4V4P
 
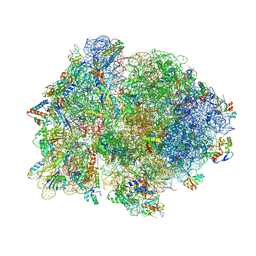 | | Crystal structure of 70S ribosome with thrS operator and tRNAs. | | Descriptor: | 16S rRNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Jenner, L, Romby, P, Rees, B, Schulze-Briese, C, Springer, M, Ehresmann, C, Ehresmann, B, Moras, D, Yusupova, G, Yusupov, M. | | Deposit date: | 2005-01-19 | | Release date: | 2014-07-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (5.5 Å) | | Cite: | Translational operator of mRNA on the ribosome: how repressor proteins exclude ribosome binding.
Science, 308, 2005
|
|
2GHJ
 
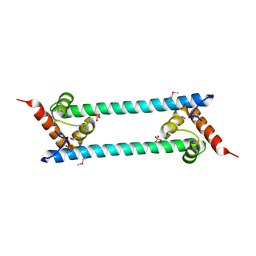 | | Crystal structure of folded and partially unfolded forms of Aquifex aeolicus ribosomal protein L20 | | Descriptor: | 50S ribosomal protein L20, SULFATE ION | | Authors: | Timsit, Y, Allemand, F, Chiaruttini, C, Springer, M. | | Deposit date: | 2006-03-27 | | Release date: | 2006-04-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Coexistence of two protein folding states in the crystal structure of ribosomal protein L20
Embo Rep., 7, 2006
|
|
1KOG
 
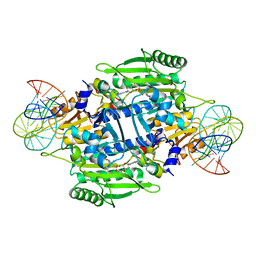 | | Crystal structure of E. coli threonyl-tRNA synthetase interacting with the essential domain of its mRNA operator | | Descriptor: | 5'-O-(N-(L-THREONYL)-SULFAMOYL)ADENOSINE, Threonyl-tRNA synthetase, Threonyl-tRNA synthetase mRNA, ... | | Authors: | Torres-Larrios, A, Dock-Bregeon, A.C, Romby, P, Rees, B, Sankaranarayanan, R, Caillet, J, Springer, M, Ehresmann, C, Ehresmann, B, Moras, D. | | Deposit date: | 2001-12-20 | | Release date: | 2002-04-26 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural basis of translational control by Escherichia coli threonyl tRNA synthetase.
Nat.Struct.Biol., 9, 2002
|
|
1FYF
 
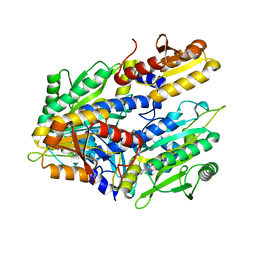 | | CRYSTAL STRUCTURE OF A TRUNCATED FORM OF THREONYL-TRNA SYNTHETASE COMPLEXED WITH A SERYL ADENYLATE ANALOG | | Descriptor: | 5'-O-(N-(L-SERYL)-SULFAMOYL)ADENOSINE, THREONYL-TRNA SYNTHETASE, ZINC ION | | Authors: | Sankaranarayanan, R, Dock-Bregeon, A.C, Moras, D. | | Deposit date: | 2000-09-29 | | Release date: | 2000-12-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Transfer RNA-mediated editing in threonyl-tRNA synthetase. The class II solution to the double discrimination problem.
Cell(Cambridge,Mass.), 103, 2000
|
|
9QD7
 
 | | The FK1 domain of FKBP51 in complex with the macrocyclic SAFit analog 15a | | Descriptor: | (2~{S},9~{S})-2-cyclohexyl-19,22-dimethoxy-11,14,17-trioxa-4-azatricyclo[16.2.2.0^{4,9}]docosa-1(21),18(22),19-triene-3,10-dione, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Brudy, C, Hausch, F. | | Deposit date: | 2025-03-06 | | Release date: | 2025-12-03 | | Last modified: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (2.264 Å) | | Cite: | Linker Modification Enables Control of Key Functional Group Orientation in Macrocycles.
J.Med.Chem., 68, 2025
|
|
9QD8
 
