8Z0X
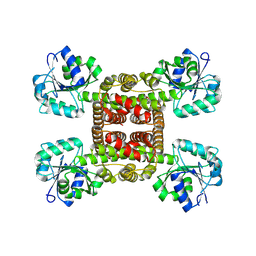 
 | | Crystal structure of glyoxylate reductase from Acetobacter aceti in the apo form | | Descriptor: | 3-hydroxyisobutyrate dehydrogenase | | Authors: | Majumder, T.R, Yoshizawa, T, Inoue, M, Aono, R, Matsumura, H, Mihara, H. | | Deposit date: | 2024-04-10 | | Release date: | 2024-10-23 | | Last modified: | 2025-05-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural insights into the mechanism underlying the dual cofactor specificity of glyoxylate reductase from Acetobacter aceti in the beta-hydroxyacid dehydrogenase family.
Biochim Biophys Acta Proteins Proteom, 1873, 2025
|
|
8Z9G
 
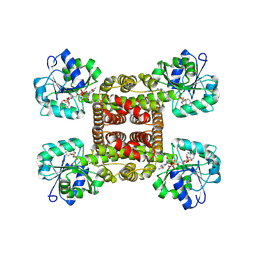 | | Crystal structure of glyoxylate reductase from Acetobacter aceti in complex with NADPH | | Descriptor: | 3-hydroxyisobutyrate dehydrogenase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Majumder, T.R, Yoshizawa, T, Inoue, M, Aono, R, Matsumura, H, Mihara, H. | | Deposit date: | 2024-04-23 | | Release date: | 2024-10-23 | | Last modified: | 2025-05-07 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Structural insights into the mechanism underlying the dual cofactor specificity of glyoxylate reductase from Acetobacter aceti in the beta-hydroxyacid dehydrogenase family.
Biochim Biophys Acta Proteins Proteom, 1873, 2025
|
|
8Z9F
 
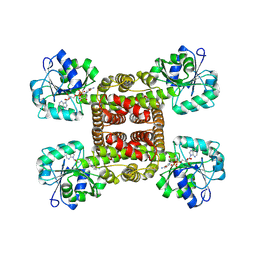 | | Crystal structure of glyoxylate reductase from Acetobacter aceti in complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, 3-hydroxyisobutyrate dehydrogenase | | Authors: | Majumder, T.R, Yoshizawa, T, Inoue, M, Aono, R, Matsumura, H, Mihara, H. | | Deposit date: | 2024-04-23 | | Release date: | 2024-10-23 | | Last modified: | 2025-05-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural insights into the mechanism underlying the dual cofactor specificity of glyoxylate reductase from Acetobacter aceti in the beta-hydroxyacid dehydrogenase family.
Biochim Biophys Acta Proteins Proteom, 1873, 2025
|
|
4GA6
 
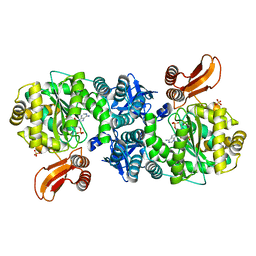 | | Crystal structure of AMP phosphorylase C-terminal deletion mutant in complex with substrates | | Descriptor: | ADENOSINE MONOPHOSPHATE, Putative thymidine phosphorylase, SULFATE ION | | Authors: | Nishitani, Y, Aono, R, Nakamura, A, Sato, T, Atomi, H, Imanaka, T, Miki, K. | | Deposit date: | 2012-07-25 | | Release date: | 2013-05-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Structure analysis of archaeal AMP phosphorylase reveals two unique modes of dimerization
J.Mol.Biol., 425, 2013
|
|
4GA5
 
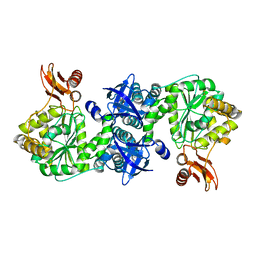 | | Crystal structure of AMP phosphorylase C-terminal deletion mutant in the apo-form | | Descriptor: | Putative thymidine phosphorylase | | Authors: | Nishitani, Y, Aono, R, Nakamura, A, Sato, T, Atomi, H, Imanaka, T, Miki, K. | | Deposit date: | 2012-07-25 | | Release date: | 2013-05-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Structure analysis of archaeal AMP phosphorylase reveals two unique modes of dimerization
J.Mol.Biol., 425, 2013
|
|
4GA4
 
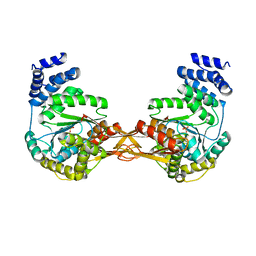 | | Crystal structure of AMP phosphorylase N-terminal deletion mutant | | Descriptor: | PHOSPHATE ION, Putative thymidine phosphorylase | | Authors: | Nishitani, Y, Aono, R, Nakamura, A, Sato, T, Atomi, H, Imanaka, T, Miki, K. | | Deposit date: | 2012-07-25 | | Release date: | 2013-05-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.51 Å) | | Cite: | Structure analysis of archaeal AMP phosphorylase reveals two unique modes of dimerization
J.Mol.Biol., 425, 2013
|
|
3VM6
 
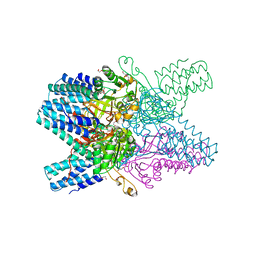 | | Crystal structure of ribose-1,5-bisphosphate isomerase from Thermococcus kodakarensis KOD1 in complex with alpha-D-ribose-1,5-bisphosphate | | Descriptor: | 1,5-di-O-phosphono-alpha-D-ribofuranose, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nakamura, A, Fujihashi, M, Aono, R, Sato, T, Nishiba, Y, Yoshida, S, Yano, A, Atomi, H, Imanaka, T, Miki, K. | | Deposit date: | 2011-12-08 | | Release date: | 2012-04-25 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Dynamic, ligand-dependent conformational change triggers reaction of ribose-1,5-bisphosphate isomerase from Thermococcus kodakarensis KOD1
J.Biol.Chem., 287, 2012
|
|
1OCC
 
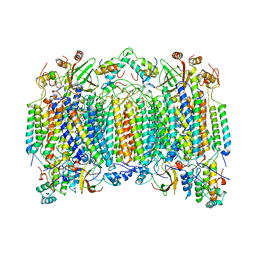 | | STRUCTURE OF BOVINE HEART CYTOCHROME C OXIDASE AT THE FULLY OXIDIZED STATE | | Descriptor: | COPPER (II) ION, CYTOCHROME C OXIDASE, HEME-A, ... | | Authors: | Tsukihara, T, Aoyama, H, Yamashita, E, Tomizaki, T, Yamaguchi, H, Shinzawa-Itoh, K, Nakashima, R, Yaono, R, Yoshikawa, S. | | Deposit date: | 1996-04-18 | | Release date: | 1996-12-07 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 A.
Science, 272, 1996
|
|
