7PXS
 
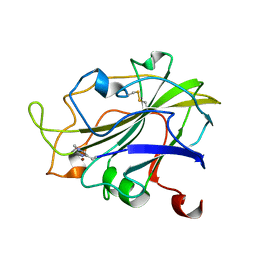 | | Room temperature X-ray structure of LPMO at 1.91x10^3 Gy | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Auxiliary activity 9, COPPER (II) ION | | 著者 | Tandrup, T, Muderspach, S.J, Banerjee, S, Lo Leggio, L. | | 登録日 | 2021-10-08 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PZ3
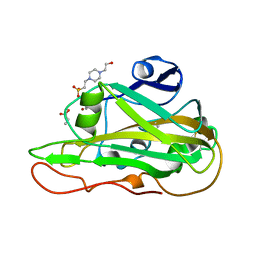 
 | | Structure of an LPMO at 5.37x10^3 Gy | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACRYLIC ACID, ... | | 著者 | Tandrup, T, Muderspach, S.J, Ipsen, J.O, Johansen, K.S, Lo Leggio, L. | | 登録日 | 2021-10-11 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7TOB
 
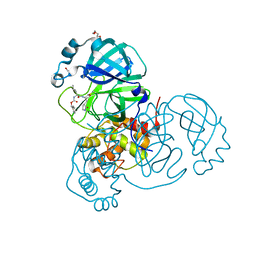 | | Crystal structure of the SARS-CoV-2 Omicron main protease (Mpro) in complex with inhibitor GC376 | | 分子名称: | (1S,2S)-2-({N-[(benzyloxy)carbonyl]-L-leucyl}amino)-1-hydroxy-3-[(3S)-2-oxopyrrolidin-3-yl]propane-1-sulfonic acid, 3C-like proteinase nsp5, DI(HYDROXYETHYL)ETHER | | 著者 | Sacco, M.D, Wang, J, Chen, Y. | | 登録日 | 2022-01-24 | | 公開日 | 2022-02-02 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | The P132H mutation in the main protease of Omicron SARS-CoV-2 decreases thermal stability without compromising catalysis or small-molecule drug inhibition.
Cell Res., 32, 2022
|
|
7TWJ
 
