2R52
 | |
1R8G
 
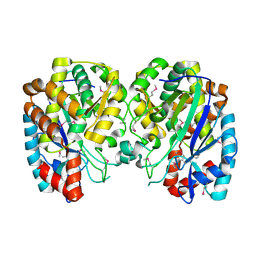 | | Structure and function of YbdK | | Descriptor: | Hypothetical protein ybdK | | Authors: | Lehmann, C, Doseeva, V, Pullalarevu, S, Krajewski, W, Howard, A, Herzberg, O, Structure 2 Function Project (S2F) | | Deposit date: | 2003-10-24 | | Release date: | 2004-08-17 | | Last modified: | 2021-07-28 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | YbdK is a carboxylate-amine ligase with a gamma-glutamyl:Cysteine ligase activity: crystal structure and enzymatic assays
PROTEINS: STRUCT.,FUNCT.,GENET., 56, 2004
|
|
1ESO
 
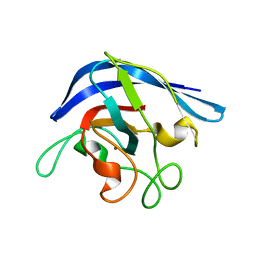 | | MONOMERIC CU,ZN SUPEROXIDE DISMUTASE FROM ESCHERICHIA COLI | | Descriptor: | COPPER (II) ION, CU, ZN SUPEROXIDE DISMUTASE, ... | | Authors: | Pesce, A, Capasso, C, Battistoni, A, Folcarelli, S, Rotilio, G, Desideri, A, Bolognesi, M. | | Deposit date: | 1997-06-27 | | Release date: | 1998-07-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Unique structural features of the monomeric Cu,Zn superoxide dismutase from Escherichia coli, revealed by X-ray crystallography.
J.Mol.Biol., 274, 1997
|
|
3EPV
 
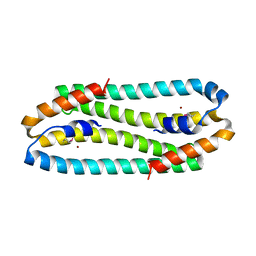 | | X-ray Structure of the Metal-sensor CnrX in both the Apo- and Copper-bound Forms | | Descriptor: | COPPER (II) ION, Nickel and cobalt resistance protein cnrR | | Authors: | Pompidor, G, Maillard, A.P, Girard, E, Gambarelli, S, Kahn, R, Coves, J. | | Deposit date: | 2008-09-30 | | Release date: | 2008-11-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.742 Å) | | Cite: | X-ray structure of the metal-sensor CnrX in both the apo- and copper-bound forms.
Febs Lett., 2008
|
|
1R54
 | | Crystal structure of the catalytic domain of human ADAM33 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ADAM 33, CALCIUM ION, ... | | Authors: | Orth, P, Reicher, P, Wang, W, Prosise, W.W, Yarosh-Tomaine, T, Hammond, G, Xiao, L, Mirza, U.A, Zou, J, Strickland, C, Taremi, S.S. | | Deposit date: | 2003-10-09 | | Release date: | 2004-10-12 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structre of the catalytic domain of human ADAM33
J.Mol.Biol., 335, 2004
|
|
2IX6
 
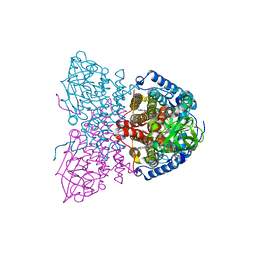 | | SHORT CHAIN SPECIFIC ACYL-COA OXIDASE FROM ARABIDOPSIS THALIANA, ACX4 | | Descriptor: | ACYL-COENZYME A OXIDASE 4, PEROXISOMAL, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Mackenzie, J, Pedersen, L, Arent, S, Henriksen, A. | | Deposit date: | 2006-07-07 | | Release date: | 2006-08-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Controlling Electron Transfer in Acyl-Coa Oxidases and Dehydrogenases: A Structural View.
J.Biol.Chem., 281, 2006
|
|
1MWW
 
