7M2M
 
 | | NMR Structure of GCAP5 | | Descriptor: | Guanylate cyclase activator 1A, MAGNESIUM ION | | Authors: | Ames, J.B, Cudia, D.L. | | Deposit date: | 2021-03-17 | | Release date: | 2021-10-20 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | NMR and EPR-DEER Structure of a Dimeric Guanylate Cyclase Activator Protein-5 from Zebrafish Photoreceptors.
Biochemistry, 60, 2021
|
|
2L7Z
 
 | | NMR Structure of A13 homedomain | | Descriptor: | Homeobox protein Hox-A13 | | Authors: | Ames, J. | | Deposit date: | 2010-12-27 | | Release date: | 2011-11-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for sequence specific DNA binding and protein dimerization of HOXA13.
Plos One, 6, 2011
|
|
2M28
 
 | | NMR structure of Ca2+ bound CaBP4 C-domain | | Descriptor: | CALCIUM ION, Calcium-binding protein 4 | | Authors: | Ames, J.B. | | Deposit date: | 2012-12-17 | | Release date: | 2014-05-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Chemical shift assignment of Ca2+ bound CaPB4
To be Published
|
|
2M29
 
 | | NMR structure of Ca2+ bound CaBP4 N-domain | | Descriptor: | CALCIUM ION, Calcium-binding protein 4 | | Authors: | Ames, J.B. | | Deposit date: | 2012-12-17 | | Release date: | 2014-05-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Chemical shift assignment of Ca2+ bound CaPB4
To be Published
|
|
1FPW
 
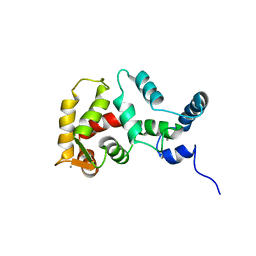 | | STRUCTURE OF YEAST FREQUENIN | | Descriptor: | CALCIUM ION, CALCIUM-BINDING PROTEIN NCS-1 | | Authors: | Ames, J.B, Hendricks, K.B, Strahl, T, Huttner, I.G, Thorner, J. | | Deposit date: | 2000-08-31 | | Release date: | 2000-10-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure and calcium-binding properties of Frq1, a novel calcium sensor in the yeast Saccharomyces cerevisiae.
Biochemistry, 39, 2000
|
|
7L8V
 
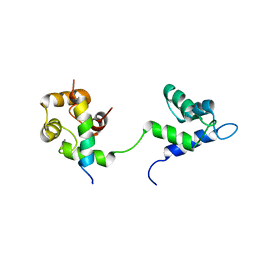 | |
1Z2K
 
 | | NMR structure of the D1 domain of the Natural Killer Cell Receptor, 2B4 | | Descriptor: | Natural killer cell receptor 2B4 | | Authors: | Ames, J.B, Vyas, V, Lusin, J.D, Mariuzza, R. | | Deposit date: | 2005-03-08 | | Release date: | 2005-05-03 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Natural Killer Cell Receptor 2B4:
Implications for Ligand Recognition
Biochemistry, 44, 2005
|
|
3CS1
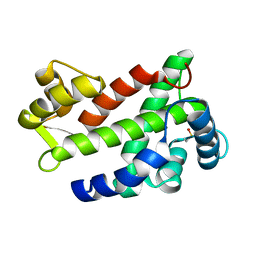 
 | | Flagellar Calcium-binding Protein (FCaBP) from T. cruzi | | Descriptor: | Flagellar calcium-binding protein | | Authors: | Ames, J.B, Ladner, J.E, Wingard, J.N, Robinson, H, Fisher, A. | | Deposit date: | 2008-04-08 | | Release date: | 2008-06-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Insights into Membrane Targeting by the Flagellar Calcium-binding Protein (FCaBP), a Myristoylated and Palmitoylated Calcium Sensor in Trypanosoma cruzi.
J.Biol.Chem., 283, 2008
|
|
2FM4
 
