3IF7
 
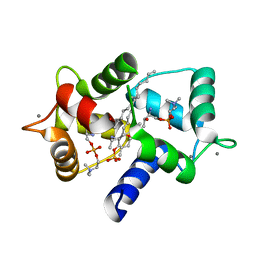 | | Structure of Calmodulin complexed with its first endogenous inhibitor, sphingosylphosphorylcholine | | Descriptor: | 2-{[(R)-{[(2S,3R,4E)-2-amino-3-hydroxyoctadec-4-en-1-yl]oxy}(hydroxy)phosphoryl]oxy}-N,N,N-trimethylethanaminium, CALCIUM ION, Calmodulin | | Authors: | Kovacs, E, Harmat, V, Toth, J, Vertessy, B.G, Modos, K, Kardos, J, Liliom, K. | | Deposit date: | 2009-07-24 | | Release date: | 2010-06-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and mechanism of calmodulin binding to a signaling sphingolipid reveal new aspects of lipid-protein interactions
Faseb J., 24, 2010
|
|
4AX4
 
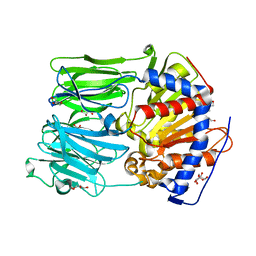 | | PROLYL OLIGOPEPTIDASE FROM PORCINE BRAIN, H680A MUTANT | | Descriptor: | GLYCEROL, PROLYL OLIGOPEPTIDASE | | Authors: | Szeltner, Z, Juhasz, T, Szamosi, I, Rea, D, Fulop, V, Modos, K, Juliano, L, Polgar, L. | | Deposit date: | 2012-06-08 | | Release date: | 2012-09-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The Loops Facing the Active Site of Prolyl Oligopeptidase are Crucial Components in Substrate Gating and Specificity.
Biochim.Biophys.Acta, 1834, 2012
|
|