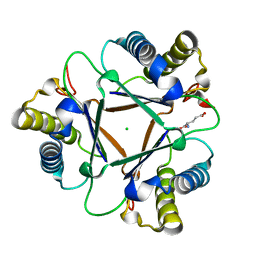 | | THE STRUCTURE OF THE HYPOTHETICAL PROTEIN HI1388.1 FROM HAEMOPHILUS INFLUENZAE REVEALS A TAUTOMERASE/MIF FOLD | | Descriptor: | CHLORIDE ION, GLUTAMIC ACID, HYPOTHETICAL PROTEIN HI1388.1 | | Authors: | Lehmann, C, Pullalarevu, S, Krajewski, W, Galkin, A, Howard, A, Herzberg, O, Structure 2 Function Project (S2F) | | Deposit date: | 2002-10-01 | | Release date: | 2003-11-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Structure of the Hypothetical Protein HI1388.1 from Haemophilus influenzae
To be Published
|
|
2IX5
 
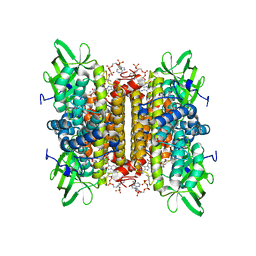 | | Short chain specific acyl-CoA oxidase from Arabidopsis thaliana, ACX4 in complex with acetoacetyl-CoA | | Descriptor: | ACETOACETYL-COENZYME A, ACYL-COENZYME A OXIDASE 4, PEROXISOMAL, ... | | Authors: | Mackenzie, J, Pedersen, L, Arent, S, Henriksen, A. | | Deposit date: | 2006-07-07 | | Release date: | 2006-08-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Controlling Electron Transfer in Acyl-Coa Oxidases and Dehydrogenases: A Structural View.
J.Biol.Chem., 281, 2006
|
|
1LCX
 
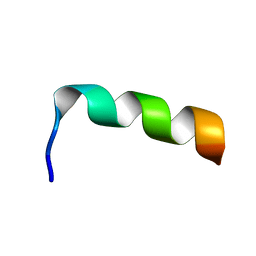 | | NMR structure of HIV-1 gp41 659-671 13mer peptide | | Descriptor: | GP41 | | Authors: | Biron, Z, Khare, S, Samson, A.O, Hayek, Y, Naider, F, Anglister, J. | | Deposit date: | 2002-04-07 | | Release date: | 2002-12-04 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A Monomeric 3(10)-Helix Is Formed in Water by a 13-Residue Peptide
Representing the Neutralizing Determinant of HIV-1 on gp41.
Biochemistry, 41, 2002
|
|
2R80
 
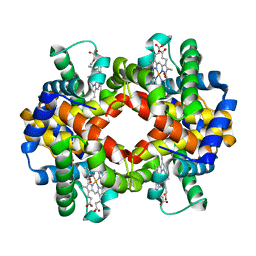 | | Pigeon Hemoglobin (OXY form) | | Descriptor: | Hemoglobin subunit alpha-A, Hemoglobin subunit beta, OXYGEN MOLECULE, ... | | Authors: | Ponnuswamy, M.N, Packianathan, C, Sundaresan, S, Neelagandan, K, Palani, K, Muller, J.J, Heinemann, U. | | Deposit date: | 2007-09-10 | | Release date: | 2008-09-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | X-ray crystal structure analysis of Hemolgobin from Pigeon (Columba Livia) at 1.44 angstrom
To be Published
|
|
1N8M
 | | Solution structure of Pi4, a four disulfide bridged scorpion toxin active on potassium channels | | Descriptor: | Potassium channel blocking toxin 4 | | Authors: | Guijarro, J.I, M'Barek, S, Olamendi-Portugal, T, Gomez-Lagunas, F, Garnier, D, Rochat, H, Possani, L.D, Sabatier, J.M, Delepierre, M. | | Deposit date: | 2002-11-21 | | Release date: | 2003-09-02 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Pi4, a short four-disulfide-bridged scorpion toxin specific of potassium channels.
Protein Sci., 12, 2003
|
|
1NNX
 