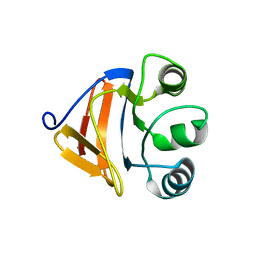 | |
2LVV
 
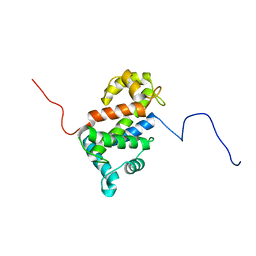 | | NMR structure of TB24 | | Descriptor: | Flagellar calcium-binding protein TB-24 | | Authors: | Ames, J. | | Deposit date: | 2012-07-11 | | Release date: | 2012-11-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the calflagin Tb24 flagellar calcium binding protein of Trypanosoma brucei.
Protein Sci., 21, 2012
|
|
1JSA
 
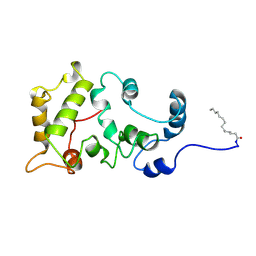 | | MYRISTOYLATED RECOVERIN WITH TWO CALCIUMS BOUND, NMR, 24 STRUCTURES | | Descriptor: | CALCIUM ION, MYRISTIC ACID, RECOVERIN | | Authors: | Ames, J.B, Ishima, R, Tanaka, T, Gordon, J.I, Stryer, L, Ikura, M. | | Deposit date: | 1997-06-04 | | Release date: | 1997-10-15 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Molecular mechanics of calcium-myristoyl switches.
Nature, 389, 1997
|
|
1JBA
 
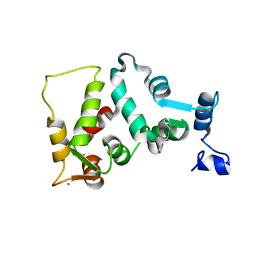 | | UNMYRISTOYLATED GCAP-2 WITH THREE CALCIUM IONS BOUND | | Descriptor: | CALCIUM ION, PROTEIN (GUANYLATE CYCLASE ACTIVATING PROTEIN 2) | | Authors: | Ames, J.B, Dizhoor, A.M, Ikura, M, Palczewski, K, Stryer, L. | | Deposit date: | 1999-04-03 | | Release date: | 1999-12-10 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of guanylyl cyclase activating protein-2, a calcium-sensitive modulator of photoreceptor guanylyl cyclases.
J.Biol.Chem., 274, 1999
|
|
1LA3
 
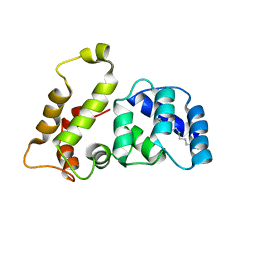 | | Solution structure of recoverin mutant, E85Q | | Descriptor: | CALCIUM ION, MYRISTIC ACID, Recoverin | | Authors: | Ames, J.B, Hamasaki, N, Molchanova, T. | | Deposit date: | 2002-03-27 | | Release date: | 2002-06-19 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Structure and calcium-binding studies of a recoverin mutant (E85Q) in an allosteric intermediate state.
Biochemistry, 41, 2002
|
|
2JU0
 
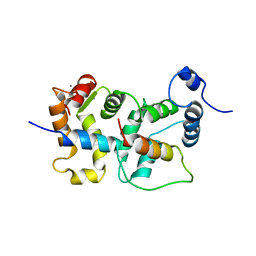 | | Structure of Yeast Frequenin bound to PdtIns 4-kinase | | Descriptor: | CALCIUM ION, Calcium-binding protein NCS-1, Phosphatidylinositol 4-kinase PIK1 | | Authors: | Ames, J. | | Deposit date: | 2007-08-11 | | Release date: | 2007-08-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural insights into activation of phosphatidylinositol 4-kinase (Pik1) by yeast frequenin (Frq1).
J.Biol.Chem., 282, 2007
|
|
2KPH
 
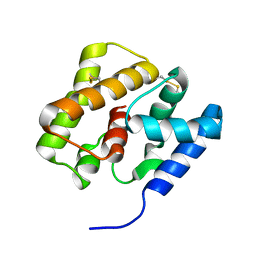 | | NMR Structure of AtraPBP1 at pH 4.5 | | Descriptor: | Pheromone binding protein | | Authors: | Ames, J. | | Deposit date: | 2009-10-15 | | Release date: | 2010-02-02 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Navel Orangeworm Moth Pheromone-Binding Protein (AtraPBP1): Implications for pH-Sensitive Pheromone Detection .
Biochemistry, 49, 2010
|
|
2JUL
 
