4HX3
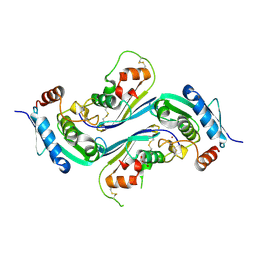 
 | | Crystal structure of Streptomyces caespitosus sermetstatin in complex with S. caespitosus snapalysin | | Descriptor: | Extracellular small neutral protease, GLYCEROL, Neutral proteinase inhibitor ScNPI, ... | | Authors: | Trillo-Muyo, S, Martinez-Rodriguez, S, Arolas, J.L, Gomis-Ruth, F.X. | | Deposit date: | 2012-11-09 | | Release date: | 2012-12-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Mechanism of action of a Janus-faced single-domain protein inhibitor simultaneously targeting two peptidase classes
CHEM SCI, 4, 2013
|
|
4HX2
 
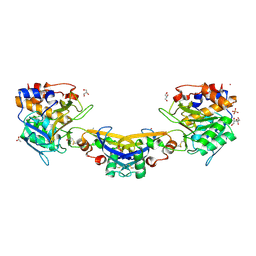 | | Crystal structure of Streptomyces caespitosus sermetstatin in complex with Bacillus licheniformis subtilisin | | Descriptor: | (2R,2'R)-3,3'-oxydipropane-1,2-diol, ACETATE ION, CACODYLATE ION, ... | | Authors: | Trillo-Muyo, S, Martinez-Rodriguez, S, Arolas, J.L, Gomis-Ruth, F.X. | | Deposit date: | 2012-11-09 | | Release date: | 2012-12-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Mechanism of action of a Janus-faced single-domain protein inhibitor simultaneously targeting two peptidase classes
CHEM SCI, 4, 2013
|
|
4HWX
 
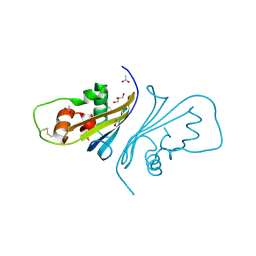 | | Crystal structure of Streptomyces caespitosus sermetstatin | | Descriptor: | ACETATE ION, GLYCEROL, Neutral proteinase inhibitor ScNPI | | Authors: | Trillo-Muyo, S, Martinez-Rodriguez, S, Arolas, J.L, Gomis-Ruth, F.X. | | Deposit date: | 2012-11-09 | | Release date: | 2012-12-05 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanism of action of a Janus-faced single-domain protein inhibitor simultaneously targeting two peptidase classes
CHEM SCI, 4, 2013
|
|
4HE5
 
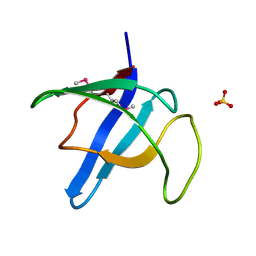 | | Crystal structure of the selenomethionine variant of the C-terminal domain of Geobacillus thermoleovorans putative U32 peptidase | | Descriptor: | Peptidase family U32, SULFATE ION | | Authors: | Trillo-Muyo, S, Jasilionis, A, Domagalski, M.J, Chruszcz, M, Minor, W, Kuisiene, N, Arolas, J.L, Sola, M, Gomis-Ruth, F.X. | | Deposit date: | 2012-10-03 | | Release date: | 2012-11-14 | | Last modified: | 2022-04-13 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Ultratight crystal packing of a 10 kDa protein.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4HE6
 
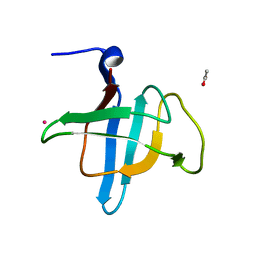 | | Crystal structure of the C-terminal domain of Geobacillus thermoleovorans putative U32 peptidase | | Descriptor: | ACETATE ION, Peptidase family U32, UNKNOWN ATOM OR ION | | Authors: | Trillo-Muyo, S, Jasilionis, A, Domagalski, M.J, Chruszcz, M, Minor, W, Kuisiene, N, Arolas, J.L, Sola, M, Gomis-Ruth, F.X. | | Deposit date: | 2012-10-03 | | Release date: | 2012-11-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Ultratight crystal packing of a 10 kDa protein.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
8S03
 
 | | NMR solution structure of the CysD2 domain of MUC2 | | Descriptor: | CALCIUM ION, Mucin-2 | | Authors: | Recktenwald, C, Karlsson, B.G, Garcia-Bonnete, M.-J, Katona, G, Jensen, M, Lymer, R, Baeckstroem, M, Johansson, M.E.V, Hansson, G.C, Trillo-Muyo, S. | | Deposit date: | 2024-02-13 | | Release date: | 2024-04-17 | | Method: | SOLUTION NMR | | Cite: | The second CysD domain of MUC2 and role in mucin organization by transglutaminase-based cross-linking
To Be Published
|
|
4IN9
 
