6KKJ
 
 | |
7W8S
 
 | | Structure of SARS-CoV-2 spike receptor-binding domain Y453F mutation complexed with American mink ACE2 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme 2, Spike protein S1, ... | | 著者 | Su, C, Qi, J.X, Gao, G.F. | | 登録日 | 2021-12-08 | | 公開日 | 2022-08-17 | | 最終更新日 | 2023-04-12 | | 実験手法 | ELECTRON MICROSCOPY (2.85 Å) | | 主引用文献 | Molecular Basis of Mink ACE2 Binding to SARS-CoV-2 and Its Mink-Derived Variants.
J.Virol., 96, 2022
|
|
7WA1
 
 | |
5HTL
 
 | | Structure of MshE with cdg | | 分子名称: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), MSHA biogenesis protein MshE | | 著者 | Chin, K.H, Wang, Y.C. | | 登録日 | 2016-01-27 | | 公開日 | 2016-10-05 | | 最終更新日 | 2016-10-12 | | 実験手法 | X-RAY DIFFRACTION (1.371 Å) | | 主引用文献 | Nucleotide binding by the widespread high-affinity cyclic di-GMP receptor MshEN domain.
Nat Commun, 7, 2016
|
|
8HI6
 
 | |
8HI5
 
 | |
4P1J
 
 | | Crystal structure of wild type Hypocrea jecorina Cel7a in a hexagonal crystal form | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Exoglucanase 1, SAMARIUM (III) ION, ... | | 著者 | Bodenheimer, A.B, Cuneo, M.J, Swartz, P.D, Myles, D.A, Meilleur, F. | | 登録日 | 2014-02-26 | | 公開日 | 2015-03-25 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2.62 Å) | | 主引用文献 | Crystal structure of wild type Hypocrea jecorina Cel7a in a hexagonal crystal form
To Be Published
|
|
8IHO
 
 | | Crystal structures of SARS-CoV-2 papain-like protease in complex with covalent inhibitors | | 分子名称: | Papain-like protease nsp3, ZINC ION, covalent inhibitor | | 著者 | Wang, Q, Hu, H, Li, M, Xu, Y. | | 登録日 | 2023-02-23 | | 公開日 | 2024-01-03 | | 実験手法 | X-RAY DIFFRACTION (2.55 Å) | | 主引用文献 | Structure-Based Design of Potent Peptidomimetic Inhibitors Covalently Targeting SARS-CoV-2 Papain-like Protease.
Int J Mol Sci, 24, 2023
|
|
4PPM
 
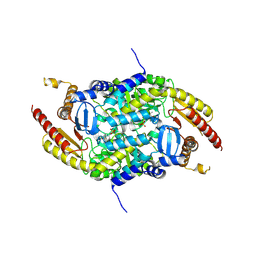 | | Crystal structure of PigE: a transaminase involved in the biosynthesis of 2-methyl-3-n-amyl-pyrrole (MAP) from Serratia sp. FS14 | | 分子名称: | Aminotransferase, MAGNESIUM ION, PYRIDOXAL-5'-PHOSPHATE | | 著者 | Lou, X.D, Ran, T.T, Xu, D.Q, Wang, W.W. | | 登録日 | 2014-02-27 | | 公開日 | 2015-01-14 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal structure of the catalytic domain of PigE: a transaminase involved in the biosynthesis of 2-methyl-3-n-amyl-pyrrole (MAP) from Serratia sp. FS14
Biochem.Biophys.Res.Commun., 447, 2014
|
|
1AKI
 
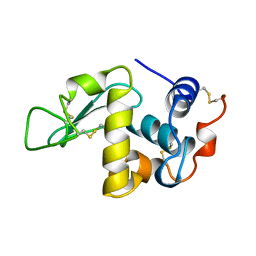 | | THE STRUCTURE OF THE ORTHORHOMBIC FORM OF HEN EGG-WHITE LYSOZYME AT 1.5 ANGSTROMS RESOLUTION | | 分子名称: | LYSOZYME | | 著者 | Carter, D, He, J, Ruble, J.R, Wright, B. | | 登録日 | 1997-05-19 | | 公開日 | 1997-11-19 | | 最終更新日 | 2023-08-02 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | The Structures of the Monoclinic and Orthorhombic Forms of Hen Egg-White Lysozyme at 6 Angstroms Resolution
Acta Crystallogr.,Sect.B, 38, 1982
|
|
7V3T
 
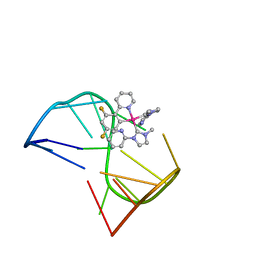 | | Solution structure of thrombin binding aptamer G-quadruplex bound a self-adaptive small molecule with rotated ligands | | 分子名称: | 11,13-bis(fluoranyl)-8-(1-methyl-3-pyridin-2-yl-imidazol-2-yl)-8-(1-methyl-3-pyridin-2-yl-imidazol-2-yl)-7$l^{4}-aza-8$l^{4}-platinatricyclo[7.4.0.0^{2,7}]trideca-1(9),2(7),3,5,10,12-hexaene, TBA G4 DNA (5'-D(*GP*GP*TP*TP*GP*GP*TP*GP*TP*GP*GP*TP*TP*GP*G)-3') | | 著者 | Liu, W, Zhu, B.C, Mao, Z.W. | | 登録日 | 2021-08-11 | | 公開日 | 2022-09-28 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of a thrombin binding aptamer complex with a non-planar platinum(ii) compound.
Chem Sci, 13, 2022
|
|
6M31
 | |
6LNZ
 
