6G8F
 
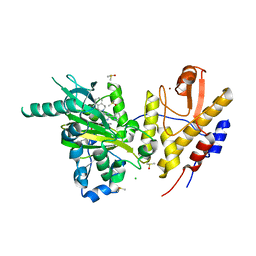 | | Crystal structure of UTX complexed with GSK-J1 | | Descriptor: | 3-[[2-pyridin-2-yl-6-(1,2,4,5-tetrahydro-3-benzazepin-3-yl)pyrimidin-4-yl]amino]propanoic acid, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Esposito, C, Sledz, P, Caflisch, A. | | Deposit date: | 2018-04-08 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.043 Å) | | Cite: | In Silico Identification of JMJD3 Demethylase Inhibitors.
J Chem Inf Model, 58, 2018
|
|
6FUK
 
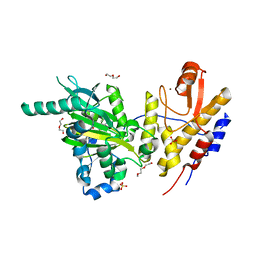 | | Crystal structure of UTX complexed with 5-carboxy-8-hydroxyquinoline | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, 8-hydroxyquinoline-5-carboxylic acid, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Esposito, C, Sledz, P, Caflisch, A. | | Deposit date: | 2018-02-27 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | In Silico Identification of JMJD3 Demethylase Inhibitors.
J Chem Inf Model, 58, 2018
|
|
6FUL
 
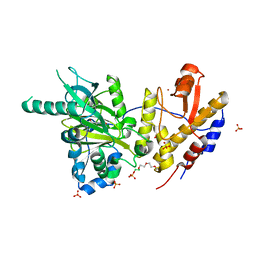 | | Crystal structure of UTX complexed with 5-hydroxy-4-keto-1-methyl-picolinate | | Descriptor: | 1-methyl-5-oxidanyl-4-oxidanylidene-pyridine-2-carboxylic acid, 2-(2-METHOXYETHOXY)ETHANOL, Lysine-specific demethylase 6A, ... | | Authors: | Esposito, C, Sledz, P, Caflisch, A. | | Deposit date: | 2018-02-27 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.649 Å) | | Cite: | In Silico Identification of JMJD3 Demethylase Inhibitors.
J Chem Inf Model, 58, 2018
|
|
2JSH
 
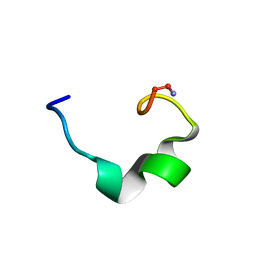 | | obestatin NMR structure in SDS/DPC micellar solution | | Descriptor: | Appetite-regulating hormone, Obestatin | | Authors: | D'Ursi, A.M, Scrima, M, Esposito, C, Campiglia, P. | | Deposit date: | 2007-07-05 | | Release date: | 2008-10-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Obestatin conformational features: a strategy to unveil obestatin's biological role?
Biochem.Biophys.Res.Commun., 363, 2007
|
|
2JSJ
 
 | | Obestatin in water solution | | Descriptor: | Appetite-regulating hormone, Obestatin | | Authors: | D'Ursi, A.M, Scrima, M, Esposito, C, Campiglia, P. | | Deposit date: | 2007-07-05 | | Release date: | 2008-10-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Obestatin conformational features: a strategy to unveil obestatin's biological role?
Biochem.Biophys.Res.Commun., 363, 2007
|
|
2JSI
 
 | | 11-23 obestatin fragment in DPC/SDS micellar solution | | Descriptor: | Appetite-regulating hormone, Obestatin | | Authors: | D'Ursi, A.M, Scrima, M, Esposito, C, Campiglia, P. | | Deposit date: | 2007-07-05 | | Release date: | 2008-10-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Obestatin conformational features: a strategy to unveil obestatin's biological role?
Biochem.Biophys.Res.Commun., 363, 2007
|
|
2O00
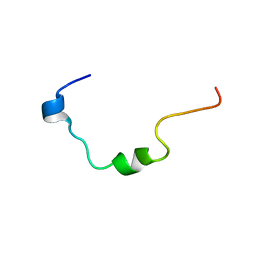 
 | | NMR structure analysis of the Penetratin conjugated Gas (374-394) peptide | | Descriptor: | Penetratin conjugated Gas (374-394) peptide | | Authors: | D'Ursi, A.M, Rovero, P, Albrizio, S, Espsito, C, D'Errico, G, Novellino, E. | | Deposit date: | 2006-11-27 | | Release date: | 2007-06-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Driving forces in the delivery of penetratin conjugated G protein fragment.
J.Med.Chem., 50, 2007
|
|
2NZZ
 
 | | NMR structure analysis of the Penetratin conjugated Gas (374-394) peptide | | Descriptor: | Penetratin conjugated Gas (374-394) peptide | | Authors: | D'Ursi, A.M, Rovero, P, Albrizio, S, Espsito, C, D'Errico, G, Novellino, E. | | Deposit date: | 2006-11-27 | | Release date: | 2007-06-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Driving forces in the delivery of penetratin conjugated G protein fragment.
J.Med.Chem., 50, 2007
|
|
6XVW
 
 | |
