1UJJ
 
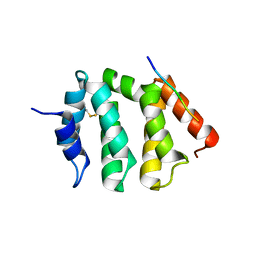 | | VHS domain of human GGA1 complexed with C-terminal peptide from BACE | | 分子名称: | ADP-ribosylation factor binding protein GGA1, C-terminal peptide from Beta-secretase | | 著者 | Shiba, T, Kametaka, S, Kawasaki, M, Shibata, M, Waguri, S, Uchiyama, Y, Wakatsuki, S. | | 登録日 | 2003-08-05 | | 公開日 | 2004-05-11 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Insights into the Phosphoregulation of beta-Secretase Sorting Signal by the VHS Domain of GGA1
TRAFFIC, 5, 2004
|
|
1UJK
 
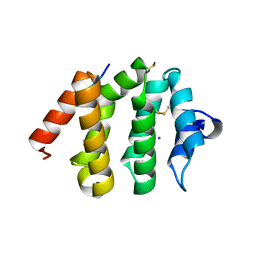 | | VHS domain of human GGA1 complexed with C-terminal phosphopeptide from BACE | | 分子名称: | ADP-ribosylation factor binding protein GGA1, C-terminal peptide from Beta-secretase, IODIDE ION | | 著者 | Shiba, T, Kametaka, S, Kawasaki, M, Shibata, M, Waguri, S, Uchiyama, Y, Wakatsuki, S. | | 登録日 | 2003-08-05 | | 公開日 | 2004-05-11 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Insights into the Phosphoregulation of beta-Secretase Sorting Signal by the VHS Domain of GGA1
TRAFFIC, 5, 2004
|
|
4ZY3
 
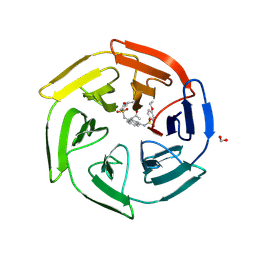 | | Crystal Structure of Keap1 in Complex with a small chemical compound, K67 | | 分子名称: | FORMIC ACID, Kelch-like ECH-associated protein 1, N,N'-[2-(2-oxopropyl)naphthalene-1,4-diyl]bis(4-ethoxybenzenesulfonamide) | | 著者 | Fukutomi, T, Iso, T, Suzuki, T, Takagi, K, Mizushima, T, Komatsu, M, Yamamoto, M. | | 登録日 | 2015-05-21 | | 公開日 | 2016-05-25 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | p62/Sqstm1 promotes malignancy of HCV-positive hepatocellular carcinoma through Nrf2-dependent metabolic reprogramming
Nat Commun, 7, 2016
|
|
3ADE
 
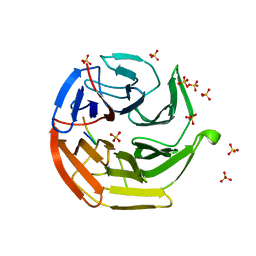 | |
3WDZ
 
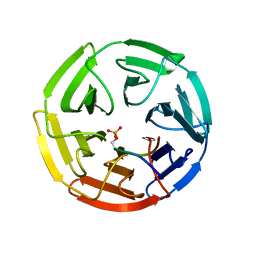 | | Crystal Structure of Keap1 in Complex with phosphorylated p62 | | 分子名称: | Kelch-like ECH-associated protein 1, Peptide from Sequestosome-1 | | 著者 | Fukutomi, T, Takagi, K, Mizushima, T, Tanaka, K, Komatsu, M, Yamamoto, M. | | 登録日 | 2013-06-26 | | 公開日 | 2013-09-04 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Phosphorylation of p62 activates the Keap1-Nrf2 pathway during selective autophagy.
Mol.Cell, 51, 2013
|
|
7W3N
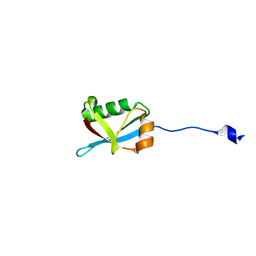 
 | |
7W3O
 
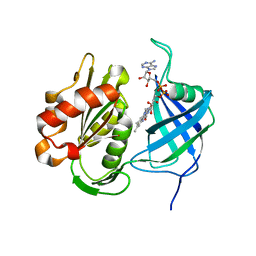 | | Crystal structure of human CYB5R3 | | 分子名称: | FLAVIN-ADENINE DINUCLEOTIDE, NADH-cytochrome b5 reductase 3 soluble form | | 著者 | Noda, N.N. | | 登録日 | 2021-11-25 | | 公開日 | 2022-12-07 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.46 Å) | | 主引用文献 | The UFM1 system regulates ER-phagy through the ufmylation of CYB5R3.
Nat Commun, 13, 2022
|
|
7P6T
 
 | | Crystal structure of the FimH-binding decoy module of human glycoprotein 2 (GP2) (crystal form III) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-ethyl-2-(hydroxymethyl)propane-1,3-diol, Isoform Alpha of Pancreatic secretory granule membrane major glycoprotein GP2 | | 著者 | Stsiapanava, A, Tunyasuvunakool, K, Jumper, J, de Sanctis, D, Jovine, L. | | 登録日 | 2021-07-17 | | 公開日 | 2022-03-16 | | 最終更新日 | 2022-03-30 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Structure of the decoy module of human glycoprotein 2 and uromodulin and its interaction with bacterial adhesin FimH.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7P6R
 
 | | Crystal structure of the FimH-binding decoy module of human glycoprotein 2 (GP2) (crystal form I) | | 分子名称: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Isoform Alpha of Pancreatic secretory granule membrane major glycoprotein GP2 | | 著者 | Stsiapanava, A, Tunyasuvunakool, K, Jumper, J, de Sanctis, D, Jovine, L. | | 登録日 | 2021-07-17 | | 公開日 | 2022-03-16 | | 最終更新日 | 2022-03-30 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Structure of the decoy module of human glycoprotein 2 and uromodulin and its interaction with bacterial adhesin FimH.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7P6S
 
 | | Crystal structure of the FimH-binding decoy module of human glycoprotein 2 (GP2) (crystal form II) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Isoform Alpha of Pancreatic secretory granule membrane major glycoprotein GP2, pentane-1,5-diol | | 著者 | Stsiapanava, A, Tunyasuvunakool, K, Jumper, J, de Sanctis, D, Jovine, L. | | 登録日 | 2021-07-17 | | 公開日 | 2022-03-16 | | 最終更新日 | 2022-03-30 | | 実験手法 | X-RAY DIFFRACTION (1.35 Å) | | 主引用文献 | Structure of the decoy module of human glycoprotein 2 and uromodulin and its interaction with bacterial adhesin FimH.
Nat.Struct.Mol.Biol., 29, 2022
|
|
