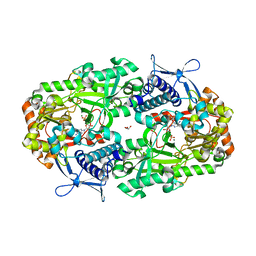
6P63
 
 | |
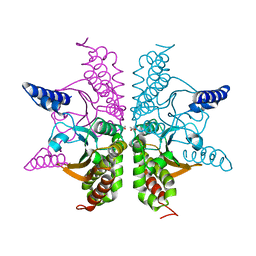
3MF3
 
 | |
4WAK
 
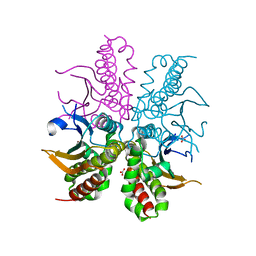 | |
6NL2
 
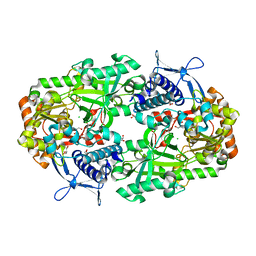 | | Apo NIS synthetase DesD variant R306Q | | Descriptor: | CHLORIDE ION, GLYCEROL, desferrioxamine E biosynthesis protein DesD | | Authors: | Hoffmann, K.M. | | Deposit date: | 2019-01-07 | | Release date: | 2020-01-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | A C-terminal Loop Mediates Cooperativity in the NIS Synthetase DesD
To Be Published
|
|
6XRC
 
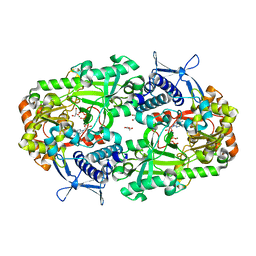 | | Apo NIS synthetase DesD variant R306Q | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Desferrioxamine E biosynthesis protein DesD, GLYCEROL, ... | | Authors: | Hoffmann, K.M. | | Deposit date: | 2020-07-11 | | Release date: | 2020-10-28 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Cofactor Complexes of DesD, a Model Enzyme in the Virulence-related NIS Synthetase Family.
Biochemistry, 59, 2020
|
|
1RPM
 
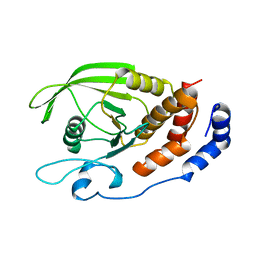 | |
4WAJ
 
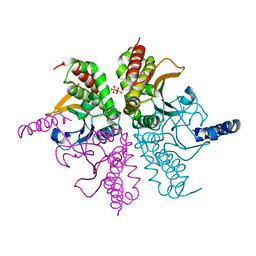 | |
3TSG
 
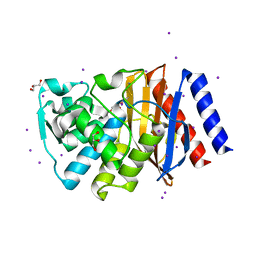 | | Crystal structure of GES-14 | | Descriptor: | Extended-spectrum beta-lactamase GES-14, GLYCEROL, IODIDE ION | | Authors: | Delbruck, H, Hoffmann, K.M.V, Bebrone, C. | | Deposit date: | 2011-09-13 | | Release date: | 2012-09-26 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Kinetic and crystallographic studies of extended-spectrum GES-11, GES-12, and GES-14 beta-lactamases.
Antimicrob.Agents Chemother., 56, 2012
|
|
3V3S
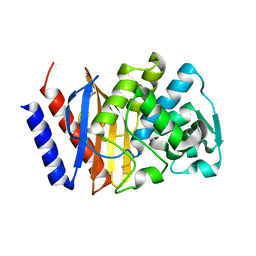 
 | | Crystal structure of GES-18 | | Descriptor: | Extended spectrum beta-lactamase GES-18 | | Authors: | Delbruck, H, Hoffmann, K.M.V, Bebrone, C. | | Deposit date: | 2011-12-14 | | Release date: | 2012-11-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | GES-18, a new carbapenem-hydrolyzing GES-Type beta-lactamase from pseudomonas aeruginosa that contains Ile80 and Ser170 residues.
Antimicrob.Agents Chemother., 57, 2013
|
|
3V3R
 