 | | Structure of karilysin MMP-like catalytic domain in complex with inhibitory tetrapeptide SWFP | | Descriptor: | GLYCEROL, Karilysin protease, POTASSIUM ION, ... | | Authors: | Guevara, T, Ksiazek, M, Skottrup, P.D, Cerda-Costa, N, Trillo-Muyo, S, de Diego, I, Riise, E, Potempa, J, Gomis-Ruth, F.X. | | Deposit date: | 2013-01-04 | | Release date: | 2013-05-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure of the catalytic domain of the Tannerella forsythia matrix metallopeptidase karilysin in complex with a tetrapeptidic inhibitor.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
8AVX
 
 | |
8AVW
 
 | | Cryo-EM structure of DrBphP in Pr state | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome,Response regulator | | Authors: | Wahlgren, W.Y, Takala, H, Westenhoff, S. | | Deposit date: | 2022-08-27 | | Release date: | 2022-12-21 | | Last modified: | 2022-12-28 | | Method: | ELECTRON MICROSCOPY (3.62 Å) | | Cite: | Structural mechanism of signal transduction in a phytochrome histidine kinase.
Nat Commun, 13, 2022
|
|
8AVV
 
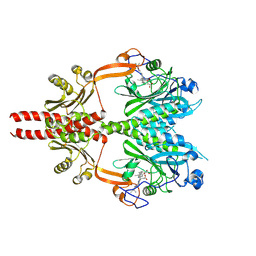 | | Cryo-EM structure of DrBphP photosensory module in Pr state | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome,Response regulator | | Authors: | Wahlgren, W.Y, Takala, H, Westenhoff, S. | | Deposit date: | 2022-08-27 | | Release date: | 2022-12-21 | | Last modified: | 2022-12-28 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural mechanism of signal transduction in a phytochrome histidine kinase.
Nat Commun, 13, 2022
|
|
7ZAO
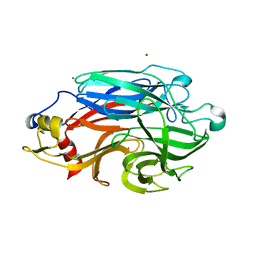 
 | | Structure of BPP43_05035 of Brachyspira pilosicoli | | Descriptor: | MAGNESIUM ION, Sialidase (Neuraminidase) family protein-like protein | | Authors: | Rajan, A, Gallego, P, Pelaseyed, T. | | Deposit date: | 2022-03-22 | | Release date: | 2023-10-11 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | BPP43_05035 is a Brachyspira pilosicoli cell surface adhesin that weakens the integrity of the epithelial barrier during infection
Biorxiv, 2024
|
|
7QCU
 
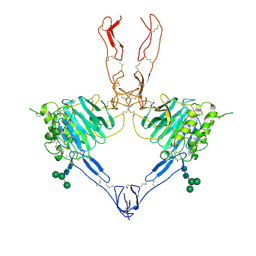 | | Structure of the MUCIN-2 Cterminal domains partially deglycosylated. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Mucin-2, ... | | Authors: | Gallego, P, Hansson, G.C. | | Deposit date: | 2021-11-25 | | Release date: | 2023-03-08 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.25 Å) | | Cite: | The intestinal MUC2 mucin C-terminus is stabilized by an extra disulfide bond in comparison to von Willebrand factor and other gel-forming mucins.
Nat Commun, 14, 2023
|
|
7QCN
 
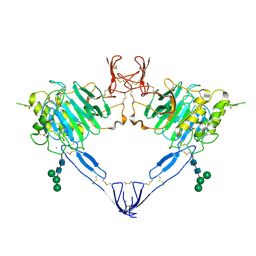 | |
7QCL
 
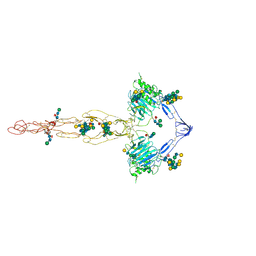 | | Structure of the MUCIN-2 Cterminal domains | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Mucin-2, ... | | Authors: | Gallego, P, Hansson, G.C. | | Deposit date: | 2021-11-24 | | Release date: | 2023-03-08 | | Last modified: | 2023-09-27 | | Method: | ELECTRON MICROSCOPY (3.36 Å) | | Cite: | The intestinal MUC2 mucin C-terminus is stabilized by an extra disulfide bond in comparison to von Willebrand factor and other gel-forming mucins.
Nat Commun, 14, 2023
|
|
