1KYJ
 
 | | Tumor Associated Mucin Motif from CD43 protein | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, Leukosialin (CD43) fragment | | Authors: | Coltart, D.M, Williams, L.J, Glunz, P.W, Sames, D, Kuduk, S.D, Schwarz, J.B, Chen, X.-T, Royyuru, A.K, Danishefsky, S.D, Live, D.H. | | Deposit date: | 2002-02-04 | | Release date: | 2002-02-20 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Principles of Mucin Architecture: Structural Studies on Synthetic Glycopeptides Bearing Clustered Mono-, Di-, Tri-, and Hexasaccharide Glycodomains
J.Am.Chem.Soc., 124, 2002
|
|
3C1X
 
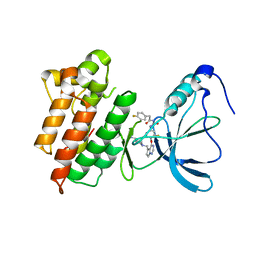 | | Crystal structure of the tyrosine kinase domain of the hepatocyte growth factor receptor c-MET in complex with a Pyrrolotriazine based inhibitor | | Descriptor: | Hepatocyte growth factor receptor, N-{[4-({5-[(4-aminopiperidin-1-yl)methyl]pyrrolo[2,1-f][1,2,4]triazin-4-yl}oxy)-3-fluorophenyl]carbamoyl}-2-(4-fluorophenyl)acetamide | | Authors: | Sack, J. | | Deposit date: | 2008-01-24 | | Release date: | 2008-03-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Identification of pyrrolo[2,1-f][1,2,4]triazine-based inhibitors of Met kinase.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3F82
 
 | |
3L8V
 
 | |
6JQW
 
 | |
6JQX
 
 | |
