7NH9
 
 | |
6GYF
 
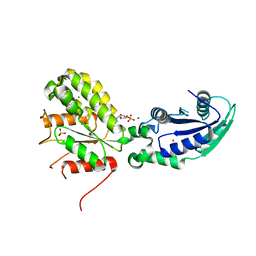 | | Crystal structure of NadR protein in complex with NR | | Descriptor: | BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE, MAGNESIUM ION, Nicotinamide-nucleotide adenylyltransferase NadR family / Ribosylnicotinamide kinase, ... | | Authors: | Singh, R, Stetsenko, A, Jaehme, M, Guskov, A, Slotboom, D.J. | | Deposit date: | 2018-06-29 | | Release date: | 2019-07-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural and Functional Characterization of NadR fromLactococcus lactis.
Molecules, 25, 2020
|
|
6GYE
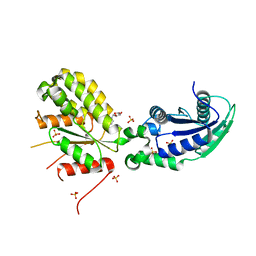 
 | | Crystal structure of NadR protein in complex with NR | | Descriptor: | Nicotinamide riboside, Nicotinamide-nucleotide adenylyltransferase NadR family / Ribosylnicotinamide kinase, SULFATE ION | | Authors: | Singh, R, Stetsenko, A, Jaehme, M, Guskov, A, Slotboom, D.J. | | Deposit date: | 2018-06-29 | | Release date: | 2019-07-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and Functional Characterization of NadR fromLactococcus lactis.
Molecules, 25, 2020
|
|
6GZO
 
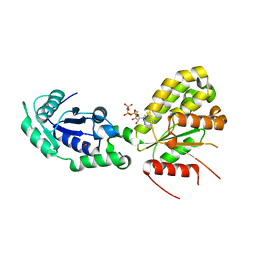 | | Crystal structure of NadR protein in complex with NAD and AMP-PNP | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Nicotinamide-nucleotide adenylyltransferase NadR family / Ribosylnicotinamide kinase, PHOSPHATE ION, ... | | Authors: | Singh, R, Stetsenko, A, Jaehme, M, Guskov, A, Slotboom, D.J. | | Deposit date: | 2018-07-04 | | Release date: | 2019-07-17 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and Functional Characterization of NadR fromLactococcus lactis.
Molecules, 25, 2020
|
|
7P4I
 
 | |
9X67
 
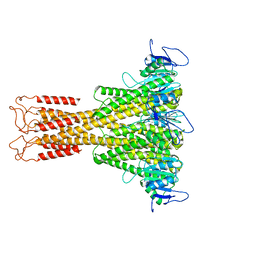
 | | Cryo-EM structure of the type I pilus from Escherichia Coli and the surrounding water network | | Descriptor: | Type-1 fimbrial protein, A chain | | Authors: | Petrova, T.E, Glukhov, A.S, Stetsenko, A, Guskov, A, Gabdulkhakov, A.G. | | Deposit date: | 2025-10-15 | | Release date: | 2026-03-11 | | Method: | ELECTRON MICROSCOPY (1.85 Å) | | Cite: | Determination of the water network surrounding the type I pilus from Escherichia coli by cryo-electron microscopy.
Febs J., 2026
|
|
5N9Y
 
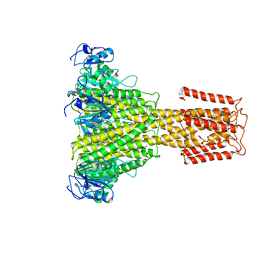 | | The full-length structure of ZntB | | Descriptor: | Zinc transport protein ZntB | | Authors: | Cornelius, G, Stetsenko, A, Scheres, S.H.W, Slotboom, D.J, Guskov, A. | | Deposit date: | 2017-02-27 | | Release date: | 2017-11-15 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | The structural basis of proton driven zinc transport by ZntB.
Nat Commun, 8, 2017
|
|
8BYV
 