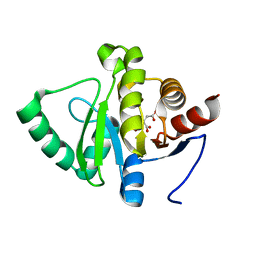 | |
7TWH
 
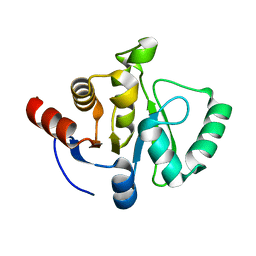 | |
7TWN
 
 | |
7TWS
 
 | |
7TWR
 
 | |
7TWQ
 
 | |
7TWI
 
 | |
7TWG
 
 | |
7TX4
 
 | |
7TWF
 
 | |
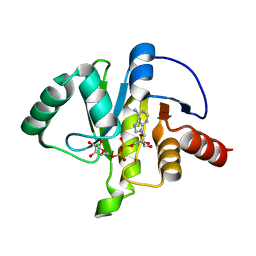
7TX5
 
 | |
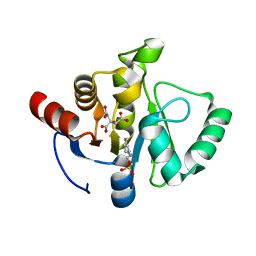
7TWO
 
 | |
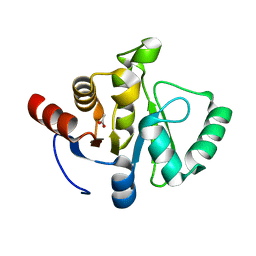
7TWP
 
 | |
5TY5
 
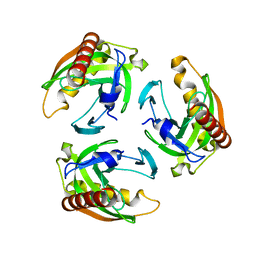 | | Neutron structure from microgravity-grown crystals of Inorganic Pyrophosphatase from Thermococcus theoreducens | | 分子名称: | Inorganic pyrophosphatase | | 著者 | Inoguchi, N, Coates, L, Morris, M.L, Singhal, A, Monaco, D.A, Garcia-Ruiz, J.M, Pusey, M.L, Ng, J.D. | | 登録日 | 2016-11-18 | | 公開日 | 2017-11-22 | | 最終更新日 | 2023-10-04 | | 実験手法 | NEUTRON DIFFRACTION (2.3 Å) | | 主引用文献 | Structure-function analysis of the neutron crystallographic structure of inorganic pyrophosphatase determined from microgravity-grown crystals
Acta Crystallogr.,Sect.A, 73, 2017
|
|
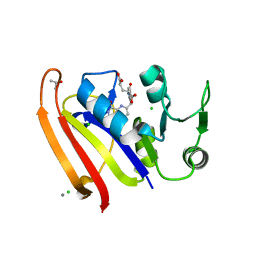
5UJX
 
 | |
7PQR
 
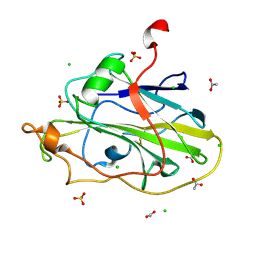 | | LsAA9A expressed in E. coli | | 分子名称: | ACETATE ION, Auxiliary activity 9, CHLORIDE ION, ... | | 著者 | Muderspach, S.J, Metherall, J, Ipsen, J, Rollan, C.H, Norholm, M, Johansen, K.S, Lo Leggio, L. | | 登録日 | 2021-09-20 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.3 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PXW
 
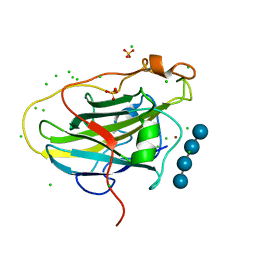 | | LPMO, expressed in E.coli, in complex with Cellotetraose | | 分子名称: | Auxiliary activity 9, CHLORIDE ION, COPPER (II) ION, ... | | 著者 | Banerjee, S, Muderspach, S.J, Tandrup, T, Ipsen, J, Rollan, C.H, Norholm, M, Johansen, K.S, Lo Leggio, L. | | 登録日 | 2021-10-08 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PXI
 
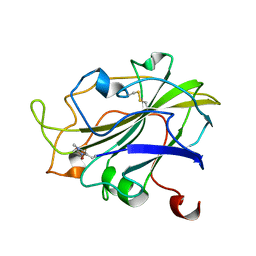 | | X-ray structure of LPMO at 7.88x10^3 Gy | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Auxiliary activity 9, COPPER (II) ION | | 著者 | Tandrup, T, Lo Leggio, L. | | 登録日 | 2021-10-08 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.63 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PXJ
 
 | | X-ray structure of LPMO at 5.99x10^4 Gy | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Auxiliary activity 9, COPPER (II) ION | | 著者 | Tandrup, T, Lo Leggio, L. | | 登録日 | 2021-10-08 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.75 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PYE
 
 | |
7PXK
 
 | | X-ray structure of LPMO at 1.39x10^5 Gy | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Auxiliary activity 9, COPPER (II) ION | | 著者 | Tandrup, T, Lo Leggio, L. | | 登録日 | 2021-10-08 | | 公開日 | 2022-08-24 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Changes in active-site geometry on X-ray photoreduction of a lytic polysaccharide monooxygenase active-site copper and saccharide binding.
Iucrj, 9, 2022
|
|
7PYD
 
 | |
