8RP4
 
 | | Alpha-Methylacyl-CoA racemase from Mycobacterium tuberculosis (D156A mutant) | | Descriptor: | 1,2-ETHANEDIOL, Alpha-methylacyl-CoA racemase, DI(HYDROXYETHYL)ETHER | | Authors: | Mojanaga, O.O, Acharya, K.R, Lloyd, M.D. | | Deposit date: | 2024-01-12 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | alpha-Methylacyl-CoA Racemase from Mycobacterium tuberculosis -Detailed Kinetic and Structural Characterization of the Active Site.
Biomolecules, 14, 2024
|
|
8RP3
 
 | |
8RP5
 
 | | Alpha-Methylacyl-CoA racemase from Mycobacterium tuberculosis (E241A mutant) | | Descriptor: | 1,2-ETHANEDIOL, Alpha-methylacyl-CoA racemase, TRIETHYLENE GLYCOL | | Authors: | Mojanaga, O.O, Acharya, K.R, Lloyd, M.D. | | Deposit date: | 2024-01-12 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | alpha-Methylacyl-CoA Racemase from Mycobacterium tuberculosis -Detailed Kinetic and Structural Characterization of the Active Site.
Biomolecules, 14, 2024
|
|
5ARJ
 
 | |
2J0T
 
 | | Crystal Structure of the Catalytic Domain of MMP-1 in Complex with the Inhibitory Domain of TIMP-1 | | Descriptor: | CALCIUM ION, INTERSTITIAL COLLAGENASE, METALLOPROTEINASE INHIBITOR 1, ... | | Authors: | Iyer, S, Wei, S, Brew, K, Acharya, K.R. | | Deposit date: | 2006-08-04 | | Release date: | 2006-10-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Crystal Structure of the Catalytic Domain of Matrix Metalloproteinase-1 in Complex with the Inhibitory Domain of Tissue Inhibitor of Metalloproteinase-1.
J.Biol.Chem., 282, 2007
|
|
2IVD
 
 | | Structure of protoporphyrinogen oxidase from Myxococcus xanthus with acifluorfen | | Descriptor: | (3S)-3-[(2S,3S,4R)-3,4-DIMETHYLTETRAHYDROFURAN-2-YL]BUTYL LAURATE, 5-[2-CHLORO-4-(TRIFLUOROMETHYL)PHENOXY]-2-NITROBENZOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Corradi, H.R, Corrigall, A.V, Boix, E, Mohan, C.G, Sturrock, E.D, Meissner, P.N, Acharya, K.R. | | Deposit date: | 2006-06-12 | | Release date: | 2006-10-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Protoporphyrinogen Oxidase from Myxococcus Xanthus and its Complex with the Inhibitor Acifluorfen.
J.Biol.Chem., 281, 2006
|
|
2YDM
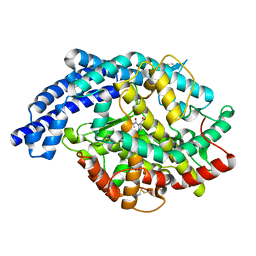 
 | | Structural characterization of angiotensin-I converting enzyme in complex with a selenium analogue of captopril | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ANGIOTENSIN CONVERTING ENZYME, CHLORIDE ION, ... | | Authors: | Akif, M, Masuyer, G, Schwager, S.L.U, Bhuyan, B.J, Mugesh, G, Sturrock, E.D, Acharya, K.R. | | Deposit date: | 2011-03-22 | | Release date: | 2011-09-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structural Characterization of Angiotensin-I Converting Enzyme in Complex with a Selenium Analogue of Captopril
FEBS J., 278, 2011
|
|
5AM9
 
 | | Crystal structure of the Angiotensin-1 converting enzyme N-domain in complex with amyloid-beta 10-16 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ANGIOTENSIN-CONVERTING ENZYME, CALCIUM ION, ... | | Authors: | Masuyer, G, Larmuth, K.M, Douglas, R.G, Sturrock, E.D, Acharya, K.R. | | Deposit date: | 2015-03-10 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Kinetic and Structural Characterisation of Amyloid-Beta Metabolism by Human Angiotensin-1- Converting Enzyme (Ace)
FEBS J., 283, 2016
|
|
2GAL
 