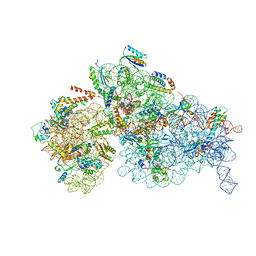 | | Cryo-EM structure of a Staphylococus aureus 30S-RbfA complex | | Descriptor: | 16S ribosomal RNA, 30S ribosomal protein S10, 30S ribosomal protein S11, ... | | Authors: | Bikmullin, A.G, Fatkhullin, B, Stetsenko, A, Guskov, A, Yusupov, M. | | Deposit date: | 2022-12-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (2.89 Å) | | Cite: | Yet Another Similarity between Mitochondrial and Bacterial Ribosomal Small Subunit Biogenesis Obtained by Structural Characterization of RbfA from S. aureus.
Int J Mol Sci, 24, 2023
|
|
9R22
 
 | | Cryo-EM structure of the light-driven proton pump PsPR in detergent micelle | | Descriptor: | DODECYL-BETA-D-MALTOSIDE, EICOSANE, RETINAL, ... | | Authors: | Kovalev, K, Stetsenko, A, Guskov, A. | | Deposit date: | 2025-04-29 | | Release date: | 2025-07-23 | | Last modified: | 2025-09-17 | | Method: | ELECTRON MICROSCOPY (2.48 Å) | | Cite: | Structural basis for no retinal binding in flotillin-associated rhodopsins.
Structure, 33, 2025
|
|
9R23
 
 | |
9R21
 
 | | Cryo-EM structure of the flotillin-associated rhodopsin PsFAR in detergent micelle | | Descriptor: | EICOSANE, flotillin-associated rhodopsin | | Authors: | Kovalev, K, Stetsenko, A, Marin, E, Guskov, A. | | Deposit date: | 2025-04-29 | | Release date: | 2025-07-23 | | Last modified: | 2025-09-17 | | Method: | ELECTRON MICROSCOPY (2.56 Å) | | Cite: | Structural basis for no retinal binding in flotillin-associated rhodopsins.
Structure, 33, 2025
|
|
9G1Z
 
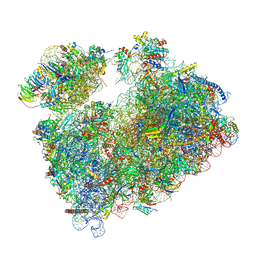 | | Structure of Candida albicans 80S ribosome in complex with mefloquine (non-rotated state) | | Descriptor: | (11R,12S)- Mefloquine, 18S rRNA, 25S rRNA, ... | | Authors: | Kolosova, O, Zgadzay, Y, Stetsenko, A, Atamas, A, Jenner, L.B, Guskov, A, Yusupov, M. | | Deposit date: | 2024-07-10 | | Release date: | 2025-04-23 | | Last modified: | 2025-05-07 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Mechanism of read-through enhancement by aminoglycosides and mefloquine.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8Q5I
 
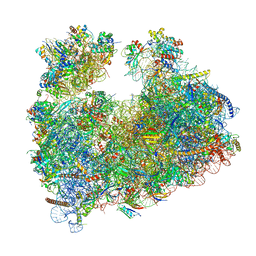 | | Structure of Candida albicans 80S ribosome in complex with cephaeline | | Descriptor: | 18S ribosomal RNA, 25S rRNA, 40S ribosomal protein S0, ... | | Authors: | Kolosova, O, Zgadzay, Y, Stetsenko, A, Atamas, A, Guskov, A, Yusupov, M. | | Deposit date: | 2023-08-09 | | Release date: | 2023-09-13 | | Last modified: | 2024-01-31 | | Method: | ELECTRON MICROSCOPY (2.45 Å) | | Cite: | Structural characterization of cephaeline binding to the eukaryotic ribosome using Cryo-Electron Microscopy
Biopolym Cell, 2023
|
|
8QQZ
 
