1AQ5
 
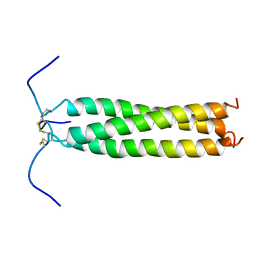 | | HIGH-RESOLUTION SOLUTION NMR STRUCTURE OF THE TRIMERIC COILED-COIL DOMAIN OF CHICKEN CARTILAGE MATRIX PROTEIN, 20 STRUCTURES | | Descriptor: | CARTILAGE MATRIX PROTEIN | | Authors: | Dames, S.A, Wiltscheck, R, Kammerer, R.A, Engel, J, Alexandrescu, A.T. | | Deposit date: | 1997-08-07 | | Release date: | 1998-02-11 | | Last modified: | 2021-01-13 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a parallel homotrimeric coiled coil.
Nat.Struct.Biol., 5, 1998
|
|
1HCE
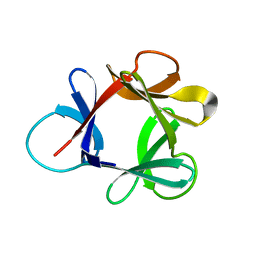 
 | | STRUCTURE OF HISACTOPHILIN IS SIMILAR TO INTERLEUKIN-1 BETA AND FIBROBLAST GROWTH FACTOR | | Descriptor: | HISACTOPHILIN | | Authors: | Habazettl, J, Gondol, D, Wiltscheck, R, Otlewski, J, Schleicher, M, Holak, T.A. | | Deposit date: | 1994-07-12 | | Release date: | 1994-09-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of hisactophilin is similar to interleukin-1 beta and fibroblast growth factor.
Nature, 359, 1992
|
|
1HCD
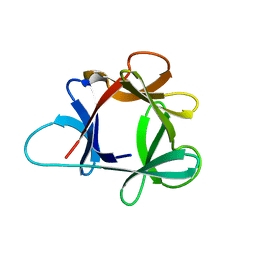 
 | | STRUCTURE OF HISACTOPHILIN IS SIMILAR TO INTERLEUKIN-1 BETA AND FIBROBLAST GROWTH FACTOR | | Descriptor: | HISACTOPHILIN | | Authors: | Habazettl, J, Gondol, D, Wiltscheck, R, Otlewski, J, Schleicher, M, Holak, T.A. | | Deposit date: | 1994-05-03 | | Release date: | 1994-10-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of hisactophilin is similar to interleukin-1 beta and fibroblast growth factor.
Nature, 359, 1992
|
|
1BIP
 
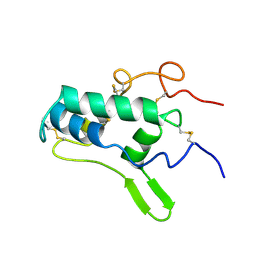 | |
1NMF
 
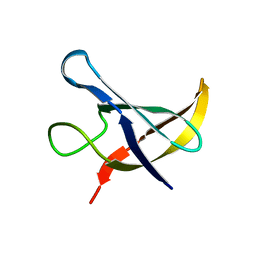 | |
1NMG
 
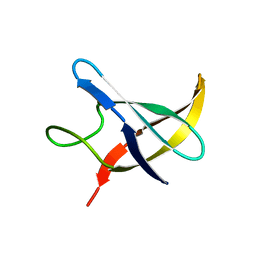 | |
1SVR
 
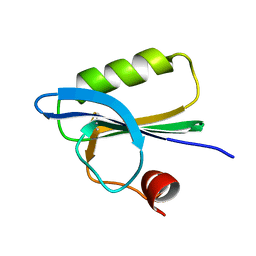 | |
1SVQ
 
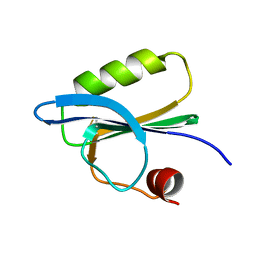 | |
1B1U
 
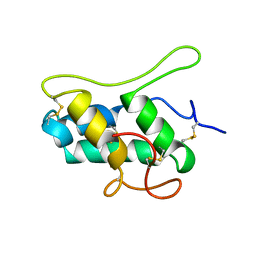 | |
1SVY
 
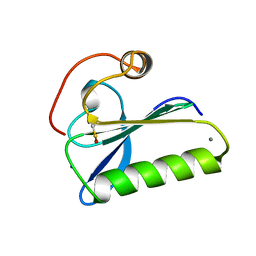 | | SEVERIN DOMAIN 2, 1.75 ANGSTROM CRYSTAL STRUCTURE | | Descriptor: | CALCIUM ION, SEVERIN, SODIUM ION | | Authors: | Puius, Y.A, Fedorov, E.V, Eichinger, L, Sullivan, M, Schleicher, M, Almo, S.C. | | Deposit date: | 1998-08-10 | | Release date: | 1999-08-10 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Mapping the functional surface of domain 2 in the gelsolin superfamily.
Biochemistry, 39, 2000
|
|
