1DYK
 
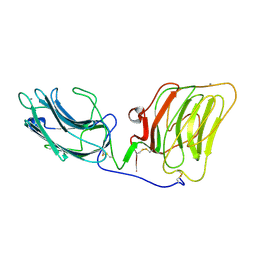 | | Laminin alpha 2 chain LG4-5 domain pair | | Descriptor: | CALCIUM ION, LAMININ ALPHA 2 CHAIN | | Authors: | Tisi, D, Talts, J.F, Timple, R, Hohenester, E. | | Deposit date: | 2000-02-01 | | Release date: | 2001-02-04 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the C-Terminal Laminin G-Like Domain Pair of the Laminin Alpha 2 Chain Harbouring Binding Sites for Alpha-Dystroglycan and Heparin
Embo J., 19, 2000
|
|
1H30
 
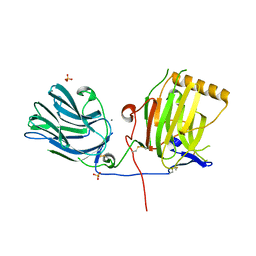 | | C-terminal LG domain pair of human Gas6 | | Descriptor: | CALCIUM ION, GROWTH-ARREST-SPECIFIC PROTEIN, SULFATE ION | | Authors: | Sasaki, T, Knyazev, P.G, Cheburkin, Y, Gohring, W, Tisi, D, Ullrich, A, Timpl, R, Hohenester, E. | | Deposit date: | 2002-08-21 | | Release date: | 2003-01-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of a Carboxy-Terminal Fragment of Growth-Arrest-Specific Protein Gas6: Receptor Tyrosine Kinase Activation by Laminin G-Like Domains
J.Biol.Chem., 277, 2002
|
|
1GL4
 
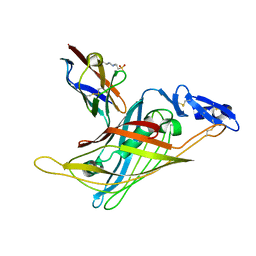 | | Nidogen-1 G2/Perlecan IG3 Complex | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, BASEMENT MEMBRANE-SPECIFIC HEPARAN SULFATE PROTEOGLYCAN CORE PROTEIN, NIDOGEN-1, ... | | Authors: | Kvansakul, M, Hopf, M, Ries, A, Timpl, R, Hohenester, E. | | Deposit date: | 2001-08-23 | | Release date: | 2001-11-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for the High-Affinity Interaction of Nidogen-1 with Immunoglobulin-Like Domain 3 of Perlecan
Embo J., 20, 2001
|
|
1UX6
 
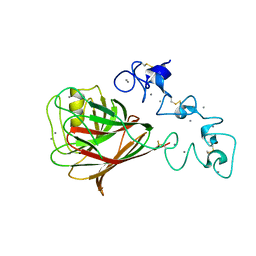 | |
2VRA
 
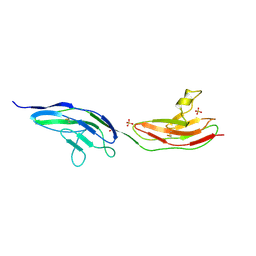 | | Drosophila Robo IG1-2 (monoclinic form) | | Descriptor: | 2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, ROUNDABOUT 1, SULFATE ION | | Authors: | Fukuhara, N, Howitt, J.A, Hussain, S, Hohenester, E. | | Deposit date: | 2008-03-28 | | Release date: | 2008-04-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural and Functional Analysis of Slit and Heparin Binding to Immunoglobulin-Like Domains 1 and 2 of Drosophila Robo
J.Biol.Chem., 283, 2008
|
|
2VKW
 
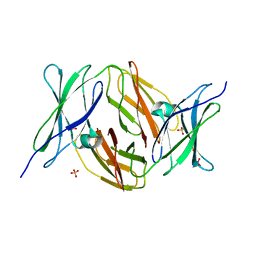 | | Human NCAM, FN3 domains 1 and 2 | | Descriptor: | NEURAL CELL ADHESION MOLECULE 1,140 KDA ISOFORM, SULFATE ION | | Authors: | Carafoli, F, Saffell, J.L, Hohenester, E. | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-26 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the Tandem Fibronectin Type 3 Domains of Neural Cell Adhesion Molecule
J.Mol.Biol., 377, 2008
|
|
6FZV
 
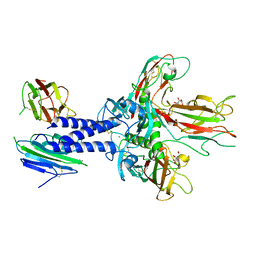 | |
2WJS
 
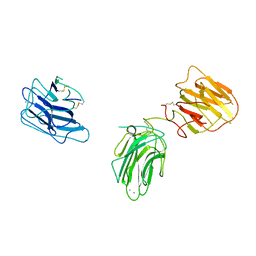 | |
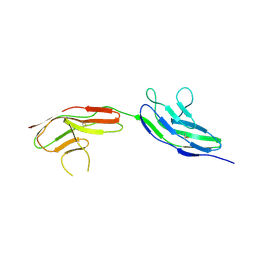
2VR9
 | |
2VKX
 
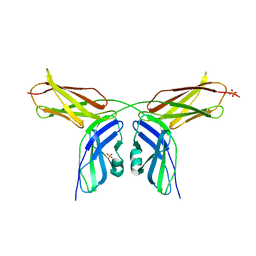 | | Human NCAM, FN3 domains 1 and 2, M610R mutant | | Descriptor: | NEURAL CELL ADHESION MOLECULE, SULFATE ION | | Authors: | Carafoli, F, Saffell, J.L, Hohenester, E. | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-26 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the Tandem Fibronectin Type 3 Domains of Neural Cell Adhesion Molecule
J.Mol.Biol., 377, 2008
|
|
1OAS
 