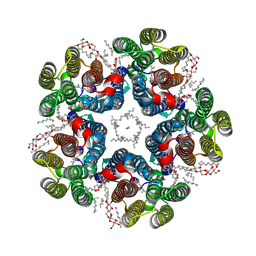 | | Cryo-EM structure of the light-driven sodium pump ErNaR in the pentameric form at pH 8.0 | | Descriptor: | Bacteriorhodopsin-like protein, DODECYL-BETA-D-MALTOSIDE, EICOSANE | | Authors: | Kovalev, K, Podoliak, E, Lamm, G.H.U, Marin, E, Stetsenko, A, Guskov, A. | | Deposit date: | 2023-10-06 | | Release date: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.63 Å) | | Cite: | A subgroup of light-driven sodium pumps with an additional Schiff base counterion.
Nat Commun, 15, 2024
|
|
8QR0
 
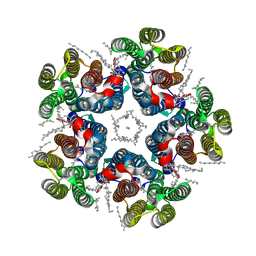 | | Cryo-EM structure of the light-driven sodium pump ErNaR in the pentameric form at pH 4.3 | | Descriptor: | Bacteriorhodopsin-like protein, DODECYL-BETA-D-MALTOSIDE, EICOSANE | | Authors: | Kovalev, K, Podoliak, E, Lamm, G.H.U, Marin, E, Stetsenko, A, Guskov, A. | | Deposit date: | 2023-10-06 | | Release date: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | A subgroup of light-driven sodium pumps with an additional Schiff base counterion.
Nat Commun, 15, 2024
|
|
8R0M
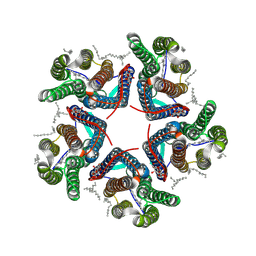 
 | | Cryo-EM structure of the microbial rhodopsin CryoR1 at pH 8.0 in detergent | | Descriptor: | DODECYL-BETA-D-MALTOSIDE, EICOSANE, RETINAL, ... | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.87 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
8R0N
 
 | | Cryo-EM structure of the microbial rhodopsin CryoR1 at pH 10.5 in detergent in the ground state | | Descriptor: | DODECYL-BETA-D-MALTOSIDE, EICOSANE, RETINAL, ... | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
8R0L
 
 | | Cryo-EM structure of the microbial rhodopsin CryoR1 at pH 8.0 in nanodisc | | Descriptor: | EICOSANE, RETINAL, Rhodopsin | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.43 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
8R0O
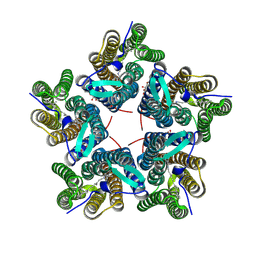 
 | | Cryo-EM structure of the microbial rhodopsin CryoR1 at pH 10.5 in detergent in the M state | | Descriptor: | DODECYL-BETA-D-MALTOSIDE, RETINAL, Rhodopsin | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
8R0K
 
 | | Cryo-EM structure of the microbial rhodopsin CryoR1 at pH 4.3 in detergent | | Descriptor: | EICOSANE, RETINAL, Rhodopsin | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.94 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
8R0P
 
 | | Cryo-EM structure of the microbial rhodopsin CryoR2 at pH 8.0 in detergent | | Descriptor: | EICOSANE, RETINAL, microbial rhodopsin CryoR2 | | Authors: | Kovalev, K, Marin, E, Stetsenko, A, Guskov, A, Lamm, G.H.U. | | Deposit date: | 2023-10-31 | | Release date: | 2025-05-14 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.44 Å) | | Cite: | CryoRhodopsins: A comprehensive characterization of a group of microbial rhodopsins from cold environments.
Sci Adv, 11, 2025
|
|
7Q0F
 
