6M75
 
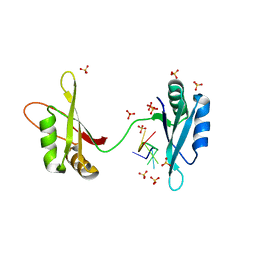 | | C-Myc DNA binding protein complex | | Descriptor: | DNA (5'-D(*TP*CP*TP*TP*AP*TP*T)-3'), RNA-binding motif, single-stranded-interacting protein 1, ... | | Authors: | Aggarwal, P, Bhavesh, N.S. | | Deposit date: | 2020-03-17 | | Release date: | 2021-03-17 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Hinge like domain motion facilitates human RBMS1 protein binding to proto-oncogene c-myc promoter.
Nucleic Acids Res., 49, 2021
|
|
7C36
 
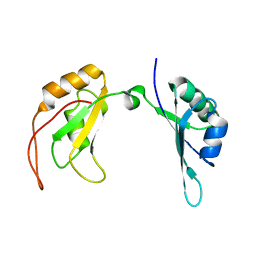 | | c-Myc DNA binding protein structure | | Descriptor: | RNA-binding motif, single-stranded-interacting protein 1 | | Authors: | Aggarwal, P, Bhavesh, N.S. | | Deposit date: | 2020-05-11 | | Release date: | 2021-05-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Hinge like domain motion facilitates human RBMS1 protein binding to proto-oncogene c-myc promoter.
Nucleic Acids Res., 49, 2021
|
|
2R8Z
 
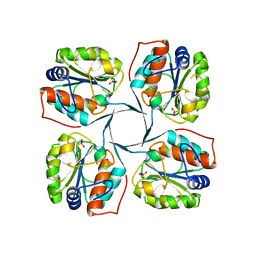 | | Crystal structure of YrbI phosphatase from Escherichia coli in complex with a phosphate and a calcium ion | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CALCIUM ION, PHOSPHATE ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8Y
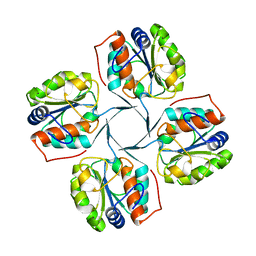 
 | | Crystal structure of YrbI phosphatase from Escherichia coli in a complex with Ca | | Descriptor: | CALCIUM ION, CHLORIDE ION, YrbI from Escherichia coli | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8X
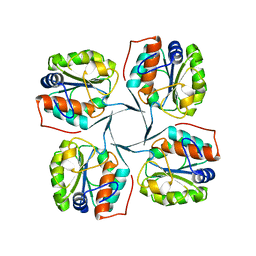 
 | | Crystal structure of YrbI phosphatase from Escherichia coli | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CHLORIDE ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R.W, Biswas, T. | | Deposit date: | 2007-09-11 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
2R8E
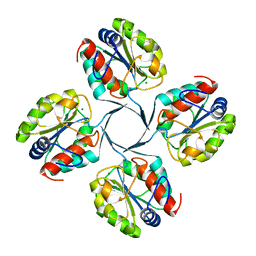 
 | | Crystal structure of YrbI from Escherichia coli in complex with Mg | | Descriptor: | 3-deoxy-D-manno-octulosonate 8-phosphate phosphatase, CHLORIDE ION, MAGNESIUM ION | | Authors: | Tsodikov, O.V, Aggarwal, P, Rubin, J.R, Stuckey, J.A, Woodard, R, Biswas, T. | | Deposit date: | 2007-09-10 | | Release date: | 2008-09-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The Tail of KdsC: CONFORMATIONAL CHANGES CONTROL THE ACTIVITY OF A HALOACID DEHALOGENASE SUPERFAMILY PHOSPHATASE.
J.Biol.Chem., 284, 2009
|
|
5ZVN
 | | Structure of [beta Glc-T9,K7]indolicidin, a glycosylated analogue of indolicidin | | Descriptor: | beta-D-glucopyranose, glycosylated analogue of Indolicidin | | Authors: | Dwivedi, R, Aggarwal, P, Kaur, K.J, Bhavesh, N.S. | | Deposit date: | 2018-05-11 | | Release date: | 2019-06-19 | | Last modified: | 2020-07-29 | | Method: | SOLUTION NMR | | Cite: | Design of therapeutically improved analogue of the antimicrobial peptide, indolicidin, using a glycosylation strategy.
Amino Acids, 51, 2019
|
|
5ZVF
 
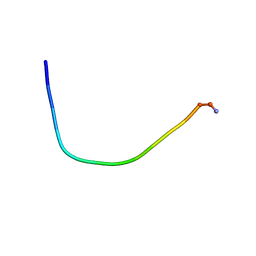 | | Structure of [T9,K7]indolicidin, a non glycosylated analogue of indolicidin | | Descriptor: | non glycosylated analogue of Indolicidin | | Authors: | Dwivedi, R, Aggarwal, P, Kaur, K.J, Bhavesh, N.S. | | Deposit date: | 2018-05-10 | | Release date: | 2019-06-19 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Design of therapeutically improved analogue of the antimicrobial peptide, indolicidin, using a glycosylation strategy.
Amino Acids, 51, 2019
|
|
