1HIL
 
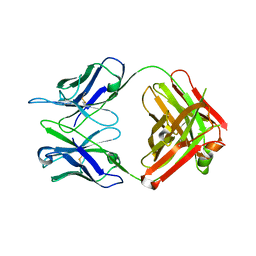 | |
1RIN
 
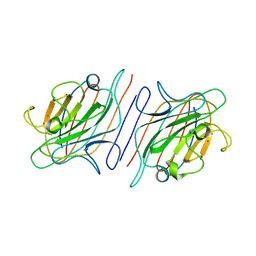 | | X-RAY CRYSTAL STRUCTURE OF A PEA LECTIN-TRIMANNOSIDE COMPLEX AT 2.6 ANGSTROMS RESOLUTION | | Descriptor: | CALCIUM ION, MANGANESE (II) ION, PEA LECTIN, ... | | Authors: | Rini, J.M, Hardman, K.D, Einspahr, H, Suddath, F.L, Carver, J.P. | | Deposit date: | 1993-01-27 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | X-ray crystal structure of a pea lectin-trimannoside complex at 2.6 A resolution.
J.Biol.Chem., 268, 1993
|
|
1HIN
 
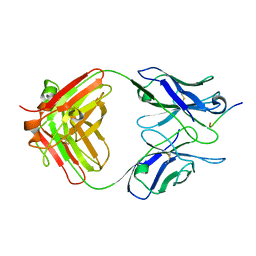 | |
1GGI
 
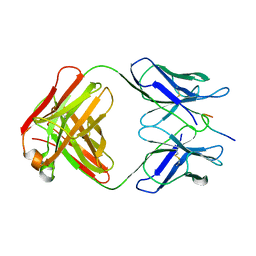 | | CRYSTAL STRUCTURE OF AN HIV-1 NEUTRALIZING ANTIBODY 50.1 IN COMPLEX WITH ITS V3 LOOP PEPTIDE ANTIGEN | | Descriptor: | HIV-1 V3 LOOP PEPTIDE ANTIGEN, IGG2A 50.1 FAB (HEAVY CHAIN), IGG2A 50.1 FAB (LIGHT CHAIN) | | Authors: | Stanfield, R.L, Rini, J.M, Wilson, I.A. | | Deposit date: | 1993-04-02 | | Release date: | 1993-10-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of a human immunodeficiency virus type 1 neutralizing antibody, 50.1, in complex with its V3 loop peptide antigen.
Proc.Natl.Acad.Sci.USA, 90, 1993
|
|
2AM4
 
 | |
2APC
 
 | |
2AM3
 
 | |
2AM5
 
 | |
9N13
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein D1 Domain (mutant T31VPR34 to GGGG) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.34 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N15
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein D1 Domain in complex with 9OAc-GD3(mutant T31VPR34 to GGGG) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N14
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein in complex with 9OAc-GD3(mutant T31VPR34 to GGGG) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.08 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N16
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein (Deletion 33Pro34Arg) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.01 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N12
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein (mutant T31VPR34 to GGGG) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.18 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N18
 
 | | Cryo-EM structure of HCoV-HKU1 spike glycoprotein in complex with 9OAc-GD3 tetrasacchride (Deletion 33Pro34Arg) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N17
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein D1 Domain (Deletion 33Pro34Arg) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9N19
 
 | | Cryo-EM structure of HCoV-HKU1spike glycoprotein D1 Domain in complex with 9OAc-GD3 tetrasacchride (Deletion 33Pro34Arg) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2025-01-24 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
6ATK
 
 | |
9BT9
 
 | | Cryo-EM Structure of HKU1 spike D1 Domain (Active state, locally refined) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.11 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BT1
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein in complex with 9O-acetyl GD3 sialoglycan (UUU state) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.22 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BTD
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein in complex with 9O-acetyl GD3 sialoglycan (DDA state) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.43 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BT2
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein D1 (Down State, locally refined) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (1.93 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BTA
 
 | | Cryo-EM density map of HKU1 spike glycoprotein D1 domain in complex with 9O-acetyl GD3 sialoglycan (Down_alt state, locally refined) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.39 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BTC
 
 | | Cryo-EM density map of HKU1 spike glycoprotein D1 domain in complex with 9O-acetyl GD3 sialoglycan (Up state, locally refined) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.41 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BSY
 
 | | Cryo-EM structure of HCoV-HKU1 glycoprotein in complex with 9O-acetyl GD3 sialoglycan (DAU state) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 5-acetamido-8-O-(5-acetamido-9-O-acetyl-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranonosyl)-3,5-dideoxy-D-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.58 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
9BSW
 
 | | Cryo-EM structure of HCoV-HKU1 Spike glycoprotein (DDA state) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Jin, M, Rini, J.M. | | Deposit date: | 2024-05-14 | | Release date: | 2025-05-14 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.06 Å) | | Cite: | Human coronavirus HKU1 spike structures reveal the basis for sialoglycan specificity and carbohydrate-promoted conformational changes.
Nat Commun, 16, 2025
|
|