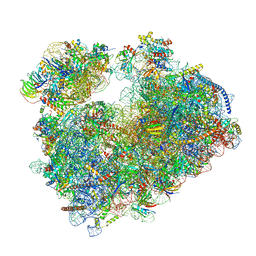 | | Structure of Candida albicans 80S ribosome in complex with phyllanthoside | | Descriptor: | 18S ribosomal RNA, 25S ribosomal RNA, 3-O-acetyl-2-O-(3-O-acetyl-6-deoxy-beta-D-glucopyranosyl)-6-deoxy-1-O-{[(2R,2'S,3a'R,4''S,5''R,6'S,7a'S)-5''-methyl-4''-{[(2E)-3-phenylprop-2-enoyl]oxy}decahydrodispiro[oxirane-2,3'-[1]benzofuran-2',2''-pyran]-6'-yl]carbonyl}-beta-D-glucopyranose, ... | | Authors: | Zgadzay, Y, Kolosova, O, Stetsenko, A, Jenner, L, Guskov, A, Yusupova, G, Yusupov, M. | | Deposit date: | 2021-10-14 | | Release date: | 2022-05-18 | | Last modified: | 2022-06-08 | | Method: | ELECTRON MICROSCOPY (2.64 Å) | | Cite: | E-site drug specificity of the human pathogen Candida albicans ribosome.
Sci Adv, 8, 2022
|
|
7Q08
 
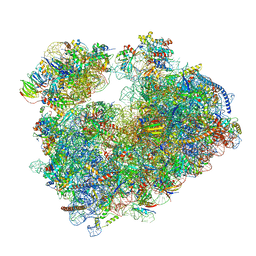 | | Structure of Candida albicans 80S ribosome in complex with cycloheximide | | Descriptor: | 18S ribosomal RNA, 25S ribosomal RNA, 4-{(2R)-2-[(1S,3S,5S)-3,5-dimethyl-2-oxocyclohexyl]-2-hydroxyethyl}piperidine-2,6-dione, ... | | Authors: | Zgadzay, Y, Kolosova, O, Stetsenko, A, Jenner, L, Guskov, A, Yusupova, G, Yusupov, M. | | Deposit date: | 2021-10-14 | | Release date: | 2022-05-25 | | Last modified: | 2022-06-08 | | Method: | ELECTRON MICROSCOPY (2.56 Å) | | Cite: | E-site drug specificity of the human pathogen Candida albicans ribosome.
Sci Adv, 8, 2022
|
|
7Q0R
 
 | | Structure of the Candida albicans 80S ribosome in complex with blasticidin s | | Descriptor: | 18S ribosomal RNA, 25S ribosomal RNA, 40S ribosomal protein S0, ... | | Authors: | Kolosova, O, Zgadzay, Y, Stetsenko, A, Jenner, L, Guskov, A, Yusupova, G, Yusupov, M. | | Deposit date: | 2021-10-16 | | Release date: | 2022-05-25 | | Last modified: | 2022-06-08 | | Method: | ELECTRON MICROSCOPY (2.67 Å) | | Cite: | E-site drug specificity of the human pathogen Candida albicans ribosome.
Sci Adv, 8, 2022
|
|
7PZY
 
 | | Structure of the vacant Candida albicans 80S ribosome | | Descriptor: | 1,4-DIAMINOBUTANE, 18S ribosomal RNA, 25S ribosomal RNA, ... | | Authors: | Zgadzay, Y, Kolosova, O, Stetsenko, A, Jenner, L, Guskov, A, Yusupova, G, Yusupov, M. | | Deposit date: | 2021-10-13 | | Release date: | 2022-05-18 | | Last modified: | 2022-06-08 | | Method: | ELECTRON MICROSCOPY (2.32 Å) | | Cite: | E-site drug specificity of the human pathogen Candida albicans ribosome.
Sci Adv, 8, 2022
|
|