 | | The FK1 domain of FKBP51 in complex with the macrocyclic SAFit analog 29a | | Descriptor: | (2~{S},9~{S},12~{S})-2-cyclohexyl-19,22-dimethoxy-12-methyl-11,14,17-trioxa-4-azatricyclo[16.2.2.0^{4,9}]docosa-1(21),18(22),19-triene-3,10-dione, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Brudy, C, Hausch, F. | | Deposit date: | 2025-03-06 | | Release date: | 2025-12-03 | | Last modified: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Linker Modification Enables Control of Key Functional Group Orientation in Macrocycles.
J.Med.Chem., 68, 2025
|
|
9QD9
 
 | | The FK1 domain of FKBP51 in complex with the macrocyclic SAFit analog 29c | | Descriptor: | (2~{S},9~{S},12~{S})-2-cyclohexyl-20,23-dimethoxy-12-methyl-11,14,18-trioxa-4-azatricyclo[17.2.2.0^{4,9}]tricosa-1(22),19(23),20-triene-3,10-dione, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Brudy, C, Hausch, F. | | Deposit date: | 2025-03-06 | | Release date: | 2025-12-03 | | Last modified: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Linker Modification Enables Control of Key Functional Group Orientation in Macrocycles.
J.Med.Chem., 68, 2025
|
|
9QDA
 
 | | The FK1 domain of FKBP51 in complex with the macrocyclic SAFit analog 29b | | Descriptor: | (2~{S},9~{S},12~{R})-2-cyclohexyl-19,22-dimethoxy-12-methyl-11,14,17-trioxa-4-azatricyclo[16.2.2.0^{4,9}]docosa-1(21),18(22),19-triene-3,10-dione, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Brudy, C, Hausch, F. | | Deposit date: | 2025-03-06 | | Release date: | 2025-12-03 | | Last modified: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Linker Modification Enables Control of Key Functional Group Orientation in Macrocycles.
J.Med.Chem., 68, 2025
|
|
9QDB
 
 | | The FK1 domain of FKBP51 in complex with the macrocyclic SAFit analog 29d | | Descriptor: | (2~{S},9~{S},12~{R})-2-cyclohexyl-20,23-dimethoxy-12-methyl-11,14,18-trioxa-4-azatricyclo[17.2.2.0^{4,9}]tricosa-1(22),19(23),20-triene-3,10-dione, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Brudy, C, Hausch, F. | | Deposit date: | 2025-03-06 | | Release date: | 2025-12-03 | | Last modified: | 2025-12-24 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Linker Modification Enables Control of Key Functional Group Orientation in Macrocycles.
J.Med.Chem., 68, 2025
|
|
8CCA
 
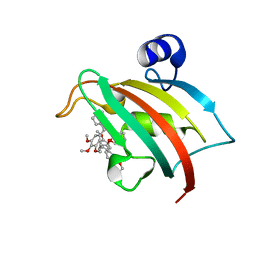 | | The Fk1 domain of FKBP51 in complex with SAFit1 | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-cyclohexyl-2-(3,4,5-trimethoxyphenyl)ethanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCD
 
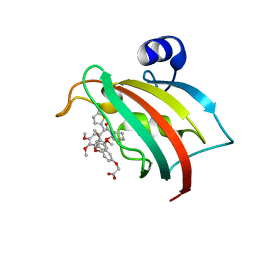 | | The Fk1 domain of FKBP51 in complex with 2-(3-((R)-1-(((S)-1-((S)-2-(5-chlorothiophen-2-yl)-2-(3,4,5-trimethoxyphenyl)acetyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-(5-chloranylthiophen-2-yl)-2-(3,4,5-trimethoxyphenyl)ethanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCB
 
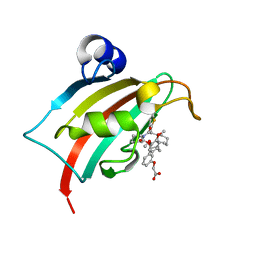 | | The Fk1 domain of FKBP51 in complex with 2-(3-((1R)-1-(((2S)-1-(2-(5-chlorothiophen-2-yl)-2-cyclohexylacetyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-(5-chloranylthiophen-2-yl)-2-cyclohexyl-ethanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCF
 