 | |
5ARL
 
 | |
5AMB
 
 | | Crystal structure of the Angiotensin-1 converting enzyme N-domain in complex with amyloid-beta 35-42 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, AMYLOID BETA A4 PROTEIN, ANGIOTENSIN-CONVERTING ENZYME, ... | | Authors: | Masuyer, G, Larmuth, K.M, Douglas, R.G, Sturrock, E.D, Acharya, K.R. | | Deposit date: | 2015-03-10 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The Kinetic and Structural Characterisation of Amyloid-Beta Metabolism by Human Angiotensin-1- Converting Enzyme (Ace)
FEBS J., 283, 2016
|
|
5AMC
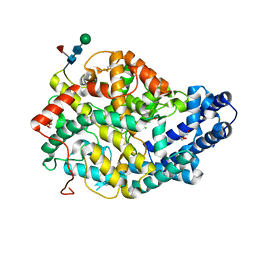 
 | | Crystal structure of the Angiotensin-1 converting enzyme N-domain in complex with amyloid-beta fluorogenic fragment 4-10 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ANGIOTENSIN-CONVERTING ENZYME, CHLORIDE ION, ... | | Authors: | Masuyer, G, Larmuth, K.M, Douglas, R.G, Sturrock, E.D, Acharya, K.R. | | Deposit date: | 2015-03-10 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The Kinetic and Structural Characterisation of Amyloid-Beta Metabolism by Human Angiotensin-1- Converting Enzyme (Ace)
FEBS J., 283, 2016
|
|
5AMA
 
 | | Crystal structure of the Angiotensin-1 converting enzyme N-domain in complex with amyloid-beta 1-16 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ANGIOTENSIN-CONVERTING ENZYME, ... | | Authors: | Masuyer, G, Larmuth, K.M, Douglas, R.G, Sturrock, E.D, Acharya, K.R. | | Deposit date: | 2015-03-10 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Kinetic and Structural Characterisation of Amyloid-Beta Metabolism by Human Angiotensin-1- Converting Enzyme (Ace)
FEBS J., 283, 2016
|
|
5AR6
 
 | | crystal structure of porcine RNase 4 | | Descriptor: | 1,2-ETHANEDIOL, RIBONUCLEASE 4 | | Authors: | Liang, S, Acharya, K.R. | | Deposit date: | 2015-09-24 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Structural Basis of Substrate Specificity in Porcine Rnase 4.
FEBS J., 283, 2016
|
|
2JCK
 
 | | Crystal structure of alpha-1,3 Galactosyltransferase (R365K) in complex with UDP and 2 manganese ion | | Descriptor: | MANGANESE (II) ION, N-ACETYLLACTOSAMINIDE ALPHA-1,3-GALACTOSYLTRANSFERASE, URIDINE-5'-DIPHOSPHATE | | Authors: | Jamaluddin, H, Tumbale, P, Withers, S.G, Acharya, K.R, Brew, K. | | Deposit date: | 2006-12-26 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Conformational Changes Induced by Binding Udp-2F-Galactose to Alpha-1,3 Galactosyltransferase-Implications for Catalysis.
J.Mol.Biol., 369, 2007
|
|
2JCL
 
 | | Crystal structure of alpha-1,3 Galactosyltransferase (R365K) in the absence of ligands | | Descriptor: | N-ACETYLLACTOSAMINIDE ALPHA-1,3-GALACTOSYLTRANSFERASE, SULFATE ION | | Authors: | Jamaluddin, H, Tumbale, P, Withers, S.G, Acharya, K.R, Brew, K. | | Deposit date: | 2006-12-26 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Conformational Changes Induced by Binding Udp-2F-Galactose to Alpha-1,3 Galactosyltransferase-Implications for Catalysis.
J.Mol.Biol., 369, 2007
|
|
2JCJ
 
 | | Crystal structure of alpha-1,3 Galactosyltransferase (C-terminus truncated mutant-C3) in complex with UDP and Tris | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLYCEROL, ... | | Authors: | Jamaluddin, H, Tumbale, P, Withers, S.G, Acharya, K.R, Brew, K. | | Deposit date: | 2006-12-25 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Conformational Changes Induced by Binding Udp-2F-Galactose to Alpha-1,3 Galactosyltransferase-Implications for Catalysis.
J.Mol.Biol., 369, 2007
|
|
2JCO
 