 | | Crystal Structure of GES-11 | | Descriptor: | Extended spectrum class A beta-lactamase GES-11, IODIDE ION, SODIUM ION | | Authors: | Delbruck, H, Hoffmann, K.M.V, Bebrone, C. | | Deposit date: | 2011-12-14 | | Release date: | 2012-09-26 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Kinetic and crystallographic studies of extended-spectrum GES-11, GES-12, and GES-14 beta-lactamases.
Antimicrob.Agents Chemother., 56, 2012
|
|
4FR7
 
 | |
4FSB
 
 | |
1T2J
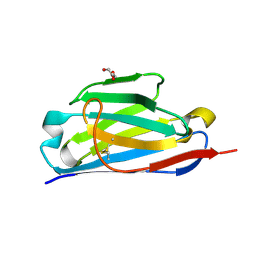 
 | |
3SW3
 
 | |
3T9M
 
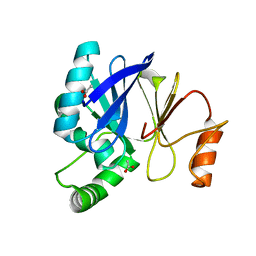 | |
3F9O
 
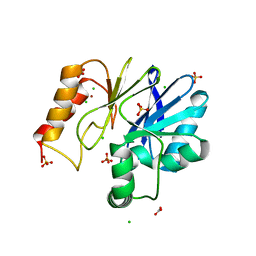 | |
3FAI
 
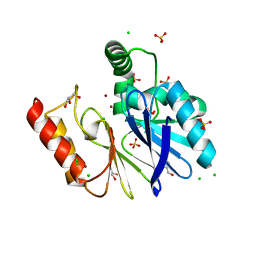 | | The Di Zinc Carbapenemase CphA N220G mutant | | Descriptor: | Beta-lactamase, CHLORIDE ION, GLYCEROL, ... | | Authors: | Delbruck, H, Hoffmann, K.M.V. | | Deposit date: | 2008-11-17 | | Release date: | 2009-09-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The structure of the dizinc subclass B2 metallo-beta-lactamase CphA reveals that the second inhibitory zinc ion binds in the histidine site
Antimicrob.Agents Chemother., 53, 2009
|
|
4WAM
 
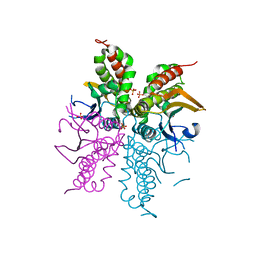 | |
3E2X
 
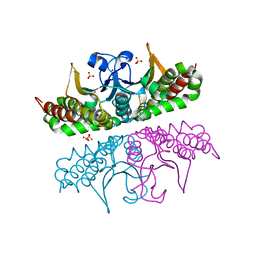 | | H. influenzae beta-carbonic anhydrase, variant V47A | | Descriptor: | Carbonic anhydrase 2, SULFATE ION, ZINC ION | | Authors: | Rowlett, R.S, Lee, J. | | Deposit date: | 2008-08-06 | | Release date: | 2009-08-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Evidence for a bicarbonate "escort" site in Haemophilus influenzae beta-carbonic anhydrase .
Biochemistry, 49, 2010
|
|
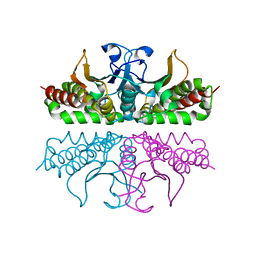
3E31
 
 | |
3E3I
 
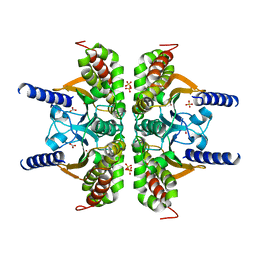 | |
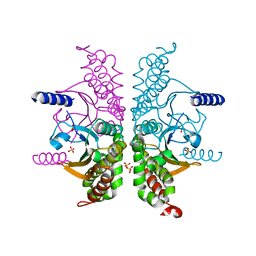
3E3G
 
 | |
3E3F
 
 | |
4J6D
 
 | |
4J6B
 
 | |