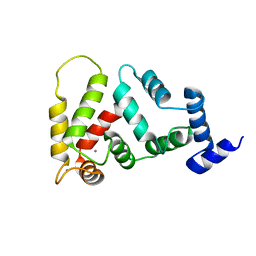 | | NMR Structure of DREAM | | Descriptor: | CALCIUM ION, Calsenilin | | Authors: | Ames, J. | | Deposit date: | 2007-08-30 | | Release date: | 2008-04-22 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | NMR structure of DREAM: Implications for Ca(2+)-dependent DNA binding and protein dimerization.
Biochemistry, 47, 2008
|
|
2I94
 
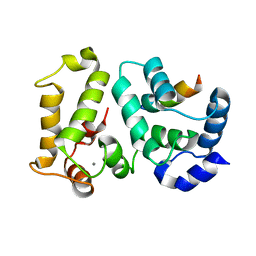 | | NMR Structure of recoverin bound to rhodopsin kinase | | Descriptor: | CALCIUM ION, Recoverin, Rhodopsin kinase | | Authors: | Ames, J.B. | | Deposit date: | 2006-09-05 | | Release date: | 2006-10-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Calcium-induced Inhibition of Rhodopsin Kinase by Recoverin.
J.Biol.Chem., 281, 2006
|
|
2K7C
 
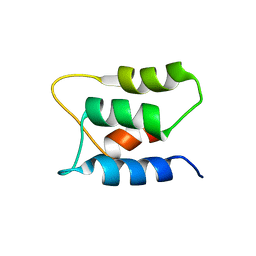 | |
2K7D
 
 | | NMR Structure of Ca2+-bound CaBP1 C-domain | | Descriptor: | CALCIUM ION, Calcium-binding protein 1 | | Authors: | Ames, J. | | Deposit date: | 2008-08-08 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Ca2+-bound CaBP1 C-domain
To be Published
|
|
2K7B
 
 | | NMR structure of Mg2+-bound CaBP1 N-domain | | Descriptor: | Calcium-binding protein 1, MAGNESIUM ION | | Authors: | Ames, J. | | Deposit date: | 2008-08-08 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR structure of Mg2+-bound CaBP1 N-domain
To be Published
|
|
2LAP
 
 | | NMR structure of Ca2+-bound CaBP1 C-domain with RDC | | Descriptor: | CALCIUM ION, Calcium-binding protein 1 | | Authors: | Ames, J. | | Deposit date: | 2011-03-16 | | Release date: | 2012-02-08 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | 1H, 15N, and 13C chemical shift assignments for CaBP1 with 3 Ca2+ bound
To be Published
|
|
2L2C
 
 | |
2L2E
 
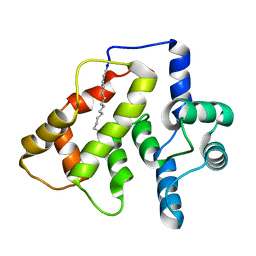 | |
2LAN
 
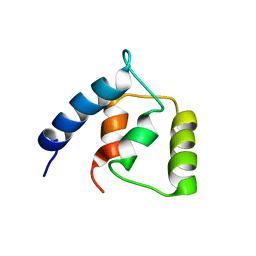 | | NMR structure of Ca2+-bound CaBP1 N-domain with RDC | | Descriptor: | CALCIUM ION, Calcium-binding protein 1 | | Authors: | Ames, J. | | Deposit date: | 2011-03-16 | | Release date: | 2012-02-08 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | 1H, 15N, and 13C chemical shift assignments for CaBP1 with 3 Ca2+ bound
To be Published
|
|
1U0J
 
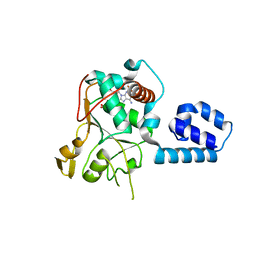 | | Crystal Structure of AAV2 Rep40-ADP complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA replication protein | | Authors: | James, J.A, Aggarwal, A.K, Linden, R.M, Escalante, C.R. | | Deposit date: | 2004-07-13 | | Release date: | 2004-08-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of adeno-associated virus type 2 Rep40-ADP complex: Insight into
nucleotide recognition and catalysis by superfamily 3 helicases
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