 | | Crystal structure of wild type alpha-1,3 Galactosyltransferase in the absence of ligands | | Descriptor: | GLYCEROL, N-ACETYLLACTOSAMINIDE ALPHA-1,3-GALACTOSYL TRANSFERASE | | Authors: | Jamaluddin, H, Tumbale, P, Withers, S.G, Acharya, K.R, Brew, K. | | Deposit date: | 2006-12-28 | | Release date: | 2007-06-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Conformational Changes Induced by Binding Udp-2F-Galactose to Alpha-1,3 Galactosyltransferase-Implications for Catalysis.
J.Mol.Biol., 369, 2007
|
|
1W4P
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | 2'-DEOXYURIDINE 3'-MONOPHOSPHATE, PANCREATIC RIBONUCLEASE A | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
1W4Q
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | 2'-FLUORO-2'-DEOXYURIDINE 3'-MONOPHOSPHATE, PANCREATIC RIBONUCLEASE A | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
1W4O
 
 | | Binding of Nonnatural 3'-Nucleotides to Ribonuclease A | | Descriptor: | PANCREATIC RIBONUCLEASE A, URACIL ARABINOSE-3'-PHOSPHATE | | Authors: | Jenkins, C.L, Thiyagarajan, N, Sweeney, R.Y, Guy, M.P, Kelemen, B.R, Acharya, K.R, Raines, R.T. | | Deposit date: | 2004-07-27 | | Release date: | 2005-02-02 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Binding of Non-Natural 3'-Nucleotides to Ribonuclease A
FEBS J., 272, 2005
|
|
1XPZ
 
 | | Structure of human carbonic anhydrase II with 4-[4-O-sulfamoylbenzyl)(4-cyanophenyl)amino]-4H-[1,2,4]-triazole | | Descriptor: | 4-{[(4-CYANOPHENYL)(4H-1,2,4-TRIAZOL-4-YL)AMINO]METHYL}PHENYL SULFAMATE, Carbonic anhydrase II, ZINC ION | | Authors: | Lloyd, M.D, Thiyagarajan, N, Ho, Y.T, Woo, L.W.L, Sutcliffe, O.B, Purohit, A, Reed, M.J, Acharya, K.R, Potter, B.V.L. | | Deposit date: | 2004-10-11 | | Release date: | 2005-05-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | First Crystal Structures of Human Carbonic Anhydrase II in Complex with Dual Aromatase-Steroid Sulfatase Inhibitors(,)
Biochemistry, 44, 2005
|
|
1XQ0
 
 | | Structure of human carbonic anhydrase II with 4-[(3-bromo-4-O-sulfamoylbenzyl)(4-cyanophenyl)amino]-4H-[1,2,4]-triazole | | Descriptor: | 2-BROMO-4-{[(4-CYANOPHENYL)(4H-1,2,4-TRIAZOL-4-YL)AMINO]METHYL}PHENYL SULFAMATE, Carbonic anhydrase II, ZINC ION | | Authors: | Lloyd, M.D, Thiyagarajan, N, Ho, Y.T, Woo, L.W.L, Sutcliffe, O.B, Purohit, A, Reed, M.J, Acharya, K.R, Potter, B.V.L. | | Deposit date: | 2004-10-11 | | Release date: | 2005-05-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | First Crystal Structures of Human Carbonic Anhydrase II in Complex with Dual Aromatase-Steroid Sulfatase Inhibitors(,)
Biochemistry, 44, 2005
|
|
1OJZ
 
 | | The crystal structure of C3stau2 from S. aureus with NAD | | Descriptor: | ADP-RIBOSYLTRANSFERASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Evans, H.R, Sutton, J.M, Holloway, D.E, Ayriss, J, Shone, C.C, Acharya, K.R. | | Deposit date: | 2003-07-16 | | Release date: | 2003-08-28 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | The Crystal Structure of C3Stau2 from Staphylococcus Aureus and its Complex with Nad
J.Biol.Chem., 278, 2003
|
|
1HI2
 
 | | Eosinophil-derived Neurotoxin (EDN) - Sulphate Complex | | Descriptor: | EOSINOPHIL-DERIVED NEUROTOXIN, SULFATE ION | | Authors: | Leonidas, D.D, Boix, E, Prill, R, Suzuki, M, Turton, R, Minson, K, Swaminathan, G.J, Youle, R.J, Acharya, K.R. | | Deposit date: | 2001-01-02 | | Release date: | 2001-05-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Mapping the Ribonucleolytic Active Site of Eosinophil-Derived Neurotoxin (Edn): High Resolution Crystal Structures of Edn Complexes with Adenylic Nucleotide Inhibitors
J.Biol.Chem., 276, 2001
|
|