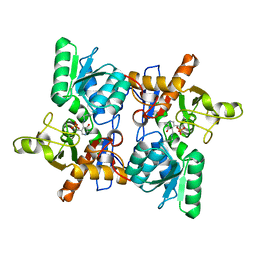 | | O-ACETYLSERINE SULFHYDRYLASE FROM SALMONELLA TYPHIMURIUM | | Descriptor: | O-ACETYLSERINE SULFHYDRYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Burkhard, P, Rao, G.S.J, Hohenester, E, Schnackerz, K.D, Cook, P.F, Jansonius, J.N. | | Deposit date: | 1999-01-29 | | Release date: | 2000-01-28 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Three-dimensional structure of O-acetylserine sulfhydrylase from Salmonella typhimurium.
J.Mol.Biol., 283, 1998
|
|
1O70
 
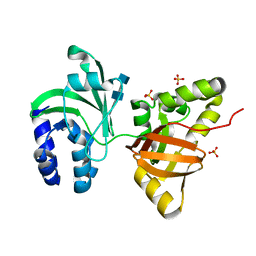 | |
6EJC
 
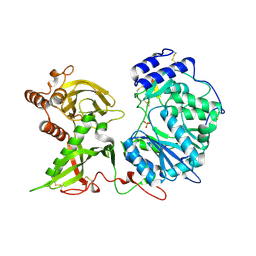 | |
6EJD
 
 | |
6EJB
 
 | |
6EJ7
 
 | |
6FOA
 
 | | Human Xylosyltransferase 1 apo structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PHOSPHATE ION, SODIUM ION, ... | | Authors: | Briggs, D.C, Hohenester, E. | | Deposit date: | 2018-02-06 | | Release date: | 2018-05-02 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.869 Å) | | Cite: | Structural Basis for the Initiation of Glycosaminoglycan Biosynthesis by Human Xylosyltransferase 1.
Structure, 26, 2018
|
|
6EJ9
 
 | |
6EJA
 
 | |
6EJ8
 
 | |
6EJE
 
 | |
1DKA
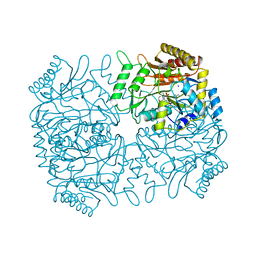 
 | | DIALKYLGLYCINE DECARBOXYLASE STRUCTURE: BIFUNCTIONAL ACTIVE SITE AND ALKALI METAL BINDING SITES | | Descriptor: | 2,2-DIALKYLGLYCINE DECARBOXYLASE (PYRUVATE), 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, POTASSIUM ION, ... | | Authors: | Toney, M.D, Hohenester, E, Jansonius, J.N. | | Deposit date: | 1993-06-18 | | Release date: | 1994-10-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Dialkylglycine decarboxylase structure: bifunctional active site and alkali metal sites.
Science, 261, 1993
|
|
2OAT
 
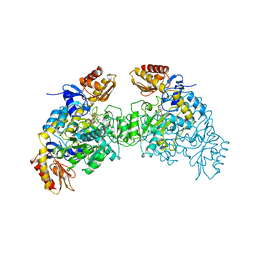 | | ORNITHINE AMINOTRANSFERASE COMPLEXED WITH 5-FLUOROMETHYLORNITHINE | | Descriptor: | 1-AMINO-7-(2-METHYL-3-OXIDO-5-((PHOSPHONOXY)METHYL)-4-PYRIDOXAL-5-OXO-6-HEPTENATE, ORNITHINE AMINOTRANSFERASE | | Authors: | Storici, P, Schirmer, T. | | Deposit date: | 1998-05-07 | | Release date: | 1998-12-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of human ornithine aminotransferase complexed with the highly specific and potent inhibitor 5-fluoromethylornithine.
J.Mol.Biol., 285, 1999
|
|
1OAT
 
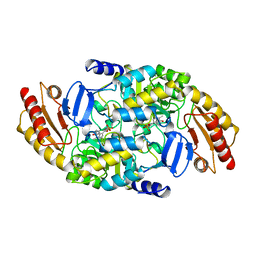 | | ORNITHINE AMINOTRANSFERASE | | Descriptor: | ORNITHINE AMINOTRANSFERASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Shen, B.W, Schirmer, T, Jansonius, J.N. | | Deposit date: | 1997-03-26 | | Release date: | 1998-04-01 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of human recombinant ornithine aminotransferase.
J.Mol.Biol., 277, 1998
|
|
1FCJ
 
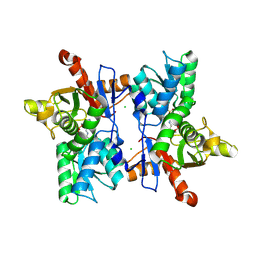 | | CRYSTAL STRUCTURE OF OASS COMPLEXED WITH CHLORIDE AND SULFATE | | Descriptor: | CHLORIDE ION, O-ACETYLSERINE SULFHYDRYLASE, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Burkhard, P, Tai, C, Jansonius, J.N, Cook, P.F. | | Deposit date: | 2000-07-18 | | Release date: | 2000-10-18 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Identification of an allosteric anion-binding site on O-acetylserine sulfhydrylase: structure of the enzyme with chloride bound.
J.Mol.Biol., 303, 2000
|
|
