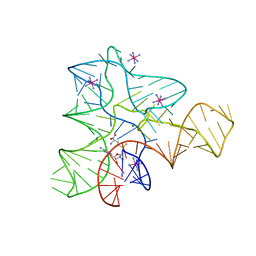 | |
3SD3
 
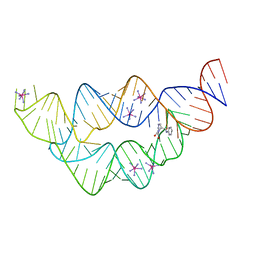 | | The structure of the tetrahydrofolate riboswitch containing a U25C mutation | | Descriptor: | IRIDIUM HEXAMMINE ION, N-[4-({[(6S)-2-amino-5-formyl-4-oxo-3,4,5,6,7,8-hexahydropteridin-6-yl]methyl}amino)benzoyl]-L-glutamic acid, Tetrahydrofolate riboswitch | | Authors: | Reyes, F.E, Trausch, J.J, Ceres, P, Batey, R.T. | | Deposit date: | 2011-06-08 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The structure of a tetrahydrofolate-sensing riboswitch reveals two ligand binding sites in a single aptamer.
Structure, 19, 2011
|
|
4L81
 
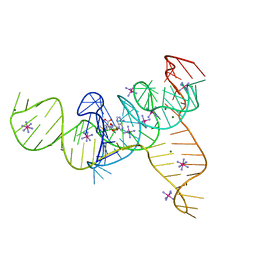 | | Structure of the SAM-I/IV riboswitch (env87(deltaU92, deltaG93)) | | Descriptor: | COBALT HEXAMMINE(III), MAGNESIUM ION, S-ADENOSYLMETHIONINE, ... | | Authors: | Trausch, J.J, Reyes, F.E, Edwards, A.L, Batey, R.T. | | Deposit date: | 2013-06-15 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural basis for diversity in the SAM clan of riboswitches.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
6P2H
 
 | |
3NPN
 
 | |
3NPQ
 
 | |
4XWF
 
 | |
4XW7
 
 | |
6DN2
 
 | | CRYSTAL STRUCTURE OF THE FMN RIBOSWITCH BOUND TO BRX1354 SPLIT RNA | | Descriptor: | 4-{benzyl[2-(7,8-dimethyl-2,4-dioxo-3,4-dihydrobenzo[g]pteridin-10(2H)-yl)ethyl]amino}butanoic acid, MAGNESIUM ION, POTASSIUM ION, ... | | Authors: | Vicens, Q, Mondragon, E, Reyes, F.E, Berman, J, Kaur, H, Kells, K, Wickens, P, Wilson, J, Gadwood, R, Schostarez, H, Suto, R.K, Coish, P, Blount, K.F, Batey, R.T. | | Deposit date: | 2018-06-05 | | Release date: | 2018-09-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | Structure-Activity Relationship of Flavin Analogues That Target the Flavin Mononucleotide Riboswitch.
ACS Chem. Biol., 13, 2018
|
|
6DN3
 
 | | CRYSTAL STRUCTURE OF THE FMN RIBOSWITCH BOUND TO BRX1555 SPLIT RNA | | Descriptor: | 7,8-dimethyl-2,4-dioxo-10-(3-phenylpropyl)-1,2,3,4-tetrahydrobenzo[g]pteridin-10-ium, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Vicens, Q, Mondragon, E, Reyes, F.E, Berman, J, Kaur, H, Kells, K, Wickens, P, Wilson, J, Gadwood, R, Schostarez, H, Suto, R.K, Coish, P, Blount, K.F, Batey, R.T. | | Deposit date: | 2018-06-05 | | Release date: | 2018-09-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure-Activity Relationship of Flavin Analogues That Target the Flavin Mononucleotide Riboswitch.
ACS Chem. Biol., 13, 2018
|
|
6UC8
 
 | | Guanine riboswitch bound to 8-aminoguanine | | Descriptor: | 8-AMINOGUANINE, ACETATE ION, COBALT HEXAMMINE(III), ... | | Authors: | Matyjasik, M.M, Hall, S.D, Batey, R.T. | | Deposit date: | 2019-09-15 | | Release date: | 2020-07-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | High Affinity Binding of N2-Modified Guanine Derivatives Significantly Disrupts the Ligand Binding Pocket of the Guanine Riboswitch.
Molecules, 25, 2020
|
|
6UC9
 
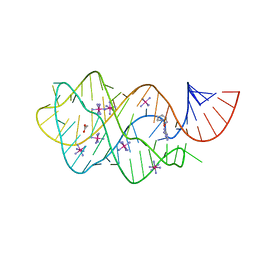 | | Guanine riboswitch bound to O6-cyclohexylmethyl guanine | | Descriptor: | 6-O-CYCLOHEXYLMETHYL GUANINE, ACETATE ION, COBALT HEXAMMINE(III), ... | | Authors: | Matyjasik, M.M, Batey, R.T. | | Deposit date: | 2019-09-15 | | Release date: | 2020-07-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.941 Å) | | Cite: | High Affinity Binding of N2-Modified Guanine Derivatives Significantly Disrupts the Ligand Binding Pocket of the Guanine Riboswitch.
Molecules, 25, 2020
|
|
6UBU
 
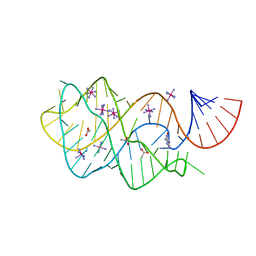 | |
6UC7
 
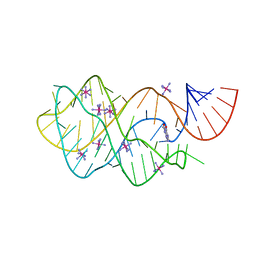 | | Structure of guanine riboswitch bound to N2-acetyl guanine | | Descriptor: | COBALT HEXAMMINE(III), N-(6-oxo-6,9-dihydro-3H-purin-2-yl)acetamide, guanine riboswitch | | Authors: | Matyjasik, M.M, Batey, R.T. | | Deposit date: | 2019-09-15 | | Release date: | 2020-07-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.798 Å) | | Cite: | High Affinity Binding of N2-Modified Guanine Derivatives Significantly Disrupts the Ligand Binding Pocket of the Guanine Riboswitch.
Molecules, 25, 2020
|
|
1ZJR
 
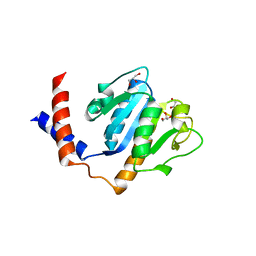 | | Crystal Structure of A. aeolicus TrmH/SpoU tRNA modifying enzyme | | Descriptor: | GLYCEROL, SULFATE ION, tRNA (Guanosine-2'-O-)-methyltransferase | | Authors: | Pleshe, E, Truesdell, J, Batey, R.T. | | Deposit date: | 2005-04-30 | | Release date: | 2005-08-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure of a class II TrmH tRNA-modifying enzyme from Aquifex aeolicus.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
4LVZ
 
 | |
