9DFQ
 
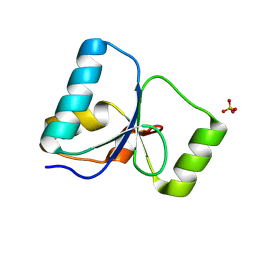 | | PARP4 BRCT domain F39Q mutant | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4, SULFATE ION | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2024-08-30 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure and mutagenesis of a nucleic acid-binding BRCT domain in human PARP4
J.Biol.Chem., 2025
|
|
9DEV
 
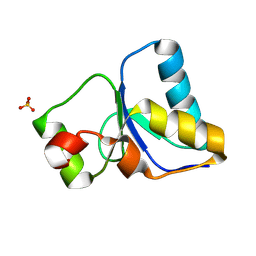 | | PARP4 BRCT domain | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4, SULFATE ION | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2024-08-29 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure and mutagenesis of a nucleic acid-binding BRCT domain in human PARP4
J.Biol.Chem., 2025
|
|
9DFO
 
 | | PARP4 BRCT domain K31Q mutant | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4, SULFATE ION | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2024-08-30 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure and mutagenesis of a nucleic acid-binding BRCT domain in human PARP4
J.Biol.Chem., 2025
|
|
9DFP
 
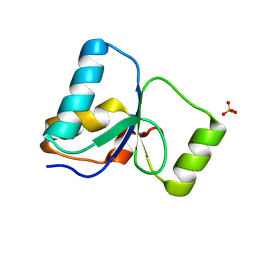 | | PARP4 BRCT domain K23/24Q mutant | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4, SULFATE ION | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2024-08-30 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Crystal structure and mutagenesis of a nucleic acid-binding BRCT domain in human PARP4
J.Biol.Chem., 2025
|
|
9DFR
 
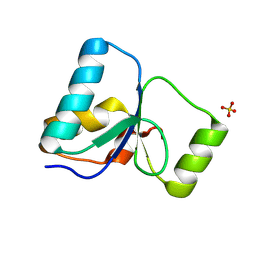 | | PARP4 BRCT domain F39A mutant | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4, SULFATE ION | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2024-08-30 | | Release date: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure and mutagenesis of a nucleic acid-binding BRCT domain in human PARP4
J.Biol.Chem., 2025
|
|
8SWZ
 
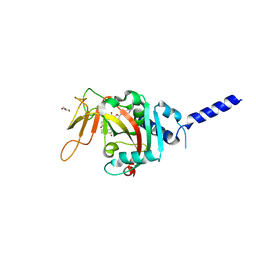 | | PARP4 ART domain bound to EB47 | | Descriptor: | 2-[4-[(2S,3S,4R,5R)-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]carbonylpiperazin-1-yl]-N-(1-oxidanylidene-2,3-dihydroisoindol-4-yl)ethanamide, GLYCEROL, Protein mono-ADP-ribosyltransferase PARP4 | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2023-05-19 | | Release date: | 2023-11-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP.
Nucleic Acids Res., 51, 2023
|
|
8SWY
 
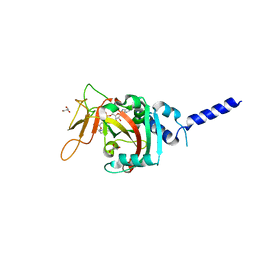 | | PARP4 ART domain bound to NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, GLYCEROL, Protein mono-ADP-ribosyltransferase PARP4 | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2023-05-19 | | Release date: | 2023-11-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP.
Nucleic Acids Res., 51, 2023
|
|
8SX1
 
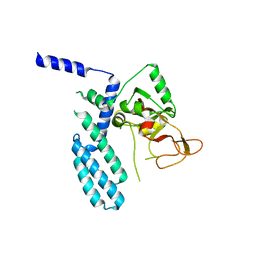 | | PARP4 catalytic domain | | Descriptor: | Protein mono-ADP-ribosyltransferase PARP4 | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2023-05-19 | | Release date: | 2023-11-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (4.2 Å) | | Cite: | Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP.
Nucleic Acids Res., 51, 2023
|
|
8SX2
 
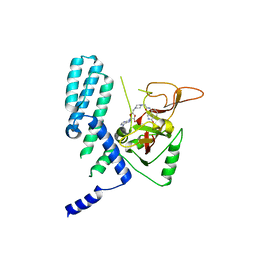 | | PARP4 catalytic domain bound to EB47 | | Descriptor: | 2-[4-[(2S,3S,4R,5R)-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]carbonylpiperazin-1-yl]-N-(1-oxidanylidene-2,3-dihydroisoindol-4-yl)ethanamide, Protein mono-ADP-ribosyltransferase PARP4 | | Authors: | Frigon, L, Pascal, J.M. | | Deposit date: | 2023-05-19 | | Release date: | 2023-11-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP.
Nucleic Acids Res., 51, 2023
|
|
8SK0
 
 | | Crystal structure of EvdS6 decarboxylase in ligand bound state | | Descriptor: | CITRATE ANION, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Sharma, P, Frigo, L, Dulin, C.C, Bachmann, B.O, Iverson, T.M. | | Deposit date: | 2023-04-18 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | EvdS6 is a bifunctional decarboxylase from the everninomicin gene cluster.
J.Biol.Chem., 299, 2023
|
|
8SHH
 
 | | Crystal structure of EvdS6 decarboxylase in ligand free state | | Descriptor: | DI(HYDROXYETHYL)ETHER, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, dTDP-glucose 4,6-dehydratase | | Authors: | Sharma, P, Frigo, L, Dulin, C.C, Bachmann, B.O, Iverson, T.M. | | Deposit date: | 2023-04-14 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | EvdS6 is a bifunctional decarboxylase from the everninomicin gene cluster.
J.Biol.Chem., 299, 2023
|
|