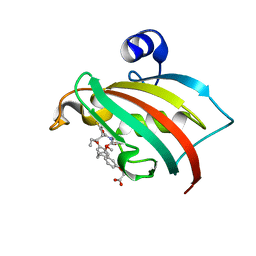 | | The Fk1 domain of FKBP51 in complex with 2-(3-((R)-1-(((S)-1-((S)-2-(5-chlorothiophen-2-yl)pent-4-enoyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-(5-chloranylthiophen-2-yl)pent-4-enoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCH
 
 | | The Fk1 domain of FKBP51 in complex with 2-(3-((1R)-1-(((2S)-1-(2-cyclohexyl-2-(thiophen-2-yl)acetyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-cyclohexyl-2-thiophen-2-yl-ethanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCG
 
 | | The Fk1 domain of FKBP51 in complex with (2R,5S,12S)-12-(thiophen-2-yl)-2-[2-(3,4-dimethoxyphenyl)ethyl]-15,15-dimethyl-3,19-dioxa-10,13,16-triazatricyclo[18.3.1.0^5,^10]tetracosa-1(24),20,22-triene-4,11,14,17-tetrone | | Descriptor: | (2~{R},5~{S},12~{S})-2-[2-(3,4-dimethoxyphenyl)ethyl]-15,15-dimethyl-12-thiophen-2-yl-3,19-dioxa-10,13,16-triazatricyclo[18.3.1.0^{5,10}]tetracosa-1(24),20,22-triene-4,11,14,17-tetrone, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCE
 | | The Fk1 domain of FKBP51 in complex with 2-(3-((R)-1-(((S)-1-((S)-2-(5-chlorothiophen-2-yl)butanoyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-(5-chloranylthiophen-2-yl)butanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, GLYCEROL, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
8CCC
 
 | | The Fk1 domain of FKBP51 in complex with 2-(3-((R)-1-(((S)-1-((S)-2-(5-chlorothiophen-2-yl)propanoyl)piperidine-2-carbonyl)oxy)-3-(3,4-dimethoxyphenyl)propyl)phenoxy)acetic acid | | Descriptor: | 2-[3-[(1~{R})-1-[(2~{S})-1-[(2~{S})-2-(5-chloranylthiophen-2-yl)propanoyl]piperidin-2-yl]carbonyloxy-3-(3,4-dimethoxyphenyl)propyl]phenoxy]ethanoic acid, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Meyners, C, Knaup, F.H, Walz, C.M, Hausch, F. | | Deposit date: | 2023-01-27 | | Release date: | 2023-04-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure-Based Discovery of a New Selectivity-Enabling Motif for the FK506-Binding Protein 51.
J.Med.Chem., 66, 2023
|
|
1QF6
 
 | | STRUCTURE OF E. COLI THREONYL-TRNA SYNTHETASE COMPLEXED WITH ITS COGNATE TRNA | | Descriptor: | ADENOSINE MONOPHOSPHATE, THREONINE TRNA, THREONYL-TRNA SYNTHETASE, ... | | Authors: | Sankaranarayanan, R, Dock-Bregeon, A.C, Rees, B, Moras, D. | | Deposit date: | 1999-04-06 | | Release date: | 1999-05-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The structure of threonyl-tRNA synthetase-tRNA(Thr) complex enlightens its repressor activity and reveals an essential zinc ion in the active site
Cell(Cambridge,Mass.), 97, 1999
|
|
1GYZ
 | |
9DNL
 
 | |
9DNC
 
 | |
9DN8
 
 | | RamR variant S2.3 complexed with 1S-1-phenyl-1,2,3,4-tetrahydroisoquinoline | | Descriptor: | (1S)-1-phenyl-1,2,3,4-tetrahydroisoquinoline, SULFATE ION, Transcriptional regulator RamR | | Authors: | Kim, W, Zhang, Y.J. | | Deposit date: | 2024-09-17 | | Release date: | 2026-04-08 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Using stereoselective biosensors to evolve asymmetric biocatalysts
To be published
|
|
9DNH
 
 | |
9DNK
 
 | | RamR variant R2.1 complexed with 1R-1-phenyl-1,2,3,4-tetrahydroisoquinoline | | Descriptor: | (1R)-1-phenyl-1,2,3,4-tetrahydroisoquinoline, Engineered RamR Variant R2.1 | | Authors: | Kim, W, Zhang, Y.J. | | Deposit date: | 2024-09-17 | | Release date: | 2026-04-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Using stereoselective biosensors to evolve asymmetric biocatalysts
To be published
|
|
9DN9
 
 | | RamR variant S2.4 complexed with 1S-1-phenyl-1,2,3,4-tetrahydroisoquinoline | | Descriptor: | (1S)-1-phenyl-1,2,3,4-tetrahydroisoquinoline, SULFATE ION, Transcriptional regulator RamR | | Authors: | Kim, W, Zhang, Y.J. | | Deposit date: | 2024-09-17 | | Release date: | 2026-04-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Using stereoselective biosensors to evolve asymmetric biocatalysts
To be published
|
|