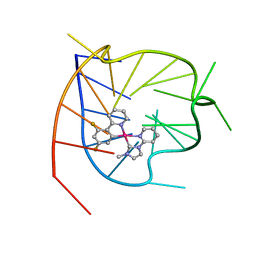 | |
2LKG
 | | WSA major conformation | | 分子名称: | Acetylcholine receptor | | 著者 | Xu, Y, Mowrey, D, Cui, T, Perez-Aguilar, J.M, Saven, J.G, Eckenhoff, R, Tang, P. | | 登録日 | 2011-10-11 | | 公開日 | 2012-01-11 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | NMR structure and dynamics of a designed water-soluble transmembrane domain of nicotinic acetylcholine receptor.
Biochim.Biophys.Acta, 1818, 2011
|
|
8H6C
 
 | |
8H6D
 
 | |
8H65
 
 | |
8H66
 
 | |
5YEJ
 
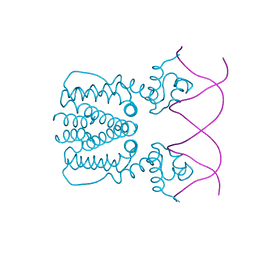 | | Crystal structure of BioQ with its naturel double-stranded DNA operator | | 分子名称: | DNA (5'-D(*AP*CP*CP*TP*GP*AP*AP*CP*AP*CP*CP*GP*TP*TP*CP*AP*AP*GP*T)-3'), DNA (5'-D(*AP*CP*TP*TP*GP*AP*AP*CP*GP*GP*TP*GP*TP*TP*CP*AP*GP*GP*T)-3'), TetR family transcriptional regulator | | 著者 | Yan, L, Guan, Z.Y, Zou, T.T. | | 登録日 | 2017-09-17 | | 公開日 | 2018-09-26 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.698 Å) | | 主引用文献 | Structural insights into operator recognition by BioQ in the Mycobacterium smegmatis biotin synthesis pathway.
Biochim Biophys Acta Gen Subj, 1862, 2018
|
|
5YEK
 
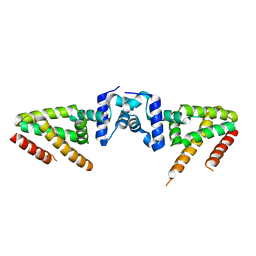 | | Crystal structure of BioQ | | 分子名称: | TetR family transcriptional regulator | | 著者 | Yan, L, Guan, Z.Y, Zou, T.T. | | 登録日 | 2017-09-17 | | 公開日 | 2018-09-26 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.191 Å) | | 主引用文献 | Structural insights into operator recognition by BioQ in the Mycobacterium smegmatis biotin synthesis pathway.
Biochim Biophys Acta Gen Subj, 1862, 2018
|
|
5Y5R
 
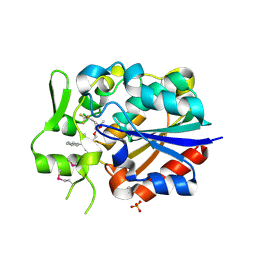 | | Crystal structure of a novel Pyrethroid Hydrolase PytH with BIF | | 分子名称: | (2-methyl-3-phenyl-phenyl)methyl (1~{S})-3-[(~{E})-2-chloranyl-3,3,3-tris(fluoranyl)prop-1-enyl]-2,2-dimethyl-cyclopropane-1-carboxylate, Pyrethroid hydrolase, SULFATE ION | | 著者 | Xu, D.Q, Ran, T.T, Wang, W.W. | | 登録日 | 2017-08-09 | | 公開日 | 2018-08-15 | | 実験手法 | X-RAY DIFFRACTION (1.899 Å) | | 主引用文献 | Structure and Catalytic Mechanism of a Novel Pyrethroid Hydrolase from Sphingobium faniae JZ-2
To Be Published
|
|
5ZJK
 
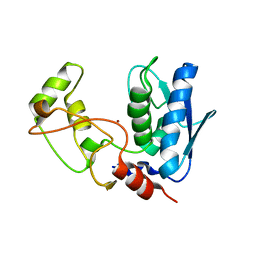 | | Structure of myroilysin | | 分子名称: | Myroilysin, PHOSPHATE ION, ZINC ION | | 著者 | Li, W.D, Ran, T.T, Xu, D.Q, Wang, W.W. | | 登録日 | 2018-03-20 | | 公開日 | 2019-03-20 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Crystal structure of mature myroilysin and implication for its activation mechanism.
Int.J.Biol.Macromol., 2019
|
|
6AEO
 
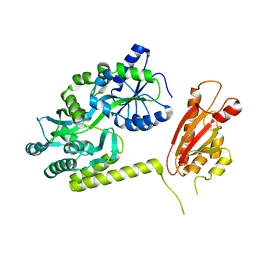 | | TssL periplasmic domain | | 分子名称: | GLYCEROL, Maltose/maltodextrin-binding periplasmic protein,TssL | | 著者 | Ran, T.T, Wang, W.W, Wang, X.B, Xu, D.Q. | | 登録日 | 2018-08-06 | | 公開日 | 2019-06-12 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal structure of the periplasmic domain of TssL, a key membrane component of Type VI secretion system.
Int.J.Biol.Macromol., 120, 2018
|
|
7DCD
 
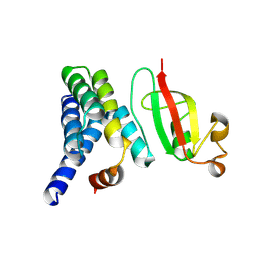 | |
7EFT
 | |