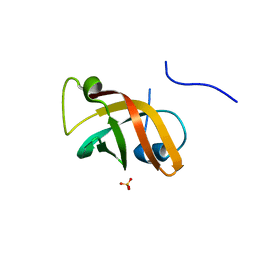 | | Structure of the hypothetical protein ygiW from E. coli. | | Descriptor: | Protein ygiW, SULFATE ION | | Authors: | Lehmann, C, Galkin, A, Pullalarevu, S, Sarikaya, E, Krajewski, W, Lim, K, Howard, A, Herzberg, O, Structure 2 Function Project (S2F) | | Deposit date: | 2003-01-14 | | Release date: | 2004-03-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structure of the hypothetical protein ygiW from E. coli.
To be Published
|
|
1RW1
 
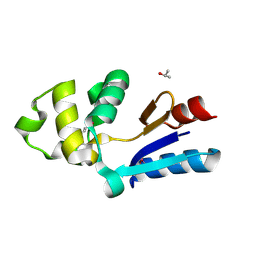 | | YFFB (PA3664) PROTEIN | | Descriptor: | ISOPROPYL ALCOHOL, conserved hypothetical protein yffB | | Authors: | Teplyakov, A, Pullalarevu, S, Obmolova, G, Doseeva, V, Galkin, A, Herzberg, O, Dauter, M, Dauter, Z, Gilliland, G.L, Structure 2 Function Project (S2F) | | Deposit date: | 2003-12-15 | | Release date: | 2004-11-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.02 Å) | | Cite: | Crystal structure of the YffB protein from Pseudomonas aeruginosa suggests a glutathione-dependent thiol reductase function.
Bmc Struct.Biol., 4, 2004
|
|
2F2V
 
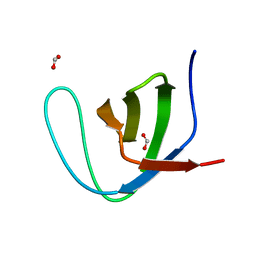 | | alpha-spectrin SH3 domain A56G mutant | | Descriptor: | FORMIC ACID, Spectrin alpha chain, brain | | Authors: | Camara-Artigas, A, Conejero-Lara, F, Casares, S, Lopez-Mayorga, O, Vega, C. | | Deposit date: | 2005-11-18 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Cooperative propagation of local stability changes from low-stability and high-stability regions in a SH3 domain
Proteins, 67, 2007
|
|
2JM8
 
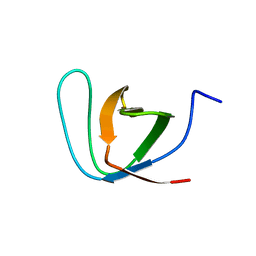 | | R21A Spc-SH3 free | | Descriptor: | Spectrin alpha chain, brain | | Authors: | van Nuland, N.A.J, Casares, S, Ab, E, Eshuis, H, Lopez-Mayorga, O, Conejero-Lara, F. | | Deposit date: | 2006-10-25 | | Release date: | 2007-04-24 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | The high-resolution NMR structure of the R21A Spc-SH3:P41 complex: Understanding the determinants of binding affinity by comparison with Abl-SH3
Bmc Struct.Biol., 7, 2007
|
|
2JMA
 
 | | R21A Spc-SH3:P41 complex | | Descriptor: | P41 peptide, Spectrin alpha chain, brain | | Authors: | van Nuland, N.A.J, Casares, S, Ab, E, Eshuis, H, Lopez-Mayorga, O, Conejero-Lara, F. | | Deposit date: | 2006-10-25 | | Release date: | 2007-04-24 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | The high-resolution NMR structure of the R21A Spc-SH3:P41 complex: Understanding the determinants of binding affinity by comparison with Abl-SH3
Bmc Struct.Biol., 7, 2007
|
|
2JM9
 | | R21A Spc-SH3 bound | | Descriptor: | Spectrin alpha chain, brain | | Authors: | van Nuland, N.A.J, Casares, S, Ab, E, Eshuis, H, Lopez-Mayorga, O, Conejero-Lara, F. | | Deposit date: | 2006-10-25 | | Release date: | 2007-04-24 | | Last modified: | 2023-12-20 | | Method: | SOLUTION NMR | | Cite: | The high-resolution NMR structure of the R21A Spc-SH3:P41 complex: Understanding the determinants of binding affinity by comparison with Abl-SH3
Bmc Struct.Biol., 7, 2007
|
|
